Showing preview only (659K chars total). Download the full file or copy to clipboard to get everything.
Repository: MKLab-ITI/CUDA
Branch: master
Commit: e567d31391d5
Files: 59
Total size: 33.1 MB
Directory structure:
gitextract_hg6xup5t/
├── README.md
├── binaries/
│ ├── linux/
│ │ ├── COPYRIGHT
│ │ ├── FAQ.html
│ │ ├── README
│ │ ├── README-GPU
│ │ ├── svm-predict
│ │ ├── svm-scale
│ │ ├── svm-train
│ │ ├── svm-train-gpu
│ │ ├── tools/
│ │ │ ├── README
│ │ │ ├── checkdata.py
│ │ │ ├── easy.py
│ │ │ ├── grid.py
│ │ │ └── subset.py
│ │ └── train_set
│ └── windows/
│ ├── x64/
│ │ ├── COPYRIGHT
│ │ ├── FAQ.html
│ │ ├── README
│ │ ├── README-GPU
│ │ ├── tools/
│ │ │ ├── README
│ │ │ ├── checkdata.py
│ │ │ ├── easy.py
│ │ │ ├── grid.py
│ │ │ └── subset.py
│ │ └── train_set
│ └── x86/
│ ├── COPYRIGHT
│ ├── FAQ.html
│ ├── README
│ ├── README-GPU
│ ├── tools/
│ │ ├── README
│ │ ├── checkdata.py
│ │ ├── easy.py
│ │ ├── grid.py
│ │ └── subset.py
│ └── train_set
└── src/
├── linux/
│ ├── COPYRIGHT
│ ├── Makefile
│ ├── README
│ ├── README-GPU
│ ├── cross_validation_with_matrix_precomputation.c
│ ├── findcudalib.mk
│ ├── kernel_matrix_calculation.c
│ ├── readme.txt
│ ├── svm-train.c
│ ├── svm.cpp
│ └── svm.h
└── windows/
├── README-GPU
├── libsvm_train_dense_gpu/
│ ├── cross_validation_with_matrix_precomputation.c
│ ├── kernel_matrix_calculation.c
│ ├── libsvm_train_dense_gpu.vcxproj
│ ├── libsvm_train_dense_gpu.vcxproj.filters
│ ├── libsvm_train_dense_gpu.vcxproj.user
│ ├── svm-train.c
│ ├── svm.cpp
│ └── svm.h
├── libsvm_train_dense_gpu.ncb
├── libsvm_train_dense_gpu.sdf
├── libsvm_train_dense_gpu.sln
└── libsvm_train_dense_gpu.suo
================================================
FILE CONTENTS
================================================
================================================
FILE: README.md
================================================
CUDA: GPU-accelerated LIBSVM
====
**LIBSVM Accelerated with GPU using the CUDA Framework**
GPU-accelerated LIBSVM is a modification of the [original LIBSVM](http://www.csie.ntu.edu.tw/~cjlin/libsvm/) that exploits the CUDA framework to significantly reduce processing time while producing identical results. The functionality and interface of LIBSVM remains the same. The modifications were done in the kernel computation, that is now performed using the GPU.
Watch a [short video](http://www.youtube.com/watch?v=Fl99tQQd55U) on the capabilities of the GPU-accelerated LIBSVM package
###CHANGELOG
V1.2
Updated to LIBSVM version 3.17
Updated to CUDA SDK v5.5
Using CUBLAS_V2 which is compatible with the CUDA SDK v4.0 and up.
### FEATURES
Mode Supported
C-SVC classification with RBF kernel
Functionality / User interface
Same as LIBSVM
### PREREQUISITES
LIBSVM prerequisites
NVIDIA Graphics card with CUDA support
Latest NVIDIA drivers for GPU
### PERFORMANCE COMPARISON
To showcase the performance gain using the GPU-accelerated LIBSVM we present an example run.
PC Setup
Quad core Intel Q6600 processor
3.5GB of DDR2 RAM
Windows-XP 32-bit OS
Input Data
TRECVID 2007 Dataset for the detection of high level features in video shots
Training vectors with a dimension of 6000
20 different feature models with a variable number of input training vectors ranging from 36 up to 3772
Classification parameters
c-svc
RBF kernel
Parameter optimization using the easy.py script provided by LIBSVM.
4 local workers
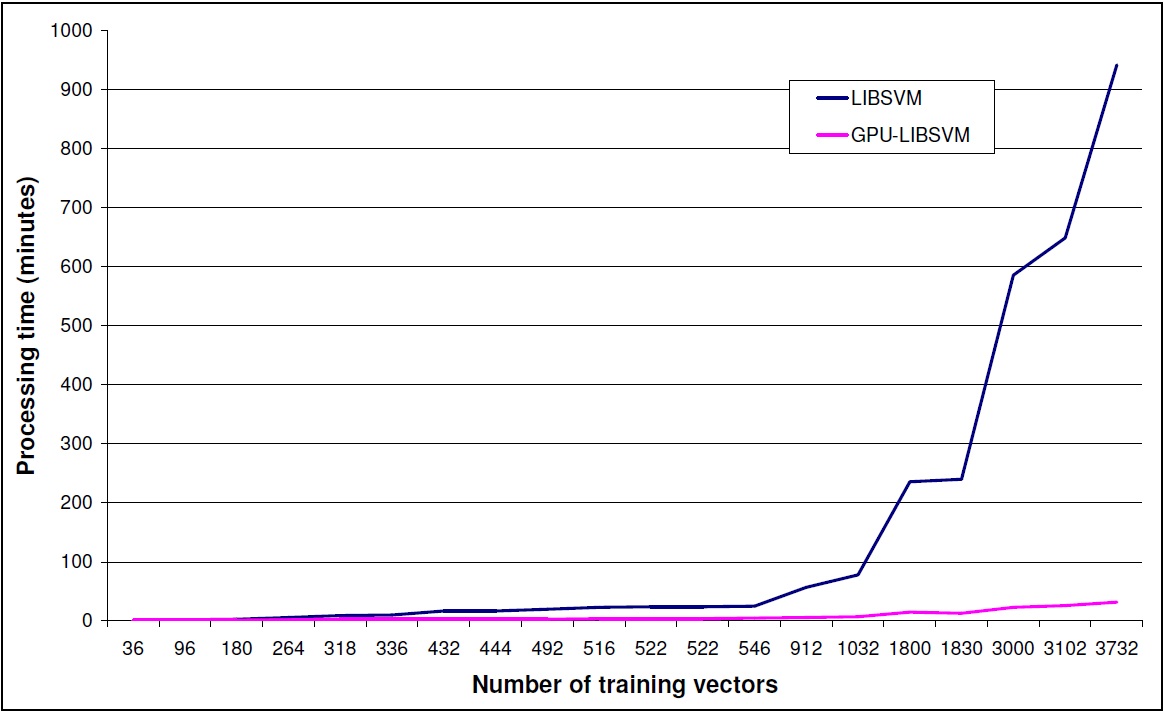
Discussion
GPU-accelerated LIBSVM gives a performance gain depending on the size of input data set.
This gain is increasing dramatically with the size of the dataset.
Please take into consideration input data size limitations that can occur from the memory
capacity of the graphics card that is used.
### PUBLICATION
A first document describing some of the work related to the GPU-Accelerated LIBSVM is the following; please cite it if you find this implementation useful in your work:
A. Athanasopoulos, A. Dimou, V. Mezaris, I. Kompatsiaris, "GPU Acceleration for Support Vector Machines", Proc. 12th International Workshop on Image Analysis for Multimedia Interactive Services (WIAMIS 2011), Delft, The Netherlands, April 2011.
### LICENSE
THIS SOFTWARE IS PROVIDED BY THE AUTHOR "AS IS" AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE AUTHOR BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
### FREQUENTLY ASKED QUESTIONS (FAQ)
* Is there a GPU-accelerated LIBSVM version for Matlab?
We are interested in porting our imlementation in Matlab but due to our workload, it is not in our immediate plans. Everybody is welcome to make the port and we can host the ported software.
* Visual Studio will not let me build the provided project.
The project has been built in both 32/64 bit mode. If you are working on 32 bits, you might need the x64 compiler for Visual Studio 2010 to build the project.
* Building the project, I get linker error messages.
Please go to the project properties and check the library settings. CUDA libraries have different names for x32 / x64. Make sure that the correct path and filenames are given.
* I have built the project but the executables will not run (The application has failed to start because its side-by-side configuration is in
correct.)
Please update the VS2010 redistributables to the PC you are running your executable and install all the latest patches for visual studio.
* My GPU-accelerated LIBSVM is running smoothly but i do not see any speed-up.
GPU-accelerated LIBSVM is giving speed-ups mainly for big datasets. In the GPU-accelerated implementation some extra time is needed to load the data to the gpu memory. If the dataset is not big enough to give a significant performance gain, the gain is lost due to the gpu-memory -> cpu-memory, cpu-memory -> gpu-memory transfer time. Please refer to the graph above to have a better understanding of the performance gain for different dataset sizes.
Problems also seem to arise when the input dataset contains values with extreme differences (e.g. 107) if no scaling is performed. Such an example is the "breast-cancer" dataset provided in the official LIBSVM page.
### ACKNOWLEDGEMENTS
This work was supported by the EU FP7 projects GLOCAL (FP7-248984) and WeKnowIt (FP7-215453)
================================================
FILE: binaries/linux/COPYRIGHT
================================================
Copyright (c) 2000-2013 Chih-Chung Chang and Chih-Jen Lin
All rights reserved.
Redistribution and use in source and binary forms, with or without
modification, are permitted provided that the following conditions
are met:
1. Redistributions of source code must retain the above copyright
notice, this list of conditions and the following disclaimer.
2. Redistributions in binary form must reproduce the above copyright
notice, this list of conditions and the following disclaimer in the
documentation and/or other materials provided with the distribution.
3. Neither name of copyright holders nor the names of its contributors
may be used to endorse or promote products derived from this software
without specific prior written permission.
THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS
``AS IS'' AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT
LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR
A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE REGENTS OR
CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL,
EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO,
PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR
PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF
LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING
NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS
SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
================================================
FILE: binaries/linux/FAQ.html
================================================
<html>
<head>
<title>LIBSVM FAQ</title>
</head>
<body bgcolor="#ffffcc">
<a name="_TOP"><b><h1><a
href=http://www.csie.ntu.edu.tw/~cjlin/libsvm>LIBSVM</a> FAQ </h1></b></a>
<b>last modified : </b>
Wed, 19 Dec 2012 13:26:34 GMT
<class="categories">
<li><a
href="#_TOP">All Questions</a>(78)</li>
<ul><b>
<li><a
href="#/Q1:_Some_sample_uses_of_libsvm">Q1:_Some_sample_uses_of_libsvm</a>(2)</li>
<li><a
href="#/Q2:_Installation_and_running_the_program">Q2:_Installation_and_running_the_program</a>(13)</li>
<li><a
href="#/Q3:_Data_preparation">Q3:_Data_preparation</a>(7)</li>
<li><a
href="#/Q4:_Training_and_prediction">Q4:_Training_and_prediction</a>(34)</li>
<li><a
href="#/Q5:_Probability_outputs">Q5:_Probability_outputs</a>(3)</li>
<li><a
href="#/Q6:_Graphic_interface">Q6:_Graphic_interface</a>(3)</li>
<li><a
href="#/Q7:_Java_version_of_libsvm">Q7:_Java_version_of_libsvm</a>(4)</li>
<li><a
href="#/Q8:_Python_interface">Q8:_Python_interface</a>(1)</li>
<li><a
href="#/Q9:_MATLAB_interface">Q9:_MATLAB_interface</a>(11)</li>
</b></ul>
</li>
<ul><ul class="headlines">
<li class="headlines_item"><a href="#faq101">Some courses which have used libsvm as a tool</a></li>
<li class="headlines_item"><a href="#faq102">Some applications/tools which have used libsvm </a></li>
<li class="headlines_item"><a href="#f201">Where can I find documents/videos of libsvm ?</a></li>
<li class="headlines_item"><a href="#f202">Where are change log and earlier versions?</a></li>
<li class="headlines_item"><a href="#f203">How to cite LIBSVM?</a></li>
<li class="headlines_item"><a href="#f204">I would like to use libsvm in my software. Is there any license problem?</a></li>
<li class="headlines_item"><a href="#f205">Is there a repository of additional tools based on libsvm?</a></li>
<li class="headlines_item"><a href="#f206">On unix machines, I got "error in loading shared libraries" or "cannot open shared object file." What happened ? </a></li>
<li class="headlines_item"><a href="#f207">I have modified the source and would like to build the graphic interface "svm-toy" on MS windows. How should I do it ?</a></li>
<li class="headlines_item"><a href="#f208">I am an MS windows user but why only one (svm-toy) of those precompiled .exe actually runs ? </a></li>
<li class="headlines_item"><a href="#f209">What is the difference between "." and "*" outputed during training? </a></li>
<li class="headlines_item"><a href="#f210">Why occasionally the program (including MATLAB or other interfaces) crashes and gives a segmentation fault?</a></li>
<li class="headlines_item"><a href="#f211">How to build a dynamic library (.dll file) on MS windows?</a></li>
<li class="headlines_item"><a href="#f212">On some systems (e.g., Ubuntu), compiling LIBSVM gives many warning messages. Is this a problem and how to disable the warning message?</a></li>
<li class="headlines_item"><a href="#f213">In LIBSVM, why you don't use certain C/C++ library functions to make the code shorter?</a></li>
<li class="headlines_item"><a href="#f301">Why sometimes not all attributes of a data appear in the training/model files ?</a></li>
<li class="headlines_item"><a href="#f302">What if my data are non-numerical ?</a></li>
<li class="headlines_item"><a href="#f303">Why do you consider sparse format ? Will the training of dense data be much slower ?</a></li>
<li class="headlines_item"><a href="#f304">Why sometimes the last line of my data is not read by svm-train?</a></li>
<li class="headlines_item"><a href="#f305">Is there a program to check if my data are in the correct format?</a></li>
<li class="headlines_item"><a href="#f306">May I put comments in data files?</a></li>
<li class="headlines_item"><a href="#f307">How to convert other data formats to LIBSVM format?</a></li>
<li class="headlines_item"><a href="#f401">The output of training C-SVM is like the following. What do they mean?</a></li>
<li class="headlines_item"><a href="#f402">Can you explain more about the model file?</a></li>
<li class="headlines_item"><a href="#f403">Should I use float or double to store numbers in the cache ?</a></li>
<li class="headlines_item"><a href="#f404">How do I choose the kernel?</a></li>
<li class="headlines_item"><a href="#f405">Does libsvm have special treatments for linear SVM?</a></li>
<li class="headlines_item"><a href="#f406">The number of free support vectors is large. What should I do?</a></li>
<li class="headlines_item"><a href="#f407">Should I scale training and testing data in a similar way?</a></li>
<li class="headlines_item"><a href="#f408">Does it make a big difference if I scale each attribute to [0,1] instead of [-1,1]?</a></li>
<li class="headlines_item"><a href="#f409">The prediction rate is low. How could I improve it?</a></li>
<li class="headlines_item"><a href="#f410">My data are unbalanced. Could libsvm handle such problems?</a></li>
<li class="headlines_item"><a href="#f411">What is the difference between nu-SVC and C-SVC?</a></li>
<li class="headlines_item"><a href="#f412">The program keeps running (without showing any output). What should I do?</a></li>
<li class="headlines_item"><a href="#f413">The program keeps running (with output, i.e. many dots). What should I do?</a></li>
<li class="headlines_item"><a href="#f414">The training time is too long. What should I do?</a></li>
<li class="headlines_item"><a href="#f4141">Does shrinking always help?</a></li>
<li class="headlines_item"><a href="#f415">How do I get the decision value(s)?</a></li>
<li class="headlines_item"><a href="#f4151">How do I get the distance between a point and the hyperplane?</a></li>
<li class="headlines_item"><a href="#f416">On 32-bit machines, if I use a large cache (i.e. large -m) on a linux machine, why sometimes I get "segmentation fault ?"</a></li>
<li class="headlines_item"><a href="#f417">How do I disable screen output of svm-train?</a></li>
<li class="headlines_item"><a href="#f418">I would like to use my own kernel. Any example? In svm.cpp, there are two subroutines for kernel evaluations: k_function() and kernel_function(). Which one should I modify ?</a></li>
<li class="headlines_item"><a href="#f419">What method does libsvm use for multi-class SVM ? Why don't you use the "1-against-the rest" method?</a></li>
<li class="headlines_item"><a href="#f4191">How does LIBSVM perform parameter selection for multi-class problems? </a></li>
<li class="headlines_item"><a href="#f420">After doing cross validation, why there is no model file outputted ?</a></li>
<li class="headlines_item"><a href="#f4201">Why my cross-validation results are different from those in the Practical Guide?</a></li>
<li class="headlines_item"><a href="#f421">On some systems CV accuracy is the same in several runs. How could I use different data partitions? In other words, how do I set random seed in LIBSVM?</a></li>
<li class="headlines_item"><a href="#f422">I would like to solve L2-loss SVM (i.e., error term is quadratic). How should I modify the code ?</a></li>
<li class="headlines_item"><a href="#f424">How do I choose parameters for one-class svm as training data are in only one class?</a></li>
<li class="headlines_item"><a href="#f427">Why the code gives NaN (not a number) results?</a></li>
<li class="headlines_item"><a href="#f428">Why on windows sometimes grid.py fails?</a></li>
<li class="headlines_item"><a href="#f429">Why grid.py/easy.py sometimes generates the following warning message?</a></li>
<li class="headlines_item"><a href="#f430">Why the sign of predicted labels and decision values are sometimes reversed?</a></li>
<li class="headlines_item"><a href="#f431">I don't know class labels of test data. What should I put in the first column of the test file?</a></li>
<li class="headlines_item"><a href="#f432">How can I use OpenMP to parallelize LIBSVM on a multicore/shared-memory computer?</a></li>
<li class="headlines_item"><a href="#f433">How could I know which training instances are support vectors?</a></li>
<li class="headlines_item"><a href="#f425">Why training a probability model (i.e., -b 1) takes a longer time?</a></li>
<li class="headlines_item"><a href="#f426">Why using the -b option does not give me better accuracy?</a></li>
<li class="headlines_item"><a href="#f427">Why using svm-predict -b 0 and -b 1 gives different accuracy values?</a></li>
<li class="headlines_item"><a href="#f501">How can I save images drawn by svm-toy?</a></li>
<li class="headlines_item"><a href="#f502">I press the "load" button to load data points but why svm-toy does not draw them ?</a></li>
<li class="headlines_item"><a href="#f503">I would like svm-toy to handle more than three classes of data, what should I do ?</a></li>
<li class="headlines_item"><a href="#f601">What is the difference between Java version and C++ version of libsvm?</a></li>
<li class="headlines_item"><a href="#f602">Is the Java version significantly slower than the C++ version?</a></li>
<li class="headlines_item"><a href="#f603">While training I get the following error message: java.lang.OutOfMemoryError. What is wrong?</a></li>
<li class="headlines_item"><a href="#f604">Why you have the main source file svm.m4 and then transform it to svm.java?</a></li>
<li class="headlines_item"><a href="#f704">Except the python-C++ interface provided, could I use Jython to call libsvm ?</a></li>
<li class="headlines_item"><a href="#f801">I compile the MATLAB interface without problem, but why errors occur while running it?</a></li>
<li class="headlines_item"><a href="#f8011">On 64bit Windows I compile the MATLAB interface without problem, but why errors occur while running it?</a></li>
<li class="headlines_item"><a href="#f802">Does the MATLAB interface provide a function to do scaling?</a></li>
<li class="headlines_item"><a href="#f803">How could I use MATLAB interface for parameter selection?</a></li>
<li class="headlines_item"><a href="#f8031">I use MATLAB parallel programming toolbox on a multi-core environment for parameter selection. Why the program is even slower?</a></li>
<li class="headlines_item"><a href="#f8032">How do I use LIBSVM with OpenMP under MATLAB?</a></li>
<li class="headlines_item"><a href="#f804">How could I generate the primal variable w of linear SVM?</a></li>
<li class="headlines_item"><a href="#f805">Is there an OCTAVE interface for libsvm?</a></li>
<li class="headlines_item"><a href="#f806">How to handle the name conflict between svmtrain in the libsvm matlab interface and that in MATLAB bioinformatics toolbox?</a></li>
<li class="headlines_item"><a href="#f807">On Windows I got an error message "Invalid MEX-file: Specific module not found" when running the pre-built MATLAB interface in the windows sub-directory. What should I do?</a></li>
<li class="headlines_item"><a href="#f808">LIBSVM supports 1-vs-1 multi-class classification. If instead I would like to use 1-vs-rest, how to implement it using MATLAB interface?</a></li>
</ul></ul>
<hr size="5" noshade />
<p/>
<a name="/Q1:_Some_sample_uses_of_libsvm"></a>
<a name="faq101"><b>Q: Some courses which have used libsvm as a tool</b></a>
<br/>
<ul>
<li><a href=http://lmb.informatik.uni-freiburg.de/lectures/svm_seminar/>Institute for Computer Science,
Faculty of Applied Science, University of Freiburg, Germany
</a>
<li> <a href=http://www.cs.vu.nl/~elena/ml.html>
Division of Mathematics and Computer Science.
Faculteit der Exacte Wetenschappen
Vrije Universiteit, The Netherlands. </a>
<li>
<a href=http://www.cae.wisc.edu/~ece539/matlab/>
Electrical and Computer Engineering Department,
University of Wisconsin-Madison
</a>
<li>
<a href=http://www.hpl.hp.com/personal/Carl_Staelin/cs236601/project.html>
Technion (Israel Institute of Technology), Israel.
<li>
<a href=http://www.cise.ufl.edu/~fu/learn.html>
Computer and Information Sciences Dept., University of Florida</a>
<li>
<a href=http://www.uonbi.ac.ke/acad_depts/ics/course_material/machine_learning/ML_and_DM_Resources.html>
The Institute of Computer Science,
University of Nairobi, Kenya.</a>
<li>
<a href=http://cerium.raunvis.hi.is/~tpr/courseware/svm/hugbunadur.html>
Applied Mathematics and Computer Science, University of Iceland.
<li>
<a href=http://chicago05.mlss.cc/tiki/tiki-read_article.php?articleId=2>
SVM tutorial in machine learning
summer school, University of Chicago, 2005.
</a>
</ul>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q1:_Some_sample_uses_of_libsvm"></a>
<a name="faq102"><b>Q: Some applications/tools which have used libsvm </b></a>
<br/>
(and maybe liblinear).
<ul>
<li>
<a href=http://people.csail.mit.edu/jjl/libpmk/>LIBPMK: A Pyramid Match Toolkit</a>
</li>
<li><a href=http://maltparser.org/>Maltparser</a>:
a system for data-driven dependency parsing
</li>
<li>
<a href=http://www.pymvpa.org/>PyMVPA: python tool for classifying neuroimages</a>
</li>
<li>
<a href=http://solpro.proteomics.ics.uci.edu/>
SOLpro: protein solubility predictor
</a>
</li>
<li>
<a href=http://bdval.campagnelab.org>
BDVal</a>: biomarker discovery in high-throughput datasets.
</li>
<li><a href=http://johel.m.free.fr/demo_045.htm>
Realtime object recognition</a>
</li>
<li><a href=http://scikit-learn.sourceforge.net/>
scikits.learn: machine learning in Python</a>
</li>
</ul>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f201"><b>Q: Where can I find documents/videos of libsvm ?</b></a>
<br/>
<p>
<ul>
<li>
Official implementation document:
<br>
C.-C. Chang and
C.-J. Lin.
LIBSVM
: a library for support vector machines.
ACM Transactions on Intelligent
Systems and Technology, 2:27:1--27:27, 2011.
<a href="http://www.csie.ntu.edu.tw/~cjlin/papers/libsvm.pdf">pdf</a>, <a href=http://www.csie.ntu.edu.tw/~cjlin/papers/libsvm.ps.gz>ps.gz</a>,
<a href=http://portal.acm.org/citation.cfm?id=1961199&CFID=29950432&CFTOKEN=30974232>ACM digital lib</a>.
<li> Instructions for using LIBSVM are in the README files in the main directory and some sub-directories.
<br>
README in the main directory: details all options, data format, and library calls.
<br>
tools/README: parameter selection and other tools
<li>
A guide for beginners:
<br>
C.-W. Hsu, C.-C. Chang, and
C.-J. Lin.
<A HREF="http://www.csie.ntu.edu.tw/~cjlin/papers/guide/guide.pdf">
A practical guide to support vector classification
</A>
<li> An <a href=http://www.youtube.com/watch?v=gePWtNAQcK8>introductory video</a>
for windows users.
</ul>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f202"><b>Q: Where are change log and earlier versions?</b></a>
<br/>
<p>See <a href="http://www.csie.ntu.edu.tw/~cjlin/libsvm/log">the change log</a>.
<p> You can download earlier versions
<a href="http://www.csie.ntu.edu.tw/~cjlin/libsvm/oldfiles">here</a>.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f203"><b>Q: How to cite LIBSVM?</b></a>
<br/>
<p>
Please cite the following paper:
<p>
Chih-Chung Chang and Chih-Jen Lin, LIBSVM
: a library for support vector machines.
ACM Transactions on Intelligent Systems and Technology, 2:27:1--27:27, 2011.
Software available at http://www.csie.ntu.edu.tw/~cjlin/libsvm
<p>
The bibtex format is
<pre>
@article{CC01a,
author = {Chang, Chih-Chung and Lin, Chih-Jen},
title = {{LIBSVM}: A library for support vector machines},
journal = {ACM Transactions on Intelligent Systems and Technology},
volume = {2},
issue = {3},
year = {2011},
pages = {27:1--27:27},
note = {Software available at \url{http://www.csie.ntu.edu.tw/~cjlin/libsvm}}
}
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f204"><b>Q: I would like to use libsvm in my software. Is there any license problem?</b></a>
<br/>
<p>
The libsvm license ("the modified BSD license")
is compatible with many
free software licenses such as GPL. Hence, it is very easy to
use libsvm in your software.
Please check the COPYRIGHT file in detail. Basically
you need to
<ol>
<li>
Clearly indicate that LIBSVM is used.
</li>
<li>
Retain the LIBSVM COPYRIGHT file in your software.
</li>
</ol>
It can also be used in commercial products.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f205"><b>Q: Is there a repository of additional tools based on libsvm?</b></a>
<br/>
<p>
Yes, see <a href="http://www.csie.ntu.edu.tw/~cjlin/libsvmtools">libsvm
tools</a>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f206"><b>Q: On unix machines, I got "error in loading shared libraries" or "cannot open shared object file." What happened ? </b></a>
<br/>
<p>
This usually happens if you compile the code
on one machine and run it on another which has incompatible
libraries.
Try to recompile the program on that machine or use static linking.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f207"><b>Q: I have modified the source and would like to build the graphic interface "svm-toy" on MS windows. How should I do it ?</b></a>
<br/>
<p>
Build it as a project by choosing "Win32 Project."
On the other hand, for "svm-train" and "svm-predict"
you want to choose "Win32 Console Project."
After libsvm 2.5, you can also use the file Makefile.win.
See details in README.
<p>
If you are not using Makefile.win and see the following
link error
<pre>
LIBCMTD.lib(wwincrt0.obj) : error LNK2001: unresolved external symbol
_wWinMain@16
</pre>
you may have selected a wrong project type.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f208"><b>Q: I am an MS windows user but why only one (svm-toy) of those precompiled .exe actually runs ? </b></a>
<br/>
<p>
You need to open a command window
and type svmtrain.exe to see all options.
Some examples are in README file.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f209"><b>Q: What is the difference between "." and "*" outputed during training? </b></a>
<br/>
<p>
"." means every 1,000 iterations (or every #data
iterations is your #data is less than 1,000).
"*" means that after iterations of using
a smaller shrunk problem,
we reset to use the whole set. See the
<a href=../papers/libsvm.pdf>implementation document</a> for details.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f210"><b>Q: Why occasionally the program (including MATLAB or other interfaces) crashes and gives a segmentation fault?</b></a>
<br/>
<p>
Very likely the program consumes too much memory than what the
operating system can provide. Try a smaller data and see if the
program still crashes.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f211"><b>Q: How to build a dynamic library (.dll file) on MS windows?</b></a>
<br/>
<p>
The easiest way is to use Makefile.win.
See details in README.
Alternatively, you can use Visual C++. Here is
the example using Visual Studio .Net 2008:
<ol>
<li>Create a Win32 empty DLL project and set (in Project->$Project_Name
Properties...->Configuration) to "Release."
About how to create a new dynamic link library, please refer to
<a href=http://msdn2.microsoft.com/en-us/library/ms235636(VS.80).aspx>http://msdn2.microsoft.com/en-us/library/ms235636(VS.80).aspx</a>
<li> Add svm.cpp, svm.h to your project.
<li> Add __WIN32__ and _CRT_SECURE_NO_DEPRECATE to Preprocessor definitions (in
Project->$Project_Name Properties...->C/C++->Preprocessor)
<li> Set Create/Use Precompiled Header to Not Using Precompiled Headers
(in Project->$Project_Name Properties...->C/C++->Precompiled Headers)
<li> Set the path for the Modulation Definition File svm.def (in
Project->$Project_Name Properties...->Linker->input
<li> Build the DLL.
<li> Rename the dll file to libsvm.dll and move it to the correct path.
</ol>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f212"><b>Q: On some systems (e.g., Ubuntu), compiling LIBSVM gives many warning messages. Is this a problem and how to disable the warning message?</b></a>
<br/>
<p>
The warning message is like
<pre>
svm.cpp:2730: warning: ignoring return value of int fscanf(FILE*, const char*, ...), declared with attribute warn_unused_result
</pre>
This is not a problem; see <a href=https://wiki.ubuntu.com/CompilerFlags#-D_FORTIFY_SOURCE=2>this page</a> for more
details of ubuntu systems.
In the future we may modify the code
so that these messages do not appear.
At this moment, to disable the warning message you can replace
<pre>
CFLAGS = -Wall -Wconversion -O3 -fPIC
</pre>
with
<pre>
CFLAGS = -Wall -Wconversion -O3 -fPIC -U_FORTIFY_SOURCE
</pre>
in Makefile.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f213"><b>Q: In LIBSVM, why you don't use certain C/C++ library functions to make the code shorter?</b></a>
<br/>
<p>
For portability, we use only features defined in ISO C89. Note that features in ISO C99 may not be available everywhere.
Even the newest gcc lacks some features in C99 (see <a href=http://gcc.gnu.org/c99status.html>http://gcc.gnu.org/c99status.html</a> for details).
If the situation changes in the future,
we might consider using these newer features.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f301"><b>Q: Why sometimes not all attributes of a data appear in the training/model files ?</b></a>
<br/>
<p>
libsvm uses the so called "sparse" format where zero
values do not need to be stored. Hence a data with attributes
<pre>
1 0 2 0
</pre>
is represented as
<pre>
1:1 3:2
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f302"><b>Q: What if my data are non-numerical ?</b></a>
<br/>
<p>
Currently libsvm supports only numerical data.
You may have to change non-numerical data to
numerical. For example, you can use several
binary attributes to represent a categorical
attribute.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f303"><b>Q: Why do you consider sparse format ? Will the training of dense data be much slower ?</b></a>
<br/>
<p>
This is a controversial issue. The kernel
evaluation (i.e. inner product) of sparse vectors is slower
so the total training time can be at least twice or three times
of that using the dense format.
However, we cannot support only dense format as then we CANNOT
handle extremely sparse cases. Simplicity of the code is another
concern. Right now we decide to support
the sparse format only.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f304"><b>Q: Why sometimes the last line of my data is not read by svm-train?</b></a>
<br/>
<p>
We assume that you have '\n' in the end of
each line. So please press enter in the end
of your last line.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f305"><b>Q: Is there a program to check if my data are in the correct format?</b></a>
<br/>
<p>
The svm-train program in libsvm conducts only a simple check of the input data. To do a
detailed check, after libsvm 2.85, you can use the python script tools/checkdata.py. See tools/README for details.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f306"><b>Q: May I put comments in data files?</b></a>
<br/>
<p>
We don't officially support this. But, cureently LIBSVM
is able to process data in the following
format:
<pre>
1 1:2 2:1 # your comments
</pre>
Note that the character ":" should not appear in your
comments.
<!--
No, for simplicity we don't support that.
However, you can easily preprocess your data before
using libsvm. For example,
if you have the following data
<pre>
test.txt
1 1:2 2:1 # proten A
</pre>
then on unix machines you can do
<pre>
cut -d '#' -f 1 < test.txt > test.features
cut -d '#' -f 2 < test.txt > test.comments
svm-predict test.feature train.model test.predicts
paste -d '#' test.predicts test.comments | sed 's/#/ #/' > test.results
</pre>
-->
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f307"><b>Q: How to convert other data formats to LIBSVM format?</b></a>
<br/>
<p>
It depends on your data format. A simple way is to use
libsvmwrite in the libsvm matlab/octave interface.
Take a CSV (comma-separated values) file
in UCI machine learning repository as an example.
We download <a href=http://archive.ics.uci.edu/ml/machine-learning-databases/spect/SPECTF.train>SPECTF.train</a>.
Labels are in the first column. The following steps produce
a file in the libsvm format.
<pre>
matlab> SPECTF = csvread('SPECTF.train'); % read a csv file
matlab> labels = SPECTF(:, 1); % labels from the 1st column
matlab> features = SPECTF(:, 2:end);
matlab> features_sparse = sparse(features); % features must be in a sparse matrix
matlab> libsvmwrite('SPECTFlibsvm.train', labels, features_sparse);
</pre>
The tranformed data are stored in SPECTFlibsvm.train.
<p>
Alternatively, you can use <a href="./faqfiles/convert.c">convert.c</a>
to convert CSV format to libsvm format.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f401"><b>Q: The output of training C-SVM is like the following. What do they mean?</b></a>
<br/>
<br>optimization finished, #iter = 219
<br>nu = 0.431030
<br>obj = -100.877286, rho = 0.424632
<br>nSV = 132, nBSV = 107
<br>Total nSV = 132
<p>
obj is the optimal objective value of the dual SVM problem.
rho is the bias term in the decision function
sgn(w^Tx - rho).
nSV and nBSV are number of support vectors and bounded support
vectors (i.e., alpha_i = C). nu-svm is a somewhat equivalent
form of C-SVM where C is replaced by nu. nu simply shows the
corresponding parameter. More details are in
<a href="http://www.csie.ntu.edu.tw/~cjlin/papers/libsvm.pdf">
libsvm document</a>.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f402"><b>Q: Can you explain more about the model file?</b></a>
<br/>
<p>
In the model file, after parameters and other informations such as labels , each line represents a support vector.
Support vectors are listed in the order of "labels" shown earlier.
(i.e., those from the first class in the "labels" list are
grouped first, and so on.)
If k is the total number of classes,
in front of a support vector in class j, there are
k-1 coefficients
y*alpha where alpha are dual solution of the
following two class problems:
<br>
1 vs j, 2 vs j, ..., j-1 vs j, j vs j+1, j vs j+2, ..., j vs k
<br>
and y=1 in first j-1 coefficients, y=-1 in the remaining
k-j coefficients.
For example, if there are 4 classes, the file looks like:
<pre>
+-+-+-+--------------------+
|1|1|1| |
|v|v|v| SVs from class 1 |
|2|3|4| |
+-+-+-+--------------------+
|1|2|2| |
|v|v|v| SVs from class 2 |
|2|3|4| |
+-+-+-+--------------------+
|1|2|3| |
|v|v|v| SVs from class 3 |
|3|3|4| |
+-+-+-+--------------------+
|1|2|3| |
|v|v|v| SVs from class 4 |
|4|4|4| |
+-+-+-+--------------------+
</pre>
See also
<a href="#f804"> an illustration using
MATLAB/OCTAVE.</a>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f403"><b>Q: Should I use float or double to store numbers in the cache ?</b></a>
<br/>
<p>
We have float as the default as you can store more numbers
in the cache.
In general this is good enough but for few difficult
cases (e.g. C very very large) where solutions are huge
numbers, it might be possible that the numerical precision is not
enough using only float.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f404"><b>Q: How do I choose the kernel?</b></a>
<br/>
<p>
In general we suggest you to try the RBF kernel first.
A recent result by Keerthi and Lin
(<a href=http://www.csie.ntu.edu.tw/~cjlin/papers/limit.pdf>
download paper here</a>)
shows that if RBF is used with model selection,
then there is no need to consider the linear kernel.
The kernel matrix using sigmoid may not be positive definite
and in general it's accuracy is not better than RBF.
(see the paper by Lin and Lin
(<a href=http://www.csie.ntu.edu.tw/~cjlin/papers/tanh.pdf>
download paper here</a>).
Polynomial kernels are ok but if a high degree is used,
numerical difficulties tend to happen
(thinking about dth power of (<1) goes to 0
and (>1) goes to infinity).
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f405"><b>Q: Does libsvm have special treatments for linear SVM?</b></a>
<br/>
<p>
No, libsvm solves linear/nonlinear SVMs by the
same way.
Some tricks may save training/testing time if the
linear kernel is used,
so libsvm is <b>NOT</b> particularly efficient for linear SVM,
especially when
C is large and
the number of data is much larger
than the number of attributes.
You can either
<ul>
<li>
Use small C only. We have shown in the following paper
that after C is larger than a certain threshold,
the decision function is the same.
<p>
<a href="http://guppy.mpe.nus.edu.sg/~mpessk/">S. S. Keerthi</a>
and
<B>C.-J. Lin</B>.
<A HREF="papers/limit.pdf">
Asymptotic behaviors of support vector machines with
Gaussian kernel
</A>
.
<I><A HREF="http://mitpress.mit.edu/journal-home.tcl?issn=08997667">Neural Computation</A></I>, 15(2003), 1667-1689.
<li>
Check <a href=http://www.csie.ntu.edu.tw/~cjlin/liblinear>liblinear</a>,
which is designed for large-scale linear classification.
</ul>
<p> Please also see our <a href=../papers/guide/guide.pdf>SVM guide</a>
on the discussion of using RBF and linear
kernels.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f406"><b>Q: The number of free support vectors is large. What should I do?</b></a>
<br/>
<p>
This usually happens when the data are overfitted.
If attributes of your data are in large ranges,
try to scale them. Then the region
of appropriate parameters may be larger.
Note that there is a scale program
in libsvm.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f407"><b>Q: Should I scale training and testing data in a similar way?</b></a>
<br/>
<p>
Yes, you can do the following:
<pre>
> svm-scale -s scaling_parameters train_data > scaled_train_data
> svm-scale -r scaling_parameters test_data > scaled_test_data
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f408"><b>Q: Does it make a big difference if I scale each attribute to [0,1] instead of [-1,1]?</b></a>
<br/>
<p>
For the linear scaling method, if the RBF kernel is
used and parameter selection is conducted, there
is no difference. Assume Mi and mi are
respectively the maximal and minimal values of the
ith attribute. Scaling to [0,1] means
<pre>
x'=(x-mi)/(Mi-mi)
</pre>
For [-1,1],
<pre>
x''=2(x-mi)/(Mi-mi)-1.
</pre>
In the RBF kernel,
<pre>
x'-y'=(x-y)/(Mi-mi), x''-y''=2(x-y)/(Mi-mi).
</pre>
Hence, using (C,g) on the [0,1]-scaled data is the
same as (C,g/2) on the [-1,1]-scaled data.
<p> Though the performance is the same, the computational
time may be different. For data with many zero entries,
[0,1]-scaling keeps the sparsity of input data and hence
may save the time.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f409"><b>Q: The prediction rate is low. How could I improve it?</b></a>
<br/>
<p>
Try to use the model selection tool grid.py in the python
directory find
out good parameters. To see the importance of model selection,
please
see my talk:
<A HREF="http://www.csie.ntu.edu.tw/~cjlin/talks/freiburg.pdf">
A practical guide to support vector
classification
</A>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f410"><b>Q: My data are unbalanced. Could libsvm handle such problems?</b></a>
<br/>
<p>
Yes, there is a -wi options. For example, if you use
<pre>
> svm-train -s 0 -c 10 -w1 1 -w-1 5 data_file
</pre>
<p>
the penalty for class "-1" is larger.
Note that this -w option is for C-SVC only.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f411"><b>Q: What is the difference between nu-SVC and C-SVC?</b></a>
<br/>
<p>
Basically they are the same thing but with different
parameters. The range of C is from zero to infinity
but nu is always between [0,1]. A nice property
of nu is that it is related to the ratio of
support vectors and the ratio of the training
error.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f412"><b>Q: The program keeps running (without showing any output). What should I do?</b></a>
<br/>
<p>
You may want to check your data. Each training/testing
data must be in one line. It cannot be separated.
In addition, you have to remove empty lines.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f413"><b>Q: The program keeps running (with output, i.e. many dots). What should I do?</b></a>
<br/>
<p>
In theory libsvm guarantees to converge.
Therefore, this means you are
handling ill-conditioned situations
(e.g. too large/small parameters) so numerical
difficulties occur.
<p>
You may get better numerical stability by replacing
<pre>
typedef float Qfloat;
</pre>
in svm.cpp with
<pre>
typedef double Qfloat;
</pre>
That is, elements in the kernel cache are stored
in double instead of single. However, this means fewer elements
can be put in the kernel cache.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f414"><b>Q: The training time is too long. What should I do?</b></a>
<br/>
<p>
For large problems, please specify enough cache size (i.e.,
-m).
Slow convergence may happen for some difficult cases (e.g. -c is large).
You can try to use a looser stopping tolerance with -e.
If that still doesn't work, you may train only a subset of the data.
You can use the program subset.py in the directory "tools"
to obtain a random subset.
<p>
If you have extremely large data and face this difficulty, please
contact us. We will be happy to discuss possible solutions.
<p> When using large -e, you may want to check if -h 0 (no shrinking) or -h 1 (shrinking) is faster.
See a related question below.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f4141"><b>Q: Does shrinking always help?</b></a>
<br/>
<p>
If the number of iterations is high, then shrinking
often helps.
However, if the number of iterations is small
(e.g., you specify a large -e), then
probably using -h 0 (no shrinking) is better.
See the
<a href=../papers/libsvm.pdf>implementation document</a> for details.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f415"><b>Q: How do I get the decision value(s)?</b></a>
<br/>
<p>
We print out decision values for regression. For classification,
we solve several binary SVMs for multi-class cases. You
can obtain values by easily calling the subroutine
svm_predict_values. Their corresponding labels
can be obtained from svm_get_labels.
Details are in
README of libsvm package.
<p>
If you are using MATLAB/OCTAVE interface, svmpredict can directly
give you decision values. Please see matlab/README for details.
<p>
We do not recommend the following. But if you would
like to get values for
TWO-class classification with labels +1 and -1
(note: +1 and -1 but not things like 5 and 10)
in the easiest way, simply add
<pre>
printf("%f\n", dec_values[0]*model->label[0]);
</pre>
after the line
<pre>
svm_predict_values(model, x, dec_values);
</pre>
of the file svm.cpp.
Positive (negative)
decision values correspond to data predicted as +1 (-1).
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f4151"><b>Q: How do I get the distance between a point and the hyperplane?</b></a>
<br/>
<p>
The distance is |decision_value| / |w|.
We have |w|^2 = w^Tw = alpha^T Q alpha = 2*(dual_obj + sum alpha_i).
Thus in svm.cpp please find the place
where we calculate the dual objective value
(i.e., the subroutine Solve())
and add a statement to print w^Tw.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f416"><b>Q: On 32-bit machines, if I use a large cache (i.e. large -m) on a linux machine, why sometimes I get "segmentation fault ?"</b></a>
<br/>
<p>
On 32-bit machines, the maximum addressable
memory is 4GB. The Linux kernel uses 3:1
split which means user space is 3G and
kernel space is 1G. Although there are
3G user space, the maximum dynamic allocation
memory is 2G. So, if you specify -m near 2G,
the memory will be exhausted. And svm-train
will fail when it asks more memory.
For more details, please read
<a href=http://groups.google.com/groups?hl=en&lr=&ie=UTF-8&selm=3BA164F6.BAFA4FB%40daimi.au.dk>
this article</a>.
<p>
The easiest solution is to switch to a
64-bit machine.
Otherwise, there are two ways to solve this. If your
machine supports Intel's PAE (Physical Address
Extension), you can turn on the option HIGHMEM64G
in Linux kernel which uses 4G:4G split for
kernel and user space. If you don't, you can
try a software `tub' which can eliminate the 2G
boundary for dynamic allocated memory. The `tub'
is available at
<a href=http://www.bitwagon.com/tub.html>http://www.bitwagon.com/tub.html</a>.
<!--
This may happen only when the cache is large, but each cached row is
not large enough. <b>Note:</b> This problem is specific to
gnu C library which is used in linux.
The solution is as follows:
<p>
In our program we have malloc() which uses two methods
to allocate memory from kernel. One is
sbrk() and another is mmap(). sbrk is faster, but mmap
has a larger address
space. So malloc uses mmap only if the wanted memory size is larger
than some threshold (default 128k).
In the case where each row is not large enough (#elements < 128k/sizeof(float)) but we need a large cache ,
the address space for sbrk can be exhausted. The solution is to
lower the threshold to force malloc to use mmap
and increase the maximum number of chunks to allocate
with mmap.
<p>
Therefore, in the main program (i.e. svm-train.c) you want
to have
<pre>
#include <malloc.h>
</pre>
and then in main():
<pre>
mallopt(M_MMAP_THRESHOLD, 32768);
mallopt(M_MMAP_MAX,1000000);
</pre>
You can also set the environment variables instead
of writing them in the program:
<pre>
$ M_MMAP_MAX=1000000 M_MMAP_THRESHOLD=32768 ./svm-train .....
</pre>
More information can be found by
<pre>
$ info libc "Malloc Tunable Parameters"
</pre>
-->
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f417"><b>Q: How do I disable screen output of svm-train?</b></a>
<br/>
<p>
For commend-line users, use the -q option:
<pre>
> ./svm-train -q heart_scale
</pre>
<p>
For library users, set the global variable
<pre>
extern void (*svm_print_string) (const char *);
</pre>
to specify the output format. You can disable the output by the following steps:
<ol>
<li>
Declare a function to output nothing:
<pre>
void print_null(const char *s) {}
</pre>
</li>
<li>
Assign the output function of libsvm by
<pre>
svm_print_string = &print_null;
</pre>
</li>
</ol>
Finally, a way used in earlier libsvm
is by updating svm.cpp from
<pre>
#if 1
void info(const char *fmt,...)
</pre>
to
<pre>
#if 0
void info(const char *fmt,...)
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f418"><b>Q: I would like to use my own kernel. Any example? In svm.cpp, there are two subroutines for kernel evaluations: k_function() and kernel_function(). Which one should I modify ?</b></a>
<br/>
<p>
An example is "LIBSVM for string data" in LIBSVM Tools.
<p>
The reason why we have two functions is as follows.
For the RBF kernel exp(-g |xi - xj|^2), if we calculate
xi - xj first and then the norm square, there are 3n operations.
Thus we consider exp(-g (|xi|^2 - 2dot(xi,xj) +|xj|^2))
and by calculating all |xi|^2 in the beginning,
the number of operations is reduced to 2n.
This is for the training. For prediction we cannot
do this so a regular subroutine using that 3n operations is
needed.
The easiest way to have your own kernel is
to put the same code in these two
subroutines by replacing any kernel.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f419"><b>Q: What method does libsvm use for multi-class SVM ? Why don't you use the "1-against-the rest" method?</b></a>
<br/>
<p>
It is one-against-one. We chose it after doing the following
comparison:
C.-W. Hsu and C.-J. Lin.
<A HREF="http://www.csie.ntu.edu.tw/~cjlin/papers/multisvm.pdf">
A comparison of methods
for multi-class support vector machines
</A>,
<I>IEEE Transactions on Neural Networks</A></I>, 13(2002), 415-425.
<p>
"1-against-the rest" is a good method whose performance
is comparable to "1-against-1." We do the latter
simply because its training time is shorter.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f4191"><b>Q: How does LIBSVM perform parameter selection for multi-class problems? </b></a>
<br/>
<p>
LIBSVM implements "one-against-one" multi-class method, so there are
k(k-1)/2 binary models, where k is the number of classes.
<p>
We can consider two ways to conduct parameter selection.
<ol>
<li>
For any two classes of data, a parameter selection procedure is conducted. Finally,
each decision function has its own optimal parameters.
</li>
<li>
The same parameters are used for all k(k-1)/2 binary classification problems.
We select parameters that achieve the highest overall performance.
</li>
</ol>
Each has its own advantages. A
single parameter set may not be uniformly good for all k(k-1)/2 decision functions.
However, as the overall accuracy is the final consideration, one parameter set
for one decision function may lead to over-fitting. In the paper
<p>
Chen, Lin, and Schölkopf,
<A HREF="../papers/nusvmtutorial.pdf">
A tutorial on nu-support vector machines.
</A>
Applied Stochastic Models in Business and Industry, 21(2005), 111-136,
<p>
they have experimentally
shown that the two methods give similar performance.
Therefore, currently the parameter selection in LIBSVM
takes the second approach by considering the same parameters for
all k(k-1)/2 models.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f420"><b>Q: After doing cross validation, why there is no model file outputted ?</b></a>
<br/>
<p>
Cross validation is used for selecting good parameters.
After finding them, you want to re-train the whole
data without the -v option.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f4201"><b>Q: Why my cross-validation results are different from those in the Practical Guide?</b></a>
<br/>
<p>
Due to random partitions of
the data, on different systems CV accuracy values
may be different.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f421"><b>Q: On some systems CV accuracy is the same in several runs. How could I use different data partitions? In other words, how do I set random seed in LIBSVM?</b></a>
<br/>
<p>
If you use GNU C library,
the default seed 1 is considered. Thus you always
get the same result of running svm-train -v.
To have different seeds, you can add the following code
in svm-train.c:
<pre>
#include <time.h>
</pre>
and in the beginning of main(),
<pre>
srand(time(0));
</pre>
Alternatively, if you are not using GNU C library
and would like to use a fixed seed, you can have
<pre>
srand(1);
</pre>
<p>
For Java, the random number generator
is initialized using the time information.
So results of two CV runs are different.
To fix the seed, after version 3.1 (released
in mid 2011), you can add
<pre>
svm.rand.setSeed(0);
</pre>
in the main() function of svm_train.java.
<p>
If you use CV to select parameters, it is recommended to use identical folds
under different parameters. In this case, you can consider fixing the seed.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f422"><b>Q: I would like to solve L2-loss SVM (i.e., error term is quadratic). How should I modify the code ?</b></a>
<br/>
<p>
It is extremely easy. Taking c-svc for example, to solve
<p>
min_w w^Tw/2 + C \sum max(0, 1- (y_i w^Tx_i+b))^2,
<p>
only two
places of svm.cpp have to be changed.
First, modify the following line of
solve_c_svc from
<pre>
s.Solve(l, SVC_Q(*prob,*param,y), minus_ones, y,
alpha, Cp, Cn, param->eps, si, param->shrinking);
</pre>
to
<pre>
s.Solve(l, SVC_Q(*prob,*param,y), minus_ones, y,
alpha, INF, INF, param->eps, si, param->shrinking);
</pre>
Second, in the class of SVC_Q, declare C as
a private variable:
<pre>
double C;
</pre>
In the constructor replace
<pre>
for(int i=0;i<prob.l;i++)
QD[i]= (Qfloat)(this->*kernel_function)(i,i);
</pre>
with
<pre>
this->C = param.C;
for(int i=0;i<prob.l;i++)
QD[i]= (Qfloat)(this->*kernel_function)(i,i)+0.5/C;
</pre>
Then in the subroutine get_Q, after the for loop, add
<pre>
if(i >= start && i < len)
data[i] += 0.5/C;
</pre>
<p>
For one-class svm, the modification is exactly the same. For SVR, you don't need an if statement like the above. Instead, you only need a simple assignment:
<pre>
data[real_i] += 0.5/C;
</pre>
<p>
For large linear L2-loss SVM, please use
<a href=../liblinear>LIBLINEAR</a>.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f424"><b>Q: How do I choose parameters for one-class svm as training data are in only one class?</b></a>
<br/>
<p>
You have pre-specified true positive rate in mind and then search for
parameters which achieve similar cross-validation accuracy.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f427"><b>Q: Why the code gives NaN (not a number) results?</b></a>
<br/>
<p>
This rarely happens, but few users reported the problem.
It seems that their
computers for training libsvm have the VPN client
running. The VPN software has some bugs and causes this
problem. Please try to close or disconnect the VPN client.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f428"><b>Q: Why on windows sometimes grid.py fails?</b></a>
<br/>
<p>
This problem shouldn't happen after version
2.85. If you are using earlier versions,
please download the latest one.
<!--
<p>
If you are using earlier
versions, the error message is probably
<pre>
Traceback (most recent call last):
File "grid.py", line 349, in ?
main()
File "grid.py", line 344, in main
redraw(db)
File "grid.py", line 132, in redraw
gnuplot.write("set term windows\n")
IOError: [Errno 22] Invalid argument
</pre>
<p>Please try to close gnuplot windows and rerun.
If the problem still occurs, comment the following
two lines in grid.py by inserting "#" in the beginning:
<pre>
redraw(db)
redraw(db,1)
</pre>
Then you get accuracy only but not cross validation contours.
-->
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f429"><b>Q: Why grid.py/easy.py sometimes generates the following warning message?</b></a>
<br/>
<pre>
Warning: empty z range [62.5:62.5], adjusting to [61.875:63.125]
Notice: cannot contour non grid data!
</pre>
<p>Nothing is wrong and please disregard the
message. It is from gnuplot when drawing
the contour.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f430"><b>Q: Why the sign of predicted labels and decision values are sometimes reversed?</b></a>
<br/>
<p>Nothing is wrong. Very likely you have two labels +1/-1 and the first instance in your data
has -1.
Think about the case of labels +5/+10. Since
SVM needs to use +1/-1, internally
we map +5/+10 to +1/-1 according to which
label appears first.
Hence a positive decision value implies
that we should predict the "internal" +1,
which may not be the +1 in the input file.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f431"><b>Q: I don't know class labels of test data. What should I put in the first column of the test file?</b></a>
<br/>
<p>Any value is ok. In this situation, what you will use is the output file of svm-predict, which gives predicted class labels.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f432"><b>Q: How can I use OpenMP to parallelize LIBSVM on a multicore/shared-memory computer?</b></a>
<br/>
<p>It is very easy if you are using GCC 4.2
or after.
<p> In Makefile, add -fopenmp to CFLAGS.
<p> In class SVC_Q of svm.cpp, modify the for loop
of get_Q to:
<pre>
#pragma omp parallel for private(j)
for(j=start;j<len;j++)
</pre>
<p> In the subroutine svm_predict_values of svm.cpp, add one line to the for loop:
<pre>
#pragma omp parallel for private(i)
for(i=0;i<l;i++)
kvalue[i] = Kernel::k_function(x,model->SV[i],model->param);
</pre>
For regression, you need to modify
class SVR_Q instead. The loop in svm_predict_values
is also different because you need
a reduction clause for the variable sum:
<pre>
#pragma omp parallel for private(i) reduction(+:sum)
for(i=0;i<model->l;i++)
sum += sv_coef[i] * Kernel::k_function(x,model->SV[i],model->param);
</pre>
<p> Then rebuild the package. Kernel evaluations in training/testing will be parallelized. An example of running this modification on
an 8-core machine using the data set
<a href=../libsvmtools/datasets/binary/ijcnn1.bz2>ijcnn1</a>:
<p> 8 cores:
<pre>
%setenv OMP_NUM_THREADS 8
%time svm-train -c 16 -g 4 -m 400 ijcnn1
27.1sec
</pre>
1 core:
<pre>
%setenv OMP_NUM_THREADS 1
%time svm-train -c 16 -g 4 -m 400 ijcnn1
79.8sec
</pre>
For this data, kernel evaluations take 80% of training time. In the above example, we assume you use csh. For bash, use
<pre>
export OMP_NUM_THREADS=8
</pre>
instead.
<p> For Python interface, you need to add the -lgomp link option:
<pre>
$(CXX) -lgomp -shared -dynamiclib svm.o -o libsvm.so.$(SHVER)
</pre>
<p> For MS Windows, you need to add /openmp in CFLAGS of Makefile.win
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f433"><b>Q: How could I know which training instances are support vectors?</b></a>
<br/>
<p>
It's very simple. Since version 3.13, you can use the function
<pre>
void svm_get_sv_indices(const struct svm_model *model, int *sv_indices)
</pre>
to get indices of support vectors. For example, in svm-train.c, after
<pre>
model = svm_train(&prob, &param);
</pre>
you can add
<pre>
int nr_sv = svm_get_nr_sv(model);
int *sv_indices = Malloc(int, nr_sv);
svm_get_sv_indices(model, sv_indices);
for (int i=0; i<nr_sv; i++)
printf("instance %d is a support vector\n", sv_indices[i]);
</pre>
<p> If you use matlab interface, you can directly check
<pre>
model.sv_indices
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q5:_Probability_outputs"></a>
<a name="f425"><b>Q: Why training a probability model (i.e., -b 1) takes a longer time?</b></a>
<br/>
<p>
To construct this probability model, we internally conduct a
cross validation, which is more time consuming than
a regular training.
Hence, in general you do parameter selection first without
-b 1. You only use -b 1 when good parameters have been
selected. In other words, you avoid using -b 1 and -v
together.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q5:_Probability_outputs"></a>
<a name="f426"><b>Q: Why using the -b option does not give me better accuracy?</b></a>
<br/>
<p>
There is absolutely no reason the probability outputs guarantee
you better accuracy. The main purpose of this option is
to provide you the probability estimates, but not to boost
prediction accuracy. From our experience,
after proper parameter selections, in general with
and without -b have similar accuracy. Occasionally there
are some differences.
It is not recommended to compare the two under
just a fixed parameter
set as more differences will be observed.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q5:_Probability_outputs"></a>
<a name="f427"><b>Q: Why using svm-predict -b 0 and -b 1 gives different accuracy values?</b></a>
<br/>
<p>
Let's just consider two-class classification here. After probability information is obtained in training,
we do not have
<p>
prob > = 0.5 if and only if decision value >= 0.
<p>
So predictions may be different with -b 0 and 1.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q6:_Graphic_interface"></a>
<a name="f501"><b>Q: How can I save images drawn by svm-toy?</b></a>
<br/>
<p>
For Microsoft windows, first press the "print screen" key on the keyboard.
Open "Microsoft Paint"
(included in Windows)
and press "ctrl-v." Then you can clip
the part of picture which you want.
For X windows, you can
use the program "xv" or "import" to grab the picture of the svm-toy window.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q6:_Graphic_interface"></a>
<a name="f502"><b>Q: I press the "load" button to load data points but why svm-toy does not draw them ?</b></a>
<br/>
<p>
The program svm-toy assumes both attributes (i.e. x-axis and y-axis
values) are in (0,1). Hence you want to scale your
data to between a small positive number and
a number less than but very close to 1.
Moreover, class labels must be 1, 2, or 3
(not 1.0, 2.0 or anything else).
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q6:_Graphic_interface"></a>
<a name="f503"><b>Q: I would like svm-toy to handle more than three classes of data, what should I do ?</b></a>
<br/>
<p>
Taking windows/svm-toy.cpp as an example, you need to
modify it and the difference
from the original file is as the following: (for five classes of
data)
<pre>
30,32c30
< RGB(200,0,200),
< RGB(0,160,0),
< RGB(160,0,0)
---
> RGB(200,0,200)
39c37
< HBRUSH brush1, brush2, brush3, brush4, brush5;
---
> HBRUSH brush1, brush2, brush3;
113,114d110
< brush4 = CreateSolidBrush(colors[7]);
< brush5 = CreateSolidBrush(colors[8]);
155,157c151
< else if(v==3) return brush3;
< else if(v==4) return brush4;
< else return brush5;
---
> else return brush3;
325d318
< int colornum = 5;
327c320
< svm_node *x_space = new svm_node[colornum * prob.l];
---
> svm_node *x_space = new svm_node[3 * prob.l];
333,338c326,331
< x_space[colornum * i].index = 1;
< x_space[colornum * i].value = q->x;
< x_space[colornum * i + 1].index = 2;
< x_space[colornum * i + 1].value = q->y;
< x_space[colornum * i + 2].index = -1;
< prob.x[i] = &x_space[colornum * i];
---
> x_space[3 * i].index = 1;
> x_space[3 * i].value = q->x;
> x_space[3 * i + 1].index = 2;
> x_space[3 * i + 1].value = q->y;
> x_space[3 * i + 2].index = -1;
> prob.x[i] = &x_space[3 * i];
397c390
< if(current_value > 5) current_value = 1;
---
> if(current_value > 3) current_value = 1;
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q7:_Java_version_of_libsvm"></a>
<a name="f601"><b>Q: What is the difference between Java version and C++ version of libsvm?</b></a>
<br/>
<p>
They are the same thing. We just rewrote the C++ code
in Java.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q7:_Java_version_of_libsvm"></a>
<a name="f602"><b>Q: Is the Java version significantly slower than the C++ version?</b></a>
<br/>
<p>
This depends on the VM you used. We have seen good
VM which leads the Java version to be quite competitive with
the C++ code. (though still slower)
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q7:_Java_version_of_libsvm"></a>
<a name="f603"><b>Q: While training I get the following error message: java.lang.OutOfMemoryError. What is wrong?</b></a>
<br/>
<p>
You should try to increase the maximum Java heap size.
For example,
<pre>
java -Xmx2048m -classpath libsvm.jar svm_train ...
</pre>
sets the maximum heap size to 2048M.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q7:_Java_version_of_libsvm"></a>
<a name="f604"><b>Q: Why you have the main source file svm.m4 and then transform it to svm.java?</b></a>
<br/>
<p>
Unlike C, Java does not have a preprocessor built-in.
However, we need some macros (see first 3 lines of svm.m4).
</ul>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q8:_Python_interface"></a>
<a name="f704"><b>Q: Except the python-C++ interface provided, could I use Jython to call libsvm ?</b></a>
<br/>
<p> Yes, here are some examples:
<pre>
$ export CLASSPATH=$CLASSPATH:~/libsvm-2.91/java/libsvm.jar
$ ./jython
Jython 2.1a3 on java1.3.0 (JIT: jitc)
Type "copyright", "credits" or "license" for more information.
>>> from libsvm import *
>>> dir()
['__doc__', '__name__', 'svm', 'svm_model', 'svm_node', 'svm_parameter',
'svm_problem']
>>> x1 = [svm_node(index=1,value=1)]
>>> x2 = [svm_node(index=1,value=-1)]
>>> param = svm_parameter(svm_type=0,kernel_type=2,gamma=1,cache_size=40,eps=0.001,C=1,nr_weight=0,shrinking=1)
>>> prob = svm_problem(l=2,y=[1,-1],x=[x1,x2])
>>> model = svm.svm_train(prob,param)
*
optimization finished, #iter = 1
nu = 1.0
obj = -1.018315639346838, rho = 0.0
nSV = 2, nBSV = 2
Total nSV = 2
>>> svm.svm_predict(model,x1)
1.0
>>> svm.svm_predict(model,x2)
-1.0
>>> svm.svm_save_model("test.model",model)
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f801"><b>Q: I compile the MATLAB interface without problem, but why errors occur while running it?</b></a>
<br/>
<p>
Your compiler version may not be supported/compatible for MATLAB.
Please check <a href=http://www.mathworks.com/support/compilers/current_release>this MATLAB page</a> first and then specify the version
number. For example, if g++ X.Y is supported, replace
<pre>
CXX = g++
</pre>
in the Makefile with
<pre>
CXX = g++-X.Y
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f8011"><b>Q: On 64bit Windows I compile the MATLAB interface without problem, but why errors occur while running it?</b></a>
<br/>
<p>
Please make sure that you use
the -largeArrayDims option in make.m. For example,
<pre>
mex -largeArrayDims -O -c svm.cpp
</pre>
Moreover, if you use Microsoft Visual Studio,
probabally it is not properly installed.
See the explanation
<a href=http://www.mathworks.com/support/compilers/current_release/win64.html#n7>here</a>.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f802"><b>Q: Does the MATLAB interface provide a function to do scaling?</b></a>
<br/>
<p>
It is extremely easy to do scaling under MATLAB.
The following one-line code scale each feature to the range
of [0,1]:
<pre>
(data - repmat(min(data,[],1),size(data,1),1))*spdiags(1./(max(data,[],1)-min(data,[],1))',0,size(data,2),size(data,2))
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f803"><b>Q: How could I use MATLAB interface for parameter selection?</b></a>
<br/>
<p>
One can do this by a simple loop.
See the following example:
<pre>
bestcv = 0;
for log2c = -1:3,
for log2g = -4:1,
cmd = ['-v 5 -c ', num2str(2^log2c), ' -g ', num2str(2^log2g)];
cv = svmtrain(heart_scale_label, heart_scale_inst, cmd);
if (cv >= bestcv),
bestcv = cv; bestc = 2^log2c; bestg = 2^log2g;
end
fprintf('%g %g %g (best c=%g, g=%g, rate=%g)\n', log2c, log2g, cv, bestc, bestg, bestcv);
end
end
</pre>
You may adjust the parameter range in the above loops.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f8031"><b>Q: I use MATLAB parallel programming toolbox on a multi-core environment for parameter selection. Why the program is even slower?</b></a>
<br/>
<p>
Fabrizio Lacalandra of University of Pisa reported this issue.
It seems the problem is caused by the screen output.
If you disable the <b>info</b> function
using <pre>#if 0,</pre> then the problem
may be solved.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f8032"><b>Q: How do I use LIBSVM with OpenMP under MATLAB?</b></a>
<br/>
<p>
In Makefile,
you need to add -fopenmp to CFLAGS and -lgomp to MEX_OPTION. For Octave, you need the same modification.
<p> However, a minor problem is that
the number of threads cannot
be specified in MATLAB. We tried Version 7.12 (R2011a) and gcc-4.6.1.
<pre>
% export OMP_NUM_THREADS=4; matlab
>> setenv('OMP_NUM_THREADS', '1');
</pre>
Then OMP_NUM_THREADS is still 4 while running the program. Please contact us if you
see how to solve this problem. You can, however,
specify the number in the source code (thanks
to comments from Ricardo Santiago-mozos):
<pre>
#pragma omp parallel for private(i) num_threads(4)
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f804"><b>Q: How could I generate the primal variable w of linear SVM?</b></a>
<br/>
<p>
Let's start from the binary class and
assume you have two labels -1 and +1.
After obtaining the model from calling svmtrain,
do the following to have w and b:
<pre>
w = model.SVs' * model.sv_coef;
b = -model.rho;
if model.Label(1) == -1
w = -w;
b = -b;
end
</pre>
If you do regression or one-class SVM, then the if statement is not needed.
<p> For multi-class SVM, we illustrate the setting
in the following example of running the iris
data, which have 3 classes
<pre>
> [y, x] = libsvmread('../../htdocs/libsvmtools/datasets/multiclass/iris.scale');
> m = svmtrain(y, x, '-t 0')
m =
Parameters: [5x1 double]
nr_class: 3
totalSV: 42
rho: [3x1 double]
Label: [3x1 double]
ProbA: []
ProbB: []
nSV: [3x1 double]
sv_coef: [42x2 double]
SVs: [42x4 double]
</pre>
sv_coef is like:
<pre>
+-+-+--------------------+
|1|1| |
|v|v| SVs from class 1 |
|2|3| |
+-+-+--------------------+
|1|2| |
|v|v| SVs from class 2 |
|2|3| |
+-+-+--------------------+
|1|2| |
|v|v| SVs from class 3 |
|3|3| |
+-+-+--------------------+
</pre>
so we need to see nSV of each classes.
<pre>
> m.nSV
ans =
3
21
18
</pre>
Suppose the goal is to find the vector w of classes
1 vs 3. Then
y_i alpha_i of training 1 vs 3 are
<pre>
> coef = [m.sv_coef(1:3,2); m.sv_coef(25:42,1)];
</pre>
and SVs are:
<pre>
> SVs = [m.SVs(1:3,:); m.SVs(25:42,:)];
</pre>
Hence, w is
<pre>
> w = SVs'*coef;
</pre>
For rho,
<pre>
> m.rho
ans =
1.1465
0.3682
-1.9969
> b = -m.rho(2);
</pre>
because rho is arranged by 1vs2 1vs3 2vs3.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f805"><b>Q: Is there an OCTAVE interface for libsvm?</b></a>
<br/>
<p>
Yes, after libsvm 2.86, the matlab interface
works on OCTAVE as well. Please use make.m by typing
<pre>
>> make
</pre>
under OCTAVE.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f806"><b>Q: How to handle the name conflict between svmtrain in the libsvm matlab interface and that in MATLAB bioinformatics toolbox?</b></a>
<br/>
<p>
The easiest way is to rename the svmtrain binary
file (e.g., svmtrain.mexw32 on 32-bit windows)
to a different
name (e.g., svmtrain2.mexw32).
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f807"><b>Q: On Windows I got an error message "Invalid MEX-file: Specific module not found" when running the pre-built MATLAB interface in the windows sub-directory. What should I do?</b></a>
<br/>
<p>
The error usually happens
when there are missing runtime components
such as MSVCR100.dll on your Windows platform.
You can use tools such as
<a href=http://www.dependencywalker.com/>Dependency
Walker</a> to find missing library files.
<p>
For example, if the pre-built MEX files are compiled by
Visual C++ 2010,
you must have installed
Microsoft Visual C++ Redistributable Package 2010
(vcredist_x86.exe). You can easily find the freely
available file from Microsoft's web site.
<p>
For 64bit Windows, the situation is similar. If
the pre-built files are by
Visual C++ 2008, then you must have
Microsoft Visual C++ Redistributable Package 2008
(vcredist_x64.exe).
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q9:_MATLAB_interface"></a>
<a name="f808"><b>Q: LIBSVM supports 1-vs-1 multi-class classification. If instead I would like to use 1-vs-rest, how to implement it using MATLAB interface?</b></a>
<br/>
<p>
Please use code in the following <a href=../libsvmtools/ovr_multiclass>directory</a>. The following example shows how to
train and test the problem dna (<a href=../libsvmtools/datasets/multiclass/dna.scale>training</a> and <a href=../libsvmtools/datasets/multiclass/dna.scale.t>testing</a>).
<p> Load, train and predict data:
<pre>
[trainY trainX] = libsvmread('./dna.scale');
[testY testX] = libsvmread('./dna.scale.t');
model = ovrtrain(trainY, trainX, '-c 8 -g 4');
[pred ac decv] = ovrpredict(testY, testX, model);
fprintf('Accuracy = %g%%\n', ac * 100);
</pre>
Conduct CV on a grid of parameters
<pre>
bestcv = 0;
for log2c = -1:2:3,
for log2g = -4:2:1,
cmd = ['-q -c ', num2str(2^log2c), ' -g ', num2str(2^log2g)];
cv = get_cv_ac(trainY, trainX, cmd, 3);
if (cv >= bestcv),
bestcv = cv; bestc = 2^log2c; bestg = 2^log2g;
end
fprintf('%g %g %g (best c=%g, g=%g, rate=%g)\n', log2c, log2g, cv, bestc, bestg, bestcv);
end
end
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<p align="middle">
<a href="http://www.csie.ntu.edu.tw/~cjlin/libsvm">LIBSVM home page</a>
</p>
</body>
</html>
================================================
FILE: binaries/linux/README
================================================
Libsvm is a simple, easy-to-use, and efficient software for SVM
classification and regression. It solves C-SVM classification, nu-SVM
classification, one-class-SVM, epsilon-SVM regression, and nu-SVM
regression. It also provides an automatic model selection tool for
C-SVM classification. This document explains the use of libsvm.
Libsvm is available at
http://www.csie.ntu.edu.tw/~cjlin/libsvm
Please read the COPYRIGHT file before using libsvm.
Table of Contents
=================
- Quick Start
- Installation and Data Format
- `svm-train' Usage
- `svm-predict' Usage
- `svm-scale' Usage
- Tips on Practical Use
- Examples
- Precomputed Kernels
- Library Usage
- Java Version
- Building Windows Binaries
- Additional Tools: Sub-sampling, Parameter Selection, Format checking, etc.
- MATLAB/OCTAVE Interface
- Python Interface
- Additional Information
Quick Start
===========
If you are new to SVM and if the data is not large, please go to
`tools' directory and use easy.py after installation. It does
everything automatic -- from data scaling to parameter selection.
Usage: easy.py training_file [testing_file]
More information about parameter selection can be found in
`tools/README.'
Installation and Data Format
============================
On Unix systems, type `make' to build the `svm-train' and `svm-predict'
programs. Run them without arguments to show the usages of them.
On other systems, consult `Makefile' to build them (e.g., see
'Building Windows binaries' in this file) or use the pre-built
binaries (Windows binaries are in the directory `windows').
The format of training and testing data file is:
<label> <index1>:<value1> <index2>:<value2> ...
.
.
.
Each line contains an instance and is ended by a '\n' character. For
classification, <label> is an integer indicating the class label
(multi-class is supported). For regression, <label> is the target
value which can be any real number. For one-class SVM, it's not used
so can be any number. The pair <index>:<value> gives a feature
(attribute) value: <index> is an integer starting from 1 and <value>
is a real number. The only exception is the precomputed kernel, where
<index> starts from 0; see the section of precomputed kernels. Indices
must be in ASCENDING order. Labels in the testing file are only used
to calculate accuracy or errors. If they are unknown, just fill the
first column with any numbers.
A sample classification data included in this package is
`heart_scale'. To check if your data is in a correct form, use
`tools/checkdata.py' (details in `tools/README').
Type `svm-train heart_scale', and the program will read the training
data and output the model file `heart_scale.model'. If you have a test
set called heart_scale.t, then type `svm-predict heart_scale.t
heart_scale.model output' to see the prediction accuracy. The `output'
file contains the predicted class labels.
For classification, if training data are in only one class (i.e., all
labels are the same), then `svm-train' issues a warning message:
`Warning: training data in only one class. See README for details,'
which means the training data is very unbalanced. The label in the
training data is directly returned when testing.
There are some other useful programs in this package.
svm-scale:
This is a tool for scaling input data file.
svm-toy:
This is a simple graphical interface which shows how SVM
separate data in a plane. You can click in the window to
draw data points. Use "change" button to choose class
1, 2 or 3 (i.e., up to three classes are supported), "load"
button to load data from a file, "save" button to save data to
a file, "run" button to obtain an SVM model, and "clear"
button to clear the window.
You can enter options in the bottom of the window, the syntax of
options is the same as `svm-train'.
Note that "load" and "save" consider dense data format both in
classification and the regression cases. For classification,
each data point has one label (the color) that must be 1, 2,
or 3 and two attributes (x-axis and y-axis values) in
[0,1). For regression, each data point has one target value
(y-axis) and one attribute (x-axis values) in [0, 1).
Type `make' in respective directories to build them.
You need Qt library to build the Qt version.
(available from http://www.trolltech.com)
You need GTK+ library to build the GTK version.
(available from http://www.gtk.org)
The pre-built Windows binaries are in the `windows'
directory. We use Visual C++ on a 32-bit machine, so the
maximal cache size is 2GB.
`svm-train' Usage
=================
Usage: svm-train [options] training_set_file [model_file]
options:
-s svm_type : set type of SVM (default 0)
0 -- C-SVC (multi-class classification)
1 -- nu-SVC (multi-class classification)
2 -- one-class SVM
3 -- epsilon-SVR (regression)
4 -- nu-SVR (regression)
-t kernel_type : set type of kernel function (default 2)
0 -- linear: u'*v
1 -- polynomial: (gamma*u'*v + coef0)^degree
2 -- radial basis function: exp(-gamma*|u-v|^2)
3 -- sigmoid: tanh(gamma*u'*v + coef0)
4 -- precomputed kernel (kernel values in training_set_file)
-d degree : set degree in kernel function (default 3)
-g gamma : set gamma in kernel function (default 1/num_features)
-r coef0 : set coef0 in kernel function (default 0)
-c cost : set the parameter C of C-SVC, epsilon-SVR, and nu-SVR (default 1)
-n nu : set the parameter nu of nu-SVC, one-class SVM, and nu-SVR (default 0.5)
-p epsilon : set the epsilon in loss function of epsilon-SVR (default 0.1)
-m cachesize : set cache memory size in MB (default 100)
-e epsilon : set tolerance of termination criterion (default 0.001)
-h shrinking : whether to use the shrinking heuristics, 0 or 1 (default 1)
-b probability_estimates : whether to train a SVC or SVR model for probability estimates, 0 or 1 (default 0)
-wi weight : set the parameter C of class i to weight*C, for C-SVC (default 1)
-v n: n-fold cross validation mode
-q : quiet mode (no outputs)
The k in the -g option means the number of attributes in the input data.
option -v randomly splits the data into n parts and calculates cross
validation accuracy/mean squared error on them.
See libsvm FAQ for the meaning of outputs.
`svm-predict' Usage
===================
Usage: svm-predict [options] test_file model_file output_file
options:
-b probability_estimates: whether to predict probability estimates, 0 or 1 (default 0); for one-class SVM only 0 is supported
model_file is the model file generated by svm-train.
test_file is the test data you want to predict.
svm-predict will produce output in the output_file.
`svm-scale' Usage
=================
Usage: svm-scale [options] data_filename
options:
-l lower : x scaling lower limit (default -1)
-u upper : x scaling upper limit (default +1)
-y y_lower y_upper : y scaling limits (default: no y scaling)
-s save_filename : save scaling parameters to save_filename
-r restore_filename : restore scaling parameters from restore_filename
See 'Examples' in this file for examples.
Tips on Practical Use
=====================
* Scale your data. For example, scale each attribute to [0,1] or [-1,+1].
* For C-SVC, consider using the model selection tool in the tools directory.
* nu in nu-SVC/one-class-SVM/nu-SVR approximates the fraction of training
errors and support vectors.
* If data for classification are unbalanced (e.g. many positive and
few negative), try different penalty parameters C by -wi (see
examples below).
* Specify larger cache size (i.e., larger -m) for huge problems.
Examples
========
> svm-scale -l -1 -u 1 -s range train > train.scale
> svm-scale -r range test > test.scale
Scale each feature of the training data to be in [-1,1]. Scaling
factors are stored in the file range and then used for scaling the
test data.
> svm-train -s 0 -c 5 -t 2 -g 0.5 -e 0.1 data_file
Train a classifier with RBF kernel exp(-0.5|u-v|^2), C=10, and
stopping tolerance 0.1.
> svm-train -s 3 -p 0.1 -t 0 data_file
Solve SVM regression with linear kernel u'v and epsilon=0.1
in the loss function.
> svm-train -c 10 -w1 1 -w-2 5 -w4 2 data_file
Train a classifier with penalty 10 = 1 * 10 for class 1, penalty 50 =
5 * 10 for class -2, and penalty 20 = 2 * 10 for class 4.
> svm-train -s 0 -c 100 -g 0.1 -v 5 data_file
Do five-fold cross validation for the classifier using
the parameters C = 100 and gamma = 0.1
> svm-train -s 0 -b 1 data_file
> svm-predict -b 1 test_file data_file.model output_file
Obtain a model with probability information and predict test data with
probability estimates
Precomputed Kernels
===================
Users may precompute kernel values and input them as training and
testing files. Then libsvm does not need the original
training/testing sets.
Assume there are L training instances x1, ..., xL and.
Let K(x, y) be the kernel
value of two instances x and y. The input formats
are:
New training instance for xi:
<label> 0:i 1:K(xi,x1) ... L:K(xi,xL)
New testing instance for any x:
<label> 0:? 1:K(x,x1) ... L:K(x,xL)
That is, in the training file the first column must be the "ID" of
xi. In testing, ? can be any value.
All kernel values including ZEROs must be explicitly provided. Any
permutation or random subsets of the training/testing files are also
valid (see examples below).
Note: the format is slightly different from the precomputed kernel
package released in libsvmtools earlier.
Examples:
Assume the original training data has three four-feature
instances and testing data has one instance:
15 1:1 2:1 3:1 4:1
45 2:3 4:3
25 3:1
15 1:1 3:1
If the linear kernel is used, we have the following new
training/testing sets:
15 0:1 1:4 2:6 3:1
45 0:2 1:6 2:18 3:0
25 0:3 1:1 2:0 3:1
15 0:? 1:2 2:0 3:1
? can be any value.
Any subset of the above training file is also valid. For example,
25 0:3 1:1 2:0 3:1
45 0:2 1:6 2:18 3:0
implies that the kernel matrix is
[K(2,2) K(2,3)] = [18 0]
[K(3,2) K(3,3)] = [0 1]
Library Usage
=============
These functions and structures are declared in the header file
`svm.h'. You need to #include "svm.h" in your C/C++ source files and
link your program with `svm.cpp'. You can see `svm-train.c' and
`svm-predict.c' for examples showing how to use them. We define
LIBSVM_VERSION and declare `extern int libsvm_version; ' in svm.h, so
you can check the version number.
Before you classify test data, you need to construct an SVM model
(`svm_model') using training data. A model can also be saved in
a file for later use. Once an SVM model is available, you can use it
to classify new data.
- Function: struct svm_model *svm_train(const struct svm_problem *prob,
const struct svm_parameter *param);
This function constructs and returns an SVM model according to
the given training data and parameters.
struct svm_problem describes the problem:
struct svm_problem
{
int l;
double *y;
struct svm_node **x;
};
where `l' is the number of training data, and `y' is an array containing
their target values. (integers in classification, real numbers in
regression) `x' is an array of pointers, each of which points to a sparse
representation (array of svm_node) of one training vector.
For example, if we have the following training data:
LABEL ATTR1 ATTR2 ATTR3 ATTR4 ATTR5
----- ----- ----- ----- ----- -----
1 0 0.1 0.2 0 0
2 0 0.1 0.3 -1.2 0
1 0.4 0 0 0 0
2 0 0.1 0 1.4 0.5
3 -0.1 -0.2 0.1 1.1 0.1
then the components of svm_problem are:
l = 5
y -> 1 2 1 2 3
x -> [ ] -> (2,0.1) (3,0.2) (-1,?)
[ ] -> (2,0.1) (3,0.3) (4,-1.2) (-1,?)
[ ] -> (1,0.4) (-1,?)
[ ] -> (2,0.1) (4,1.4) (5,0.5) (-1,?)
[ ] -> (1,-0.1) (2,-0.2) (3,0.1) (4,1.1) (5,0.1) (-1,?)
where (index,value) is stored in the structure `svm_node':
struct svm_node
{
int index;
double value;
};
index = -1 indicates the end of one vector. Note that indices must
be in ASCENDING order.
struct svm_parameter describes the parameters of an SVM model:
struct svm_parameter
{
int svm_type;
int kernel_type;
int degree; /* for poly */
double gamma; /* for poly/rbf/sigmoid */
double coef0; /* for poly/sigmoid */
/* these are for training only */
double cache_size; /* in MB */
double eps; /* stopping criteria */
double C; /* for C_SVC, EPSILON_SVR, and NU_SVR */
int nr_weight; /* for C_SVC */
int *weight_label; /* for C_SVC */
double* weight; /* for C_SVC */
double nu; /* for NU_SVC, ONE_CLASS, and NU_SVR */
double p; /* for EPSILON_SVR */
int shrinking; /* use the shrinking heuristics */
int probability; /* do probability estimates */
};
svm_type can be one of C_SVC, NU_SVC, ONE_CLASS, EPSILON_SVR, NU_SVR.
C_SVC: C-SVM classification
NU_SVC: nu-SVM classification
ONE_CLASS: one-class-SVM
EPSILON_SVR: epsilon-SVM regression
NU_SVR: nu-SVM regression
kernel_type can be one of LINEAR, POLY, RBF, SIGMOID.
LINEAR: u'*v
POLY: (gamma*u'*v + coef0)^degree
RBF: exp(-gamma*|u-v|^2)
SIGMOID: tanh(gamma*u'*v + coef0)
PRECOMPUTED: kernel values in training_set_file
cache_size is the size of the kernel cache, specified in megabytes.
C is the cost of constraints violation.
eps is the stopping criterion. (we usually use 0.00001 in nu-SVC,
0.001 in others). nu is the parameter in nu-SVM, nu-SVR, and
one-class-SVM. p is the epsilon in epsilon-insensitive loss function
of epsilon-SVM regression. shrinking = 1 means shrinking is conducted;
= 0 otherwise. probability = 1 means model with probability
information is obtained; = 0 otherwise.
nr_weight, weight_label, and weight are used to change the penalty
for some classes (If the weight for a class is not changed, it is
set to 1). This is useful for training classifier using unbalanced
input data or with asymmetric misclassification cost.
nr_weight is the number of elements in the array weight_label and
weight. Each weight[i] corresponds to weight_label[i], meaning that
the penalty of class weight_label[i] is scaled by a factor of weight[i].
If you do not want to change penalty for any of the classes,
just set nr_weight to 0.
*NOTE* Because svm_model contains pointers to svm_problem, you can
not free the memory used by svm_problem if you are still using the
svm_model produced by svm_train().
*NOTE* To avoid wrong parameters, svm_check_parameter() should be
called before svm_train().
struct svm_model stores the model obtained from the training procedure.
It is not recommended to directly access entries in this structure.
Programmers should use the interface functions to get the values.
struct svm_model
{
struct svm_parameter param; /* parameter */
int nr_class; /* number of classes, = 2 in regression/one class svm */
int l; /* total #SV */
struct svm_node **SV; /* SVs (SV[l]) */
double **sv_coef; /* coefficients for SVs in decision functions (sv_coef[k-1][l]) */
double *rho; /* constants in decision functions (rho[k*(k-1)/2]) */
double *probA; /* pairwise probability information */
double *probB;
int *sv_indices; /* sv_indices[0,...,nSV-1] are values in [1,...,num_traning_data] to indicate SVs in the training set */
/* for classification only */
int *label; /* label of each class (label[k]) */
int *nSV; /* number of SVs for each class (nSV[k]) */
/* nSV[0] + nSV[1] + ... + nSV[k-1] = l */
/* XXX */
int free_sv; /* 1 if svm_model is created by svm_load_model*/
/* 0 if svm_model is created by svm_train */
};
param describes the parameters used to obtain the model.
nr_class is the number of classes. It is 2 for regression and one-class SVM.
l is the number of support vectors. SV and sv_coef are support
vectors and the corresponding coefficients, respectively. Assume there are
k classes. For data in class j, the corresponding sv_coef includes (k-1) y*alpha vectors,
where alpha's are solutions of the following two class problems:
1 vs j, 2 vs j, ..., j-1 vs j, j vs j+1, j vs j+2, ..., j vs k
and y=1 for the first j-1 vectors, while y=-1 for the remaining k-j
vectors. For example, if there are 4 classes, sv_coef and SV are like:
+-+-+-+--------------------+
|1|1|1| |
|v|v|v| SVs from class 1 |
|2|3|4| |
+-+-+-+--------------------+
|1|2|2| |
|v|v|v| SVs from class 2 |
|2|3|4| |
+-+-+-+--------------------+
|1|2|3| |
|v|v|v| SVs from class 3 |
|3|3|4| |
+-+-+-+--------------------+
|1|2|3| |
|v|v|v| SVs from class 4 |
|4|4|4| |
+-+-+-+--------------------+
See svm_train() for an example of assigning values to sv_coef.
rho is the bias term (-b). probA and probB are parameters used in
probability outputs. If there are k classes, there are k*(k-1)/2
binary problems as well as rho, probA, and probB values. They are
aligned in the order of binary problems:
1 vs 2, 1 vs 3, ..., 1 vs k, 2 vs 3, ..., 2 vs k, ..., k-1 vs k.
sv_indices[0,...,nSV-1] are values in [1,...,num_traning_data] to
indicate support vectors in the training set.
label contains labels in the training data.
nSV is the number of support vectors in each class.
free_sv is a flag used to determine whether the space of SV should
be released in free_model_content(struct svm_model*) and
free_and_destroy_model(struct svm_model**). If the model is
generated by svm_train(), then SV points to data in svm_problem
and should not be removed. For example, free_sv is 0 if svm_model
is created by svm_train, but is 0 if created by svm_load_model.
- Function: double svm_predict(const struct svm_model *model,
const struct svm_node *x);
This function does classification or regression on a test vector x
given a model.
For a classification model, the predicted class for x is returned.
For a regression model, the function value of x calculated using
the model is returned. For an one-class model, +1 or -1 is
returned.
- Function: void svm_cross_validation(const struct svm_problem *prob,
const struct svm_parameter *param, int nr_fold, double *target);
This function conducts cross validation. Data are separated to
nr_fold folds. Under given parameters, sequentially each fold is
validated using the model from training the remaining. Predicted
labels (of all prob's instances) in the validation process are
stored in the array called target.
The format of svm_prob is same as that for svm_train().
- Function: int svm_get_svm_type(const struct svm_model *model);
This function gives svm_type of the model. Possible values of
svm_type are defined in svm.h.
- Function: int svm_get_nr_class(const svm_model *model);
For a classification model, this function gives the number of
classes. For a regression or an one-class model, 2 is returned.
- Function: void svm_get_labels(const svm_model *model, int* label)
For a classification model, this function outputs the name of
labels into an array called label. For regression and one-class
models, label is unchanged.
- Function: void svm_get_sv_indices(const struct svm_model *model, int *sv_indices)
This function outputs indices of support vectors into an array called sv_indices.
The size of sv_indices is the number of support vectors and can be obtained by calling svm_get_nr_sv.
Each sv_indices[i] is in the range of [1, ..., num_traning_data].
- Function: int svm_get_nr_sv(const struct svm_model *model)
This function gives the number of total support vector.
- Function: double svm_get_svr_probability(const struct svm_model *model);
For a regression model with probability information, this function
outputs a value sigma > 0. For test data, we consider the
probability model: target value = predicted value + z, z: Laplace
distribution e^(-|z|/sigma)/(2sigma)
If the model is not for svr or does not contain required
information, 0 is returned.
- Function: double svm_predict_values(const svm_model *model,
const svm_node *x, double* dec_values)
This function gives decision values on a test vector x given a
model, and return the predicted label (classification) or
the function value (regression).
For a classification model with nr_class classes, this function
gives nr_class*(nr_class-1)/2 decision values in the array
dec_values, where nr_class can be obtained from the function
svm_get_nr_class. The order is label[0] vs. label[1], ...,
label[0] vs. label[nr_class-1], label[1] vs. label[2], ...,
label[nr_class-2] vs. label[nr_class-1], where label can be
obtained from the function svm_get_labels. The returned value is
the predicted class for x. Note that when nr_class = 1, this
function does not give any decision value.
For a regression model, dec_values[0] and the returned value are
both the function value of x calculated using the model. For a
one-class model, dec_values[0] is the decision value of x, while
the returned value is +1/-1.
- Function: double svm_predict_probability(const struct svm_model *model,
const struct svm_node *x, double* prob_estimates);
This function does classification or regression on a test vector x
given a model with probability information.
For a classification model with probability information, this
function gives nr_class probability estimates in the array
prob_estimates. nr_class can be obtained from the function
svm_get_nr_class. The class with the highest probability is
returned. For regression/one-class SVM, the array prob_estimates
is unchanged and the returned value is the same as that of
svm_predict.
- Function: const char *svm_check_parameter(const struct svm_problem *prob,
const struct svm_parameter *param);
This function checks whether the parameters are within the feasible
range of the problem. This function should be called before calling
svm_train() and svm_cross_validation(). It returns NULL if the
parameters are feasible, otherwise an error message is returned.
- Function: int svm_check_probability_model(const struct svm_model *model);
This function checks whether the model contains required
information to do probability estimates. If so, it returns
+1. Otherwise, 0 is returned. This function should be called
before calling svm_get_svr_probability and
svm_predict_probability.
- Function: int svm_save_model(const char *model_file_name,
const struct svm_model *model);
This function saves a model to a file; returns 0 on success, or -1
if an error occurs.
- Function: struct svm_model *svm_load_model(const char *model_file_name);
This function returns a pointer to the model read from the file,
or a null pointer if the model could not be loaded.
- Function: void svm_free_model_content(struct svm_model *model_ptr);
This function frees the memory used by the entries in a model structure.
- Function: void svm_free_and_destroy_model(struct svm_model **model_ptr_ptr);
This function frees the memory used by a model and destroys the model
structure. It is equivalent to svm_destroy_model, which
is deprecated after version 3.0.
- Function: void svm_destroy_param(struct svm_parameter *param);
This function frees the memory used by a parameter set.
- Function: void svm_set_print_string_function(void (*print_func)(const char *));
Users can specify their output format by a function. Use
svm_set_print_string_function(NULL);
for default printing to stdout.
Java Version
============
The pre-compiled java class archive `libsvm.jar' and its source files are
in the java directory. To run the programs, use
java -classpath libsvm.jar svm_train <arguments>
java -classpath libsvm.jar svm_predict <arguments>
java -classpath libsvm.jar svm_toy
java -classpath libsvm.jar svm_scale <arguments>
Note that you need Java 1.5 (5.0) or above to run it.
You may need to add Java runtime library (like classes.zip) to the classpath.
You may need to increase maximum Java heap size.
Library usages are similar to the C version. These functions are available:
public class svm {
public static final int LIBSVM_VERSION=317;
public static svm_model svm_train(svm_problem prob, svm_parameter param);
public static void svm_cross_validation(svm_problem prob, svm_parameter param, int nr_fold, double[] target);
public static int svm_get_svm_type(svm_model model);
public static int svm_get_nr_class(svm_model model);
public static void svm_get_labels(svm_model model, int[] label);
public static void svm_get_sv_indices(svm_model model, int[] indices);
public static int svm_get_nr_sv(svm_model model);
public static double svm_get_svr_probability(svm_model model);
public static double svm_predict_values(svm_model model, svm_node[] x, double[] dec_values);
public static double svm_predict(svm_model model, svm_node[] x);
public static double svm_predict_probability(svm_model model, svm_node[] x, double[] prob_estimates);
public static void svm_save_model(String model_file_name, svm_model model) throws IOException
public static svm_model svm_load_model(String model_file_name) throws IOException
public static String svm_check_parameter(svm_problem prob, svm_parameter param);
public static int svm_check_probability_model(svm_model model);
public static void svm_set_print_string_function(svm_print_interface print_func);
}
The library is in the "libsvm" package.
Note that in Java version, svm_node[] is not ended with a node whose index = -1.
Users can specify their output format by
your_print_func = new svm_print_interface()
{
public void print(String s)
{
// your own format
}
};
svm.svm_set_print_string_function(your_print_func);
Building Windows Binaries
=========================
Windows binaries are in the directory `windows'. To build them via
Visual C++, use the following steps:
1. Open a DOS command box (or Visual Studio Command Prompt) and change
to libsvm directory. If environment variables of VC++ have not been
set, type
"C:\Program Files\Microsoft Visual Studio 10.0\VC\bin\vcvars32.bat"
You may have to modify the above command according which version of
VC++ or where it is installed.
2. Type
nmake -f Makefile.win clean all
3. (optional) To build shared library libsvm.dll, type
nmake -f Makefile.win lib
Another way is to build them from Visual C++ environment. See details
in libsvm FAQ.
- Additional Tools: Sub-sampling, Parameter Selection, Format checking, etc.
============================================================================
See the README file in the tools directory.
MATLAB/OCTAVE Interface
=======================
Please check the file README in the directory `matlab'.
Python Interface
================
See the README file in python directory.
Additional Information
======================
If you find LIBSVM helpful, please cite it as
Chih-Chung Chang and Chih-Jen Lin, LIBSVM : a library for support
vector machines. ACM Transactions on Intelligent Systems and
Technology, 2:27:1--27:27, 2011. Software available at
http://www.csie.ntu.edu.tw/~cjlin/libsvm
LIBSVM implementation document is available at
http://www.csie.ntu.edu.tw/~cjlin/papers/libsvm.pdf
For any questions and comments, please email cjlin@csie.ntu.edu.tw
Acknowledgments:
This work was supported in part by the National Science
Council of Taiwan via the grant NSC 89-2213-E-002-013.
The authors thank their group members and users
for many helpful discussions and comments. They are listed in
http://www.csie.ntu.edu.tw/~cjlin/libsvm/acknowledgements
================================================
FILE: binaries/linux/README-GPU
================================================
GPU-Accelerated LIBSVM is exploiting the GPU, using the CUDA interface, to
speed-up the training process. This package contains a new executable for
training classifiers "svm-train-gpu.exe" together with the original one.
The use of the new executable is exactly the same as with the original one.
This binary was built with the CUBLAS API version 2 which is compatible with SDKs from 4.0 and up.
To test the binary "svm-train-gpu" you can run the easy.py script which is located in the "tools" folder.
To observe speed improvements between CPU and GPU execution we provide a custom relatively large dataset (train_set) which can be used as an input to easy.py.
FEATURES
Mode Supported
* c-svc classification with RBF kernel
Functionality / User interface
* Same as LIBSVM
PREREQUISITES
* NVIDIA Graphics card with CUDA support
* Latest NVIDIA drivers for GPU
Additional Information
======================
If you find GPU-Accelerated LIBSVM helpful, please cite it as
A. Athanasopoulos, A. Dimou, V. Mezaris, I. Kompatsiaris, "GPU Acceleration for Support Vector Machines",
Proc. 12th International Workshop on Image Analysis for Multimedia Interactive Services (WIAMIS 2011), Delft, The Netherlands, April 2011.
Software available at http://mklab.iti.gr/project/GPU-LIBSVM
================================================
FILE: binaries/linux/tools/README
================================================
This directory includes some useful codes:
1. subset selection tools.
2. parameter selection tools.
3. LIBSVM format checking tools
Part I: Subset selection tools
Introduction
============
Training large data is time consuming. Sometimes one should work on a
smaller subset first. The python script subset.py randomly selects a
specified number of samples. For classification data, we provide a
stratified selection to ensure the same class distribution in the
subset.
Usage: subset.py [options] dataset number [output1] [output2]
This script selects a subset of the given data set.
options:
-s method : method of selection (default 0)
0 -- stratified selection (classification only)
1 -- random selection
output1 : the subset (optional)
output2 : the rest of data (optional)
If output1 is omitted, the subset will be printed on the screen.
Example
=======
> python subset.py heart_scale 100 file1 file2
From heart_scale 100 samples are randomly selected and stored in
file1. All remaining instances are stored in file2.
Part II: Parameter Selection Tools
Introduction
============
grid.py is a parameter selection tool for C-SVM classification using
the RBF (radial basis function) kernel. It uses cross validation (CV)
technique to estimate the accuracy of each parameter combination in
the specified range and helps you to decide the best parameters for
your problem.
grid.py directly executes libsvm binaries (so no python binding is needed)
for cross validation and then draw contour of CV accuracy using gnuplot.
You must have libsvm and gnuplot installed before using it. The package
gnuplot is available at http://www.gnuplot.info/
On Mac OSX, the precompiled gnuplot file needs the library Aquarterm,
which thus must be installed as well. In addition, this version of
gnuplot does not support png, so you need to change "set term png
transparent small" and use other image formats. For example, you may
have "set term pbm small color".
Usage: grid.py [grid_options] [svm_options] dataset
grid_options :
-log2c {begin,end,step | "null"} : set the range of c (default -5,15,2)
begin,end,step -- c_range = 2^{begin,...,begin+k*step,...,end}
"null" -- do not grid with c
-log2g {begin,end,step | "null"} : set the range of g (default 3,-15,-2)
begin,end,step -- g_range = 2^{begin,...,begin+k*step,...,end}
"null" -- do not grid with g
-v n : n-fold cross validation (default 5)
-svmtrain pathname : set svm executable path and name
-gnuplot {pathname | "null"} :
pathname -- set gnuplot executable path and name
"null" -- do not plot
-out {pathname | "null"} : (default dataset.out)
pathname -- set output file path and name
"null" -- do not output file
-png pathname : set graphic output file path and name (default dataset.png)
-resume [pathname] : resume the grid task using an existing output file (default pathname is dataset.out)
Use this option only if some parameters have been checked for the SAME data.
svm_options : additional options for svm-train
The program conducts v-fold cross validation using parameter C (and gamma)
= 2^begin, 2^(begin+step), ..., 2^end.
You can specify where the libsvm executable and gnuplot are using the
-svmtrain and -gnuplot parameters.
For windows users, please use pgnuplot.exe. If you are using gnuplot
3.7.1, please upgrade to version 3.7.3 or higher. The version 3.7.1
has a bug. If you use cygwin on windows, please use gunplot-x11.
If the task is terminated accidentally or you would like to change the
range of parameters, you can apply '-resume' to save time by re-using
previous results. You may specify the output file of a previous run
or use the default (i.e., dataset.out) without giving a name. Please
note that the same condition must be used in two runs. For example,
you cannot use '-v 10' earlier and resume the task with '-v 5'.
The value of some options can be "null." For example, `-log2c -1,0,1
-log2 "null"' means that C=2^-1,2^0,2^1 and g=LIBSVM's default gamma
value. That is, you do not conduct parameter selection on gamma.
Example
=======
> python grid.py -log2c -5,5,1 -log2g -4,0,1 -v 5 -m 300 heart_scale
Users (in particular MS Windows users) may need to specify the path of
executable files. You can either change paths in the beginning of
grid.py or specify them in the command line. For example,
> grid.py -log2c -5,5,1 -svmtrain "c:\Program Files\libsvm\windows\svm-train.exe" -gnuplot c:\tmp\gnuplot\binary\pgnuplot.exe -v 10 heart_scale
Output: two files
dataset.png: the CV accuracy contour plot generated by gnuplot
dataset.out: the CV accuracy at each (log2(C),log2(gamma))
The following example saves running time by loading the output file of a previous run.
> python grid.py -log2c -7,7,1 -log2g -5,2,1 -v 5 -resume heart_scale.out heart_scale
Parallel grid search
====================
You can conduct a parallel grid search by dispatching jobs to a
cluster of computers which share the same file system. First, you add
machine names in grid.py:
ssh_workers = ["linux1", "linux5", "linux5"]
and then setup your ssh so that the authentication works without
asking a password.
The same machine (e.g., linux5 here) can be listed more than once if
it has multiple CPUs or has more RAM. If the local machine is the
best, you can also enlarge the nr_local_worker. For example:
nr_local_worker = 2
Example:
> python grid.py heart_scale
[local] -1 -1 78.8889 (best c=0.5, g=0.5, rate=78.8889)
[linux5] -1 -7 83.3333 (best c=0.5, g=0.0078125, rate=83.3333)
[linux5] 5 -1 77.037 (best c=0.5, g=0.0078125, rate=83.3333)
[linux1] 5 -7 83.3333 (best c=0.5, g=0.0078125, rate=83.3333)
.
.
.
If -log2c, -log2g, or -v is not specified, default values are used.
If your system uses telnet instead of ssh, you list the computer names
in telnet_workers.
Calling grid in Python
======================
In addition to using grid.py as a command-line tool, you can use it as a
Python module.
>>> rate, param = find_parameters(dataset, options)
You need to specify `dataset' and `options' (default ''). See the following example.
> python
>>> from grid import *
>>> rate, param = find_parameters('../heart_scale', '-log2c -1,1,1 -log2g -1,1,1')
[local] 0.0 0.0 rate=74.8148 (best c=1.0, g=1.0, rate=74.8148)
[local] 0.0 -1.0 rate=77.037 (best c=1.0, g=0.5, rate=77.037)
.
.
[local] -1.0 -1.0 rate=78.8889 (best c=0.5, g=0.5, rate=78.8889)
.
.
>>> rate
78.8889
>>> param
{'c': 0.5, 'g': 0.5}
Part III: LIBSVM format checking tools
Introduction
============
`svm-train' conducts only a simple check of the input data. To do a
detailed check, we provide a python script `checkdata.py.'
Usage: checkdata.py dataset
Exit status (returned value): 1 if there are errors, 0 otherwise.
This tool is written by Rong-En Fan at National Taiwan University.
Example
=======
> cat bad_data
1 3:1 2:4
> python checkdata.py bad_data
line 1: feature indices must be in an ascending order, previous/current features 3:1 2:4
Found 1 lines with error.
================================================
FILE: binaries/linux/tools/checkdata.py
================================================
#!/usr/bin/env python
#
# A format checker for LIBSVM
#
#
# Copyright (c) 2007, Rong-En Fan
#
# All rights reserved.
#
# This program is distributed under the same license of the LIBSVM package.
#
from sys import argv, exit
import os.path
def err(line_no, msg):
print("line {0}: {1}".format(line_no, msg))
# works like float() but does not accept nan and inf
def my_float(x):
if x.lower().find("nan") != -1 or x.lower().find("inf") != -1:
raise ValueError
return float(x)
def main():
if len(argv) != 2:
print("Usage: {0} dataset".format(argv[0]))
exit(1)
dataset = argv[1]
if not os.path.exists(dataset):
print("dataset {0} not found".format(dataset))
exit(1)
line_no = 1
error_line_count = 0
for line in open(dataset, 'r'):
line_error = False
# each line must end with a newline character
if line[-1] != '\n':
err(line_no, "missing a newline character in the end")
line_error = True
nodes = line.split()
# check label
try:
label = nodes.pop(0)
if label.find(',') != -1:
# multi-label format
try:
for l in label.split(','):
l = my_float(l)
except:
err(line_no, "label {0} is not a valid multi-label form".format(label))
line_error = True
else:
try:
label = my_float(label)
except:
err(line_no, "label {0} is not a number".format(label))
line_error = True
except:
err(line_no, "missing label, perhaps an empty line?")
line_error = True
# check features
prev_index = -1
for i in range(len(nodes)):
try:
(index, value) = nodes[i].split(':')
index = int(index)
value = my_float(value)
# precomputed kernel's index starts from 0 and LIBSVM
# checks it. Hence, don't treat index 0 as an error.
if index < 0:
err(line_no, "feature index must be positive; wrong feature {0}".format(nodes[i]))
line_error = True
elif index <= prev_index:
err(line_no, "feature indices must be in an ascending order, previous/current features {0} {1}".format(nodes[i-1], nodes[i]))
line_error = True
prev_index = index
except:
err(line_no, "feature '{0}' not an <index>:<value> pair, <index> integer, <value> real number ".format(nodes[i]))
line_error = True
line_no += 1
if line_error:
error_line_count += 1
if error_line_count > 0:
print("Found {0} lines with error.".format(error_line_count))
return 1
else:
print("No error.")
return 0
if __name__ == "__main__":
exit(main())
================================================
FILE: binaries/linux/tools/easy.py
================================================
#!/usr/bin/env python
import sys
import os
from subprocess import *
if len(sys.argv) <= 1:
print('Usage: {0} training_file [testing_file]'.format(sys.argv[0]))
raise SystemExit
# svm, grid, and gnuplot executable files
is_win32 = (sys.platform == 'win32')
if not is_win32:
svmscale_exe = "../svm-scale"
svmtrain_exe = "../svm-train-gpu"
svmpredict_exe = "../svm-predict"
grid_py = "./grid.py"
gnuplot_exe = "/usr/bin/gnuplot"
else:
# example for windows
svmscale_exe = r"..\windows\svm-scale.exe"
svmtrain_exe = r"..\windows\svm-train-gpu.exe"
svmpredict_exe = r"..\windows\svm-predict.exe"
gnuplot_exe = r"c:\tmp\gnuplot\binary\pgnuplot.exe"
grid_py = r".\grid.py"
assert os.path.exists(svmscale_exe),"svm-scale executable not found"
assert os.path.exists(svmtrain_exe),"svm-train executable not found"
assert os.path.exists(svmpredict_exe),"svm-predict executable not found"
assert os.path.exists(gnuplot_exe),"gnuplot executable not found"
assert os.path.exists(grid_py),"grid.py not found"
train_pathname = sys.argv[1]
assert os.path.exists(train_pathname),"training file not found"
file_name = os.path.split(train_pathname)[1]
scaled_file = file_name + ".scale"
model_file = file_name + ".model"
range_file = file_name + ".range"
if len(sys.argv) > 2:
test_pathname = sys.argv[2]
file_name = os.path.split(test_pathname)[1]
assert os.path.exists(test_pathname),"testing file not found"
scaled_test_file = file_name + ".scale"
predict_test_file = file_name + ".predict"
cmd = '{0} -s "{1}" "{2}" > "{3}"'.format(svmscale_exe, range_file, train_pathname, scaled_file)
print('Scaling training data...')
Popen(cmd, shell = True, stdout = PIPE).communicate()
cmd = '{0} -svmtrain "{1}" -gnuplot "{2}" "{3}"'.format(grid_py, svmtrain_exe, gnuplot_exe, scaled_file)
print('Cross validation...')
f = Popen(cmd, shell = True, stdout = PIPE).stdout
line = ''
while True:
last_line = line
line = f.readline()
if not line: break
c,g,rate = map(float,last_line.split())
print('Best c={0}, g={1} CV rate={2}'.format(c,g,rate))
cmd = '{0} -c {1} -g {2} "{3}" "{4}"'.format(svmtrain_exe,c,g,scaled_file,model_file)
print('Training...')
Popen(cmd, shell = True, stdout = PIPE).communicate()
print('Output model: {0}'.format(model_file))
if len(sys.argv) > 2:
cmd = '{0} -r "{1}" "{2}" > "{3}"'.format(svmscale_exe, range_file, test_pathname, scaled_test_file)
print('Scaling testing data...')
Popen(cmd, shell = True, stdout = PIPE).communicate()
cmd = '{0} "{1}" "{2}" "{3}"'.format(svmpredict_exe, scaled_test_file, model_file, predict_test_file)
print('Testing...')
Popen(cmd, shell = True).communicate()
print('Output prediction: {0}'.format(predict_test_file))
================================================
FILE: binaries/linux/tools/grid.py
================================================
#!/usr/bin/env python
__all__ = ['find_parameters']
import os, sys, traceback, getpass, time, re
from threading import Thread
from subprocess import *
if sys.version_info[0] < 3:
from Queue import Queue
else:
from queue import Queue
telnet_workers = []
ssh_workers = []
nr_local_worker = 1
class GridOption:
def __init__(self, dataset_pathname, options):
dirname = os.path.dirname(__file__)
if sys.platform != 'win32':
self.svmtrain_pathname = os.path.join(dirname, '../svm-train-gpu')
self.gnuplot_pathname = '/usr/bin/gnuplot'
else:
# example for windows
self.svmtrain_pathname = os.path.join(dirname, r'..\windows\svm-train-gpu.exe')
# svmtrain_pathname = r'c:\Program Files\libsvm\windows\svm-train-gpu.exe'
self.gnuplot_pathname = r'c:\tmp\gnuplot\binary\pgnuplot.exe'
self.fold = 5
self.c_begin, self.c_end, self.c_step = -5, 15, 2
self.g_begin, self.g_end, self.g_step = 3, -15, -2
self.grid_with_c, self.grid_with_g = True, True
self.dataset_pathname = dataset_pathname
self.dataset_title = os.path.split(dataset_pathname)[1]
self.out_pathname = '{0}.out'.format(self.dataset_title)
self.png_pathname = '{0}.png'.format(self.dataset_title)
self.pass_through_string = ' '
self.resume_pathname = None
self.parse_options(options)
def parse_options(self, options):
if type(options) == str:
options = options.split()
i = 0
pass_through_options = []
while i < len(options):
if options[i] == '-log2c':
i = i + 1
if options[i] == 'null':
self.grid_with_c = False
else:
self.c_begin, self.c_end, self.c_step = map(float,options[i].split(','))
elif options[i] == '-log2g':
i = i + 1
if options[i] == 'null':
self.grid_with_g = False
else:
self.g_begin, self.g_end, self.g_step = map(float,options[i].split(','))
elif options[i] == '-v':
i = i + 1
self.fold = options[i]
elif options[i] in ('-c','-g'):
raise ValueError('Use -log2c and -log2g.')
elif options[i] == '-svmtrain':
i = i + 1
self.svmtrain_pathname = options[i]
elif options[i] == '-gnuplot':
i = i + 1
if options[i] == 'null':
self.gnuplot_pathname = None
else:
self.gnuplot_pathname = options[i]
elif options[i] == '-out':
i = i + 1
if options[i] == 'null':
self.out_pathname = None
else:
self.out_pathname = options[i]
elif options[i] == '-png':
i = i + 1
self.png_pathname = options[i]
elif options[i] == '-resume':
if i == (len(options)-1) or options[i+1].startswith('-'):
self.resume_pathname = self.dataset_title + '.out'
else:
i = i + 1
self.resume_pathname = options[i]
else:
pass_through_options.append(options[i])
i = i + 1
self.pass_through_string = ' '.join(pass_through_options)
if not os.path.exists(self.svmtrain_pathname):
raise IOError('svm-train executable not found')
if not os.path.exists(self.dataset_pathname):
raise IOError('dataset not found')
if self.resume_pathname and not os.path.exists(self.resume_pathname):
raise IOError('file for resumption not found')
if not self.grid_with_c and not self.grid_with_g:
raise ValueError('-log2c and -log2g should not be null simultaneously')
if self.gnuplot_pathname and not os.path.exists(self.gnuplot_pathname):
sys.stderr.write('gnuplot executable not found\n')
self.gnuplot_pathname = None
def redraw(db,best_param,gnuplot,options,tofile=False):
if len(db) == 0: return
begin_level = round(max(x[2] for x in db)) - 3
step_size = 0.5
best_log2c,best_log2g,best_rate = best_param
# if newly obtained c, g, or cv values are the same,
# then stop redrawing the contour.
if all(x[0] == db[0][0] for x in db): return
if all(x[1] == db[0][1] for x in db): return
if all(x[2] == db[0][2] for x in db): return
if tofile:
gnuplot.write(b"set term png transparent small linewidth 2 medium enhanced\n")
gnuplot.write("set output \"{0}\"\n".format(options.png_pathname.replace('\\','\\\\')).encode())
#gnuplot.write(b"set term postscript color solid\n")
#gnuplot.write("set output \"{0}.ps\"\n".format(options.dataset_title).encode().encode())
elif sys.platform == 'win32':
gnuplot.write(b"set term windows\n")
else:
gnuplot.write( b"set term x11\n")
gnuplot.write(b"set xlabel \"log2(C)\"\n")
gnuplot.write(b"set ylabel \"log2(gamma)\"\n")
gnuplot.write("set xrange [{0}:{1}]\n".format(options.c_begin,options.c_end).encode())
gnuplot.write("set yrange [{0}:{1}]\n".format(options.g_begin,options.g_end).encode())
gnuplot.write(b"set contour\n")
gnuplot.write("set cntrparam levels incremental {0},{1},100\n".format(begin_level,step_size).encode())
gnuplot.write(b"unset surface\n")
gnuplot.write(b"unset ztics\n")
gnuplot.write(b"set view 0,0\n")
gnuplot.write("set title \"{0}\"\n".format(options.dataset_title).encode())
gnuplot.write(b"unset label\n")
gnuplot.write("set label \"Best log2(C) = {0} log2(gamma) = {1} accuracy = {2}%\" \
at screen 0.5,0.85 center\n". \
format(best_log2c, best_log2g, best_rate).encode())
gnuplot.write("set label \"C = {0} gamma = {1}\""
" at screen 0.5,0.8 center\n".format(2**best_log2c, 2**best_log2g).encode())
gnuplot.write(b"set key at screen 0.9,0.9\n")
gnuplot.write(b"splot \"-\" with lines\n")
db.sort(key = lambda x:(x[0], -x[1]))
prevc = db[0][0]
for line in db:
if prevc != line[0]:
gnuplot.write(b"\n")
prevc = line[0]
gnuplot.write("{0[0]} {0[1]} {0[2]}\n".format(line).encode())
gnuplot.write(b"e\n")
gnuplot.write(b"\n") # force gnuplot back to prompt when term set failure
gnuplot.flush()
def calculate_jobs(options):
def range_f(begin,end,step):
# like range, but works on non-integer too
seq = []
while True:
if step > 0 and begin > end: break
if step < 0 and begin < end: break
seq.append(begin)
begin = begin + step
return seq
def permute_sequence(seq):
n = len(seq)
if n <= 1: return seq
mid = int(n/2)
left = permute_sequence(seq[:mid])
right = permute_sequence(seq[mid+1:])
ret = [seq[mid]]
while left or right:
if left: ret.append(left.pop(0))
if right: ret.append(right.pop(0))
return ret
c_seq = permute_sequence(range_f(options.c_begin,options.c_end,options.c_step))
g_seq = permute_sequence(range_f(options.g_begin,options.g_end,options.g_step))
if not options.grid_with_c:
c_seq = [None]
if not options.grid_with_g:
g_seq = [None]
nr_c = float(len(c_seq))
nr_g = float(len(g_seq))
i, j = 0, 0
jobs = []
while i < nr_c or j < nr_g:
if i/nr_c < j/nr_g:
# increase C resolution
line = []
for k in range(0,j):
line.append((c_seq[i],g_seq[k]))
i = i + 1
jobs.append(line)
else:
# increase g resolution
line = []
for k in range(0,i):
line.append((c_seq[k],g_seq[j]))
j = j + 1
jobs.append(line)
resumed_jobs = {}
if options.resume_pathname is None:
return jobs, resumed_jobs
for line in open(options.resume_pathname, 'r'):
line = line.strip()
rst = re.findall(r'rate=([0-9.]+)',line)
if not rst:
continue
rate = float(rst[0])
c, g = None, None
rst = re.findall(r'log2c=([0-9.-]+)',line)
if rst:
c = float(rst[0])
rst = re.findall(r'log2g=([0-9.-]+)',line)
if rst:
g = float(rst[0])
resumed_jobs[(c,g)] = rate
return jobs, resumed_jobs
class WorkerStopToken: # used to notify the worker to stop or if a worker is dead
pass
class Worker(Thread):
def __init__(self,name,job_queue,result_queue,options):
Thread.__init__(self)
self.name = name
self.job_queue = job_queue
self.result_queue = result_queue
self.options = options
def run(self):
while True:
(cexp,gexp) = self.job_queue.get()
if cexp is WorkerStopToken:
self.job_queue.put((cexp,gexp))
# print('worker {0} stop.'.format(self.name))
break
try:
c, g = None, None
if cexp != None:
c = 2.0**cexp
if gexp != None:
g = 2.0**gexp
rate = self.run_one(c,g)
if rate is None: raise RuntimeError('get no rate')
except:
# we failed, let others do that and we just quit
traceback.print_exception(sys.exc_info()[0], sys.exc_info()[1], sys.exc_info()[2])
self.job_queue.put((cexp,gexp))
sys.stderr.write('worker {0} quit.\n'.format(self.name))
break
else:
self.result_queue.put((self.name,cexp,gexp,rate))
def get_cmd(self,c,g):
options=self.options
cmdline = options.svmtrain_pathname
if options.grid_with_c:
cmdline += ' -c {0} '.format(c)
if options.grid_with_g:
cmdline += ' -g {0} '.format(g)
cmdline += ' -v {0} {1} {2} '.format\
(options.fold,options.pass_through_string,options.dataset_pathname)
return cmdline
class LocalWorker(Worker):
def run_one(self,c,g):
cmdline = self.get_cmd(c,g)
result = Popen(cmdline,shell=True,stdout=PIPE,stderr=PIPE,stdin=PIPE).stdout
for line in result.readlines():
if str(line).find('Cross') != -1:
return float(line.split()[-1][0:-1])
class SSHWorker(Worker):
def __init__(self,name,job_queue,result_queue,host,options):
Worker.__init__(self,name,job_queue,result_queue,options)
self.host = host
self.cwd = os.getcwd()
def run_one(self,c,g):
cmdline = 'ssh -x -t -t {0} "cd {1}; {2}"'.format\
(self.host,self.cwd,self.get_cmd(c,g))
result = Popen(cmdline,shell=True,stdout=PIPE,stderr=PIPE,stdin=PIPE).stdout
for line in result.readlines():
if str(line).find('Cross') != -1:
return float(line.split()[-1][0:-1])
class TelnetWorker(Worker):
def __init__(self,name,job_queue,result_queue,host,username,password,options):
Worker.__init__(self,name,job_queue,result_queue,options)
self.host = host
self.username = username
self.password = password
def run(self):
import telnetlib
self.tn = tn = telnetlib.Telnet(self.host)
tn.read_until('login: ')
tn.write(self.username + '\n')
tn.read_until('Password: ')
tn.write(self.password + '\n')
# XXX: how to know whether login is successful?
tn.read_until(self.username)
#
print('login ok', self.host)
tn.write('cd '+os.getcwd()+'\n')
Worker.run(self)
tn.write('exit\n')
def run_one(self,c,g):
cmdline = self.get_cmd(c,g)
result = self.tn.write(cmdline+'\n')
(idx,matchm,output) = self.tn.expect(['Cross.*\n'])
for line in output.split('\n'):
if str(line).find('Cross') != -1:
return float(line.split()[-1][0:-1])
def find_parameters(dataset_pathname, options=''):
def update_param(c,g,rate,best_c,best_g,best_rate,worker,resumed):
if (rate > best_rate) or (rate==best_rate and g==best_g and c<best_c):
best_rate,best_c,best_g = rate,c,g
stdout_str = '[{0}] {1} {2} (best '.format\
(worker,' '.join(str(x) for x in [c,g] if x is not None),rate)
output_str = ''
if c != None:
stdout_str += 'c={0}, '.format(2.0**best_c)
output_str += 'log2c={0} '.format(c)
if g != None:
stdout_str += 'g={0}, '.format(2.0**best_g)
output_str += 'log2g={0} '.format(g)
stdout_str += 'rate={0})'.format(best_rate)
print(stdout_str)
if options.out_pathname and not resumed:
output_str += 'rate={0}\n'.format(rate)
result_file.write(output_str)
result_file.flush()
return best_c,best_g,best_rate
options = GridOption(dataset_pathname, options);
if options.gnuplot_pathname:
gnuplot = Popen(options.gnuplot_pathname,stdin = PIPE,stdout=PIPE,stderr=PIPE).stdin
else:
gnuplot = None
# put jobs in queue
jobs,resumed_jobs = calculate_jobs(options)
job_queue = Queue(0)
result_queue = Queue(0)
for (c,g) in resumed_jobs:
result_queue.put(('resumed',c,g,resumed_jobs[(c,g)]))
for line in jobs:
for (c,g) in line:
if (c,g) not in resumed_jobs:
job_queue.put((c,g))
# hack the queue to become a stack --
# this is important when some thread
# failed and re-put a job. It we still
# use FIFO, the job will be put
# into the end of the queue, and the graph
# will only be updated in the end
job_queue._put = job_queue.queue.appendleft
# fire telnet workers
if telnet_workers:
nr_telnet_worker = len(telnet_workers)
username = getpass.getuser()
password = getpass.getpass()
for host in telnet_workers:
worker = TelnetWorker(host,job_queue,result_queue,
host,username,password,options)
worker.start()
# fire ssh workers
if ssh_workers:
for host in ssh_workers:
worker = SSHWorker(host,job_queue,result_queue,host,options)
worker.start()
# fire local workers
for i in range(nr_local_worker):
worker = LocalWorker('local',job_queue,result_queue,options)
worker.start()
# gather results
done_jobs = {}
if options.out_pathname:
if options.resume_pathname:
result_file = open(options.out_pathname, 'a')
else:
result_file = open(options.out_pathname, 'w')
db = []
best_rate = -1
best_c,best_g = None,None
for (c,g) in resumed_jobs:
rate = resumed_jobs[(c,g)]
best_c,best_g,best_rate = update_param(c,g,rate,best_c,best_g,best_rate,'resumed',True)
for line in jobs:
for (c,g) in line:
while (c,g) not in done_jobs:
(worker,c1,g1,rate1) = result_queue.get()
done_jobs[(c1,g1)] = rate1
if (c1,g1) not in resumed_jobs:
best_c,best_g,best_rate = update_param(c1,g1,rate1,best_c,best_g,best_rate,worker,False)
db.append((c,g,done_jobs[(c,g)]))
if gnuplot and options.grid_with_c and options.grid_with_g:
redraw(db,[best_c, best_g, best_rate],gnuplot,options)
redraw(db,[best_c, best_g, best_rate],gnuplot,options,True)
if options.out_pathname:
result_file.close()
job_queue.put((WorkerStopToken,None))
best_param, best_cg = {}, []
if best_c != None:
best_param['c'] = 2.0**best_c
best_cg += [2.0**best_c]
if best_g != None:
best_param['g'] = 2.0**best_g
best_cg += [2.0**best_g]
print('{0} {1}'.format(' '.join(map(str,best_cg)), best_rate))
return best_rate, best_param
if __name__ == '__main__':
def exit_with_help():
print("""\
Usage: grid.py [grid_options] [svm_options] dataset
grid_options :
-log2c {begin,end,step | "null"} : set the range of c (default -5,15,2)
begin,end,step -- c_range = 2^{begin,...,begin+k*step,...,end}
"null" -- do not grid with c
-log2g {begin,end,step | "null"} : set the range of g (default 3,-15,-2)
begin,end,step -- g_range = 2^{begin,...,begin+k*step,...,end}
"null" -- do not grid with g
-v n : n-fold cross validation (default 5)
-svmtrain pathname : set svm executable path and name
-gnuplot {pathname | "null"} :
pathname -- set gnuplot executable path and name
"null" -- do not plot
-out {pathname | "null"} : (default dataset.out)
pathname -- set output file path and name
"null" -- do not output file
-png pathname : set graphic output file path and name (default dataset.png)
-resume [pathname] : resume the grid task using an existing output file (default pathname is dataset.out)
This is experimental. Try this option only if some parameters have been checked for the SAME data.
svm_options : additional options for svm-train""")
sys.exit(1)
if len(sys.argv) < 2:
exit_with_help()
dataset_pathname = sys.argv[-1]
options = sys.argv[1:-1]
try:
find_parameters(dataset_pathname, options)
except (IOError,ValueError) as e:
sys.stderr.write(str(e) + '\n')
sys.stderr.write('Try "grid.py" for more information.\n')
sys.exit(1)
================================================
FILE: binaries/linux/tools/subset.py
================================================
#!/usr/bin/env python
import os, sys, math, random
from collections import defaultdict
if sys.version_info[0] >= 3:
xrange = range
def exit_with_help(argv):
print("""\
Usage: {0} [options] dataset subset_size [output1] [output2]
This script randomly selects a subset of the dataset.
options:
-s method : method of selection (default 0)
0 -- stratified selection (classification only)
1 -- random selection
output1 : the subset (optional)
output2 : rest of the data (optional)
If output1 is omitted, the subset will be printed on the screen.""".format(argv[0]))
exit(1)
def process_options(argv):
argc = len(argv)
if argc < 3:
exit_with_help(argv)
# default method is stratified selection
method = 0
subset_file = sys.stdout
rest_file = None
i = 1
while i < argc:
if argv[i][0] != "-":
break
if argv[i] == "-s":
i = i + 1
method = int(argv[i])
if method not in [0,1]:
print("Unknown selection method {0}".format(method))
exit_with_help(argv)
i = i + 1
dataset = argv[i]
subset_size = int(argv[i+1])
if i+2 < argc:
subset_file = open(argv[i+2],'w')
if i+3 < argc:
rest_file = open(argv[i+3],'w')
return dataset, subset_size, method, subset_file, rest_file
def random_selection(dataset, subset_size):
l = sum(1 for line in open(dataset,'r'))
return sorted(random.sample(xrange(l), subset_size))
def stratified_selection(dataset, subset_size):
labels = [line.split(None,1)[0] for line in open(dataset)]
label_linenums = defaultdict(list)
for i, label in enumerate(labels):
label_linenums[label] += [i]
l = len(labels)
remaining = subset_size
ret = []
# classes with fewer data are sampled first; otherwise
# some rare classes may not be selected
for label in sorted(label_linenums, key=lambda x: len(label_linenums[x])):
linenums = label_linenums[label]
label_size = len(linenums)
# at least one instance per class
s = int(min(remaining, max(1, math.ceil(label_size*(float(subset_size)/l)))))
if s == 0:
sys.stderr.write('''\
Error: failed to have at least one instance per class
1. You may have regression data.
2. Your classification data is unbalanced or too small.
Please use -s 1.
''')
sys.exit(-1)
remaining -= s
ret += [linenums[i] for i in random.sample(xrange(label_size), s)]
return sorted(ret)
def main(argv=sys.argv):
dataset, subset_size, method, subset_file, rest_file = process_options(argv)
#uncomment the following line to fix the random seed
#random.seed(0)
selected_lines = []
if method == 0:
selected_lines = stratified_selection(dataset, subset_size)
elif method == 1:
selected_lines = random_selection(dataset, subset_size)
#select instances based on selected_lines
dataset = open(dataset,'r')
prev_selected_linenum = -1
for i in xrange(len(selected_lines)):
for cnt in xrange(selected_lines[i]-prev_selected_linenum-1):
line = dataset.readline()
if rest_file:
rest_file.write(line)
subset_file.write(dataset.readline())
prev_selected_linenum = selected_lines[i]
subset_file.close()
if rest_file:
for line in dataset:
rest_file.write(line)
rest_file.close()
dataset.close()
if __name__ == '__main__':
main(sys.argv)
================================================
FILE: binaries/linux/train_set
================================================
[File too large to display: 10.8 MB]
================================================
FILE: binaries/windows/x64/COPYRIGHT
================================================
Copyright (c) 2000-2013 Chih-Chung Chang and Chih-Jen Lin
All rights reserved.
Redistribution and use in source and binary forms, with or without
modification, are permitted provided that the following conditions
are met:
1. Redistributions of source code must retain the above copyright
notice, this list of conditions and the following disclaimer.
2. Redistributions in binary form must reproduce the above copyright
notice, this list of conditions and the following disclaimer in the
documentation and/or other materials provided with the distribution.
3. Neither name of copyright holders nor the names of its contributors
may be used to endorse or promote products derived from this software
without specific prior written permission.
THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS
``AS IS'' AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT
LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR
A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE REGENTS OR
CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL,
EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO,
PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR
PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF
LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING
NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS
SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
================================================
FILE: binaries/windows/x64/FAQ.html
================================================
<html>
<head>
<title>LIBSVM FAQ</title>
</head>
<body bgcolor="#ffffcc">
<a name="_TOP"><b><h1><a
href=http://www.csie.ntu.edu.tw/~cjlin/libsvm>LIBSVM</a> FAQ </h1></b></a>
<b>last modified : </b>
Wed, 19 Dec 2012 13:26:34 GMT
<class="categories">
<li><a
href="#_TOP">All Questions</a>(78)</li>
<ul><b>
<li><a
href="#/Q1:_Some_sample_uses_of_libsvm">Q1:_Some_sample_uses_of_libsvm</a>(2)</li>
<li><a
href="#/Q2:_Installation_and_running_the_program">Q2:_Installation_and_running_the_program</a>(13)</li>
<li><a
href="#/Q3:_Data_preparation">Q3:_Data_preparation</a>(7)</li>
<li><a
href="#/Q4:_Training_and_prediction">Q4:_Training_and_prediction</a>(34)</li>
<li><a
href="#/Q5:_Probability_outputs">Q5:_Probability_outputs</a>(3)</li>
<li><a
href="#/Q6:_Graphic_interface">Q6:_Graphic_interface</a>(3)</li>
<li><a
href="#/Q7:_Java_version_of_libsvm">Q7:_Java_version_of_libsvm</a>(4)</li>
<li><a
href="#/Q8:_Python_interface">Q8:_Python_interface</a>(1)</li>
<li><a
href="#/Q9:_MATLAB_interface">Q9:_MATLAB_interface</a>(11)</li>
</b></ul>
</li>
<ul><ul class="headlines">
<li class="headlines_item"><a href="#faq101">Some courses which have used libsvm as a tool</a></li>
<li class="headlines_item"><a href="#faq102">Some applications/tools which have used libsvm </a></li>
<li class="headlines_item"><a href="#f201">Where can I find documents/videos of libsvm ?</a></li>
<li class="headlines_item"><a href="#f202">Where are change log and earlier versions?</a></li>
<li class="headlines_item"><a href="#f203">How to cite LIBSVM?</a></li>
<li class="headlines_item"><a href="#f204">I would like to use libsvm in my software. Is there any license problem?</a></li>
<li class="headlines_item"><a href="#f205">Is there a repository of additional tools based on libsvm?</a></li>
<li class="headlines_item"><a href="#f206">On unix machines, I got "error in loading shared libraries" or "cannot open shared object file." What happened ? </a></li>
<li class="headlines_item"><a href="#f207">I have modified the source and would like to build the graphic interface "svm-toy" on MS windows. How should I do it ?</a></li>
<li class="headlines_item"><a href="#f208">I am an MS windows user but why only one (svm-toy) of those precompiled .exe actually runs ? </a></li>
<li class="headlines_item"><a href="#f209">What is the difference between "." and "*" outputed during training? </a></li>
<li class="headlines_item"><a href="#f210">Why occasionally the program (including MATLAB or other interfaces) crashes and gives a segmentation fault?</a></li>
<li class="headlines_item"><a href="#f211">How to build a dynamic library (.dll file) on MS windows?</a></li>
<li class="headlines_item"><a href="#f212">On some systems (e.g., Ubuntu), compiling LIBSVM gives many warning messages. Is this a problem and how to disable the warning message?</a></li>
<li class="headlines_item"><a href="#f213">In LIBSVM, why you don't use certain C/C++ library functions to make the code shorter?</a></li>
<li class="headlines_item"><a href="#f301">Why sometimes not all attributes of a data appear in the training/model files ?</a></li>
<li class="headlines_item"><a href="#f302">What if my data are non-numerical ?</a></li>
<li class="headlines_item"><a href="#f303">Why do you consider sparse format ? Will the training of dense data be much slower ?</a></li>
<li class="headlines_item"><a href="#f304">Why sometimes the last line of my data is not read by svm-train?</a></li>
<li class="headlines_item"><a href="#f305">Is there a program to check if my data are in the correct format?</a></li>
<li class="headlines_item"><a href="#f306">May I put comments in data files?</a></li>
<li class="headlines_item"><a href="#f307">How to convert other data formats to LIBSVM format?</a></li>
<li class="headlines_item"><a href="#f401">The output of training C-SVM is like the following. What do they mean?</a></li>
<li class="headlines_item"><a href="#f402">Can you explain more about the model file?</a></li>
<li class="headlines_item"><a href="#f403">Should I use float or double to store numbers in the cache ?</a></li>
<li class="headlines_item"><a href="#f404">How do I choose the kernel?</a></li>
<li class="headlines_item"><a href="#f405">Does libsvm have special treatments for linear SVM?</a></li>
<li class="headlines_item"><a href="#f406">The number of free support vectors is large. What should I do?</a></li>
<li class="headlines_item"><a href="#f407">Should I scale training and testing data in a similar way?</a></li>
<li class="headlines_item"><a href="#f408">Does it make a big difference if I scale each attribute to [0,1] instead of [-1,1]?</a></li>
<li class="headlines_item"><a href="#f409">The prediction rate is low. How could I improve it?</a></li>
<li class="headlines_item"><a href="#f410">My data are unbalanced. Could libsvm handle such problems?</a></li>
<li class="headlines_item"><a href="#f411">What is the difference between nu-SVC and C-SVC?</a></li>
<li class="headlines_item"><a href="#f412">The program keeps running (without showing any output). What should I do?</a></li>
<li class="headlines_item"><a href="#f413">The program keeps running (with output, i.e. many dots). What should I do?</a></li>
<li class="headlines_item"><a href="#f414">The training time is too long. What should I do?</a></li>
<li class="headlines_item"><a href="#f4141">Does shrinking always help?</a></li>
<li class="headlines_item"><a href="#f415">How do I get the decision value(s)?</a></li>
<li class="headlines_item"><a href="#f4151">How do I get the distance between a point and the hyperplane?</a></li>
<li class="headlines_item"><a href="#f416">On 32-bit machines, if I use a large cache (i.e. large -m) on a linux machine, why sometimes I get "segmentation fault ?"</a></li>
<li class="headlines_item"><a href="#f417">How do I disable screen output of svm-train?</a></li>
<li class="headlines_item"><a href="#f418">I would like to use my own kernel. Any example? In svm.cpp, there are two subroutines for kernel evaluations: k_function() and kernel_function(). Which one should I modify ?</a></li>
<li class="headlines_item"><a href="#f419">What method does libsvm use for multi-class SVM ? Why don't you use the "1-against-the rest" method?</a></li>
<li class="headlines_item"><a href="#f4191">How does LIBSVM perform parameter selection for multi-class problems? </a></li>
<li class="headlines_item"><a href="#f420">After doing cross validation, why there is no model file outputted ?</a></li>
<li class="headlines_item"><a href="#f4201">Why my cross-validation results are different from those in the Practical Guide?</a></li>
<li class="headlines_item"><a href="#f421">On some systems CV accuracy is the same in several runs. How could I use different data partitions? In other words, how do I set random seed in LIBSVM?</a></li>
<li class="headlines_item"><a href="#f422">I would like to solve L2-loss SVM (i.e., error term is quadratic). How should I modify the code ?</a></li>
<li class="headlines_item"><a href="#f424">How do I choose parameters for one-class svm as training data are in only one class?</a></li>
<li class="headlines_item"><a href="#f427">Why the code gives NaN (not a number) results?</a></li>
<li class="headlines_item"><a href="#f428">Why on windows sometimes grid.py fails?</a></li>
<li class="headlines_item"><a href="#f429">Why grid.py/easy.py sometimes generates the following warning message?</a></li>
<li class="headlines_item"><a href="#f430">Why the sign of predicted labels and decision values are sometimes reversed?</a></li>
<li class="headlines_item"><a href="#f431">I don't know class labels of test data. What should I put in the first column of the test file?</a></li>
<li class="headlines_item"><a href="#f432">How can I use OpenMP to parallelize LIBSVM on a multicore/shared-memory computer?</a></li>
<li class="headlines_item"><a href="#f433">How could I know which training instances are support vectors?</a></li>
<li class="headlines_item"><a href="#f425">Why training a probability model (i.e., -b 1) takes a longer time?</a></li>
<li class="headlines_item"><a href="#f426">Why using the -b option does not give me better accuracy?</a></li>
<li class="headlines_item"><a href="#f427">Why using svm-predict -b 0 and -b 1 gives different accuracy values?</a></li>
<li class="headlines_item"><a href="#f501">How can I save images drawn by svm-toy?</a></li>
<li class="headlines_item"><a href="#f502">I press the "load" button to load data points but why svm-toy does not draw them ?</a></li>
<li class="headlines_item"><a href="#f503">I would like svm-toy to handle more than three classes of data, what should I do ?</a></li>
<li class="headlines_item"><a href="#f601">What is the difference between Java version and C++ version of libsvm?</a></li>
<li class="headlines_item"><a href="#f602">Is the Java version significantly slower than the C++ version?</a></li>
<li class="headlines_item"><a href="#f603">While training I get the following error message: java.lang.OutOfMemoryError. What is wrong?</a></li>
<li class="headlines_item"><a href="#f604">Why you have the main source file svm.m4 and then transform it to svm.java?</a></li>
<li class="headlines_item"><a href="#f704">Except the python-C++ interface provided, could I use Jython to call libsvm ?</a></li>
<li class="headlines_item"><a href="#f801">I compile the MATLAB interface without problem, but why errors occur while running it?</a></li>
<li class="headlines_item"><a href="#f8011">On 64bit Windows I compile the MATLAB interface without problem, but why errors occur while running it?</a></li>
<li class="headlines_item"><a href="#f802">Does the MATLAB interface provide a function to do scaling?</a></li>
<li class="headlines_item"><a href="#f803">How could I use MATLAB interface for parameter selection?</a></li>
<li class="headlines_item"><a href="#f8031">I use MATLAB parallel programming toolbox on a multi-core environment for parameter selection. Why the program is even slower?</a></li>
<li class="headlines_item"><a href="#f8032">How do I use LIBSVM with OpenMP under MATLAB?</a></li>
<li class="headlines_item"><a href="#f804">How could I generate the primal variable w of linear SVM?</a></li>
<li class="headlines_item"><a href="#f805">Is there an OCTAVE interface for libsvm?</a></li>
<li class="headlines_item"><a href="#f806">How to handle the name conflict between svmtrain in the libsvm matlab interface and that in MATLAB bioinformatics toolbox?</a></li>
<li class="headlines_item"><a href="#f807">On Windows I got an error message "Invalid MEX-file: Specific module not found" when running the pre-built MATLAB interface in the windows sub-directory. What should I do?</a></li>
<li class="headlines_item"><a href="#f808">LIBSVM supports 1-vs-1 multi-class classification. If instead I would like to use 1-vs-rest, how to implement it using MATLAB interface?</a></li>
</ul></ul>
<hr size="5" noshade />
<p/>
<a name="/Q1:_Some_sample_uses_of_libsvm"></a>
<a name="faq101"><b>Q: Some courses which have used libsvm as a tool</b></a>
<br/>
<ul>
<li><a href=http://lmb.informatik.uni-freiburg.de/lectures/svm_seminar/>Institute for Computer Science,
Faculty of Applied Science, University of Freiburg, Germany
</a>
<li> <a href=http://www.cs.vu.nl/~elena/ml.html>
Division of Mathematics and Computer Science.
Faculteit der Exacte Wetenschappen
Vrije Universiteit, The Netherlands. </a>
<li>
<a href=http://www.cae.wisc.edu/~ece539/matlab/>
Electrical and Computer Engineering Department,
University of Wisconsin-Madison
</a>
<li>
<a href=http://www.hpl.hp.com/personal/Carl_Staelin/cs236601/project.html>
Technion (Israel Institute of Technology), Israel.
<li>
<a href=http://www.cise.ufl.edu/~fu/learn.html>
Computer and Information Sciences Dept., University of Florida</a>
<li>
<a href=http://www.uonbi.ac.ke/acad_depts/ics/course_material/machine_learning/ML_and_DM_Resources.html>
The Institute of Computer Science,
University of Nairobi, Kenya.</a>
<li>
<a href=http://cerium.raunvis.hi.is/~tpr/courseware/svm/hugbunadur.html>
Applied Mathematics and Computer Science, University of Iceland.
<li>
<a href=http://chicago05.mlss.cc/tiki/tiki-read_article.php?articleId=2>
SVM tutorial in machine learning
summer school, University of Chicago, 2005.
</a>
</ul>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q1:_Some_sample_uses_of_libsvm"></a>
<a name="faq102"><b>Q: Some applications/tools which have used libsvm </b></a>
<br/>
(and maybe liblinear).
<ul>
<li>
<a href=http://people.csail.mit.edu/jjl/libpmk/>LIBPMK: A Pyramid Match Toolkit</a>
</li>
<li><a href=http://maltparser.org/>Maltparser</a>:
a system for data-driven dependency parsing
</li>
<li>
<a href=http://www.pymvpa.org/>PyMVPA: python tool for classifying neuroimages</a>
</li>
<li>
<a href=http://solpro.proteomics.ics.uci.edu/>
SOLpro: protein solubility predictor
</a>
</li>
<li>
<a href=http://bdval.campagnelab.org>
BDVal</a>: biomarker discovery in high-throughput datasets.
</li>
<li><a href=http://johel.m.free.fr/demo_045.htm>
Realtime object recognition</a>
</li>
<li><a href=http://scikit-learn.sourceforge.net/>
scikits.learn: machine learning in Python</a>
</li>
</ul>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f201"><b>Q: Where can I find documents/videos of libsvm ?</b></a>
<br/>
<p>
<ul>
<li>
Official implementation document:
<br>
C.-C. Chang and
C.-J. Lin.
LIBSVM
: a library for support vector machines.
ACM Transactions on Intelligent
Systems and Technology, 2:27:1--27:27, 2011.
<a href="http://www.csie.ntu.edu.tw/~cjlin/papers/libsvm.pdf">pdf</a>, <a href=http://www.csie.ntu.edu.tw/~cjlin/papers/libsvm.ps.gz>ps.gz</a>,
<a href=http://portal.acm.org/citation.cfm?id=1961199&CFID=29950432&CFTOKEN=30974232>ACM digital lib</a>.
<li> Instructions for using LIBSVM are in the README files in the main directory and some sub-directories.
<br>
README in the main directory: details all options, data format, and library calls.
<br>
tools/README: parameter selection and other tools
<li>
A guide for beginners:
<br>
C.-W. Hsu, C.-C. Chang, and
C.-J. Lin.
<A HREF="http://www.csie.ntu.edu.tw/~cjlin/papers/guide/guide.pdf">
A practical guide to support vector classification
</A>
<li> An <a href=http://www.youtube.com/watch?v=gePWtNAQcK8>introductory video</a>
for windows users.
</ul>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f202"><b>Q: Where are change log and earlier versions?</b></a>
<br/>
<p>See <a href="http://www.csie.ntu.edu.tw/~cjlin/libsvm/log">the change log</a>.
<p> You can download earlier versions
<a href="http://www.csie.ntu.edu.tw/~cjlin/libsvm/oldfiles">here</a>.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f203"><b>Q: How to cite LIBSVM?</b></a>
<br/>
<p>
Please cite the following paper:
<p>
Chih-Chung Chang and Chih-Jen Lin, LIBSVM
: a library for support vector machines.
ACM Transactions on Intelligent Systems and Technology, 2:27:1--27:27, 2011.
Software available at http://www.csie.ntu.edu.tw/~cjlin/libsvm
<p>
The bibtex format is
<pre>
@article{CC01a,
author = {Chang, Chih-Chung and Lin, Chih-Jen},
title = {{LIBSVM}: A library for support vector machines},
journal = {ACM Transactions on Intelligent Systems and Technology},
volume = {2},
issue = {3},
year = {2011},
pages = {27:1--27:27},
note = {Software available at \url{http://www.csie.ntu.edu.tw/~cjlin/libsvm}}
}
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f204"><b>Q: I would like to use libsvm in my software. Is there any license problem?</b></a>
<br/>
<p>
The libsvm license ("the modified BSD license")
is compatible with many
free software licenses such as GPL. Hence, it is very easy to
use libsvm in your software.
Please check the COPYRIGHT file in detail. Basically
you need to
<ol>
<li>
Clearly indicate that LIBSVM is used.
</li>
<li>
Retain the LIBSVM COPYRIGHT file in your software.
</li>
</ol>
It can also be used in commercial products.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f205"><b>Q: Is there a repository of additional tools based on libsvm?</b></a>
<br/>
<p>
Yes, see <a href="http://www.csie.ntu.edu.tw/~cjlin/libsvmtools">libsvm
tools</a>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f206"><b>Q: On unix machines, I got "error in loading shared libraries" or "cannot open shared object file." What happened ? </b></a>
<br/>
<p>
This usually happens if you compile the code
on one machine and run it on another which has incompatible
libraries.
Try to recompile the program on that machine or use static linking.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f207"><b>Q: I have modified the source and would like to build the graphic interface "svm-toy" on MS windows. How should I do it ?</b></a>
<br/>
<p>
Build it as a project by choosing "Win32 Project."
On the other hand, for "svm-train" and "svm-predict"
you want to choose "Win32 Console Project."
After libsvm 2.5, you can also use the file Makefile.win.
See details in README.
<p>
If you are not using Makefile.win and see the following
link error
<pre>
LIBCMTD.lib(wwincrt0.obj) : error LNK2001: unresolved external symbol
_wWinMain@16
</pre>
you may have selected a wrong project type.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f208"><b>Q: I am an MS windows user but why only one (svm-toy) of those precompiled .exe actually runs ? </b></a>
<br/>
<p>
You need to open a command window
and type svmtrain.exe to see all options.
Some examples are in README file.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f209"><b>Q: What is the difference between "." and "*" outputed during training? </b></a>
<br/>
<p>
"." means every 1,000 iterations (or every #data
iterations is your #data is less than 1,000).
"*" means that after iterations of using
a smaller shrunk problem,
we reset to use the whole set. See the
<a href=../papers/libsvm.pdf>implementation document</a> for details.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f210"><b>Q: Why occasionally the program (including MATLAB or other interfaces) crashes and gives a segmentation fault?</b></a>
<br/>
<p>
Very likely the program consumes too much memory than what the
operating system can provide. Try a smaller data and see if the
program still crashes.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f211"><b>Q: How to build a dynamic library (.dll file) on MS windows?</b></a>
<br/>
<p>
The easiest way is to use Makefile.win.
See details in README.
Alternatively, you can use Visual C++. Here is
the example using Visual Studio .Net 2008:
<ol>
<li>Create a Win32 empty DLL project and set (in Project->$Project_Name
Properties...->Configuration) to "Release."
About how to create a new dynamic link library, please refer to
<a href=http://msdn2.microsoft.com/en-us/library/ms235636(VS.80).aspx>http://msdn2.microsoft.com/en-us/library/ms235636(VS.80).aspx</a>
<li> Add svm.cpp, svm.h to your project.
<li> Add __WIN32__ and _CRT_SECURE_NO_DEPRECATE to Preprocessor definitions (in
Project->$Project_Name Properties...->C/C++->Preprocessor)
<li> Set Create/Use Precompiled Header to Not Using Precompiled Headers
(in Project->$Project_Name Properties...->C/C++->Precompiled Headers)
<li> Set the path for the Modulation Definition File svm.def (in
Project->$Project_Name Properties...->Linker->input
<li> Build the DLL.
<li> Rename the dll file to libsvm.dll and move it to the correct path.
</ol>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f212"><b>Q: On some systems (e.g., Ubuntu), compiling LIBSVM gives many warning messages. Is this a problem and how to disable the warning message?</b></a>
<br/>
<p>
The warning message is like
<pre>
svm.cpp:2730: warning: ignoring return value of int fscanf(FILE*, const char*, ...), declared with attribute warn_unused_result
</pre>
This is not a problem; see <a href=https://wiki.ubuntu.com/CompilerFlags#-D_FORTIFY_SOURCE=2>this page</a> for more
details of ubuntu systems.
In the future we may modify the code
so that these messages do not appear.
At this moment, to disable the warning message you can replace
<pre>
CFLAGS = -Wall -Wconversion -O3 -fPIC
</pre>
with
<pre>
CFLAGS = -Wall -Wconversion -O3 -fPIC -U_FORTIFY_SOURCE
</pre>
in Makefile.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q2:_Installation_and_running_the_program"></a>
<a name="f213"><b>Q: In LIBSVM, why you don't use certain C/C++ library functions to make the code shorter?</b></a>
<br/>
<p>
For portability, we use only features defined in ISO C89. Note that features in ISO C99 may not be available everywhere.
Even the newest gcc lacks some features in C99 (see <a href=http://gcc.gnu.org/c99status.html>http://gcc.gnu.org/c99status.html</a> for details).
If the situation changes in the future,
we might consider using these newer features.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f301"><b>Q: Why sometimes not all attributes of a data appear in the training/model files ?</b></a>
<br/>
<p>
libsvm uses the so called "sparse" format where zero
values do not need to be stored. Hence a data with attributes
<pre>
1 0 2 0
</pre>
is represented as
<pre>
1:1 3:2
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f302"><b>Q: What if my data are non-numerical ?</b></a>
<br/>
<p>
Currently libsvm supports only numerical data.
You may have to change non-numerical data to
numerical. For example, you can use several
binary attributes to represent a categorical
attribute.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f303"><b>Q: Why do you consider sparse format ? Will the training of dense data be much slower ?</b></a>
<br/>
<p>
This is a controversial issue. The kernel
evaluation (i.e. inner product) of sparse vectors is slower
so the total training time can be at least twice or three times
of that using the dense format.
However, we cannot support only dense format as then we CANNOT
handle extremely sparse cases. Simplicity of the code is another
concern. Right now we decide to support
the sparse format only.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f304"><b>Q: Why sometimes the last line of my data is not read by svm-train?</b></a>
<br/>
<p>
We assume that you have '\n' in the end of
each line. So please press enter in the end
of your last line.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f305"><b>Q: Is there a program to check if my data are in the correct format?</b></a>
<br/>
<p>
The svm-train program in libsvm conducts only a simple check of the input data. To do a
detailed check, after libsvm 2.85, you can use the python script tools/checkdata.py. See tools/README for details.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f306"><b>Q: May I put comments in data files?</b></a>
<br/>
<p>
We don't officially support this. But, cureently LIBSVM
is able to process data in the following
format:
<pre>
1 1:2 2:1 # your comments
</pre>
Note that the character ":" should not appear in your
comments.
<!--
No, for simplicity we don't support that.
However, you can easily preprocess your data before
using libsvm. For example,
if you have the following data
<pre>
test.txt
1 1:2 2:1 # proten A
</pre>
then on unix machines you can do
<pre>
cut -d '#' -f 1 < test.txt > test.features
cut -d '#' -f 2 < test.txt > test.comments
svm-predict test.feature train.model test.predicts
paste -d '#' test.predicts test.comments | sed 's/#/ #/' > test.results
</pre>
-->
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q3:_Data_preparation"></a>
<a name="f307"><b>Q: How to convert other data formats to LIBSVM format?</b></a>
<br/>
<p>
It depends on your data format. A simple way is to use
libsvmwrite in the libsvm matlab/octave interface.
Take a CSV (comma-separated values) file
in UCI machine learning repository as an example.
We download <a href=http://archive.ics.uci.edu/ml/machine-learning-databases/spect/SPECTF.train>SPECTF.train</a>.
Labels are in the first column. The following steps produce
a file in the libsvm format.
<pre>
matlab> SPECTF = csvread('SPECTF.train'); % read a csv file
matlab> labels = SPECTF(:, 1); % labels from the 1st column
matlab> features = SPECTF(:, 2:end);
matlab> features_sparse = sparse(features); % features must be in a sparse matrix
matlab> libsvmwrite('SPECTFlibsvm.train', labels, features_sparse);
</pre>
The tranformed data are stored in SPECTFlibsvm.train.
<p>
Alternatively, you can use <a href="./faqfiles/convert.c">convert.c</a>
to convert CSV format to libsvm format.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f401"><b>Q: The output of training C-SVM is like the following. What do they mean?</b></a>
<br/>
<br>optimization finished, #iter = 219
<br>nu = 0.431030
<br>obj = -100.877286, rho = 0.424632
<br>nSV = 132, nBSV = 107
<br>Total nSV = 132
<p>
obj is the optimal objective value of the dual SVM problem.
rho is the bias term in the decision function
sgn(w^Tx - rho).
nSV and nBSV are number of support vectors and bounded support
vectors (i.e., alpha_i = C). nu-svm is a somewhat equivalent
form of C-SVM where C is replaced by nu. nu simply shows the
corresponding parameter. More details are in
<a href="http://www.csie.ntu.edu.tw/~cjlin/papers/libsvm.pdf">
libsvm document</a>.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f402"><b>Q: Can you explain more about the model file?</b></a>
<br/>
<p>
In the model file, after parameters and other informations such as labels , each line represents a support vector.
Support vectors are listed in the order of "labels" shown earlier.
(i.e., those from the first class in the "labels" list are
grouped first, and so on.)
If k is the total number of classes,
in front of a support vector in class j, there are
k-1 coefficients
y*alpha where alpha are dual solution of the
following two class problems:
<br>
1 vs j, 2 vs j, ..., j-1 vs j, j vs j+1, j vs j+2, ..., j vs k
<br>
and y=1 in first j-1 coefficients, y=-1 in the remaining
k-j coefficients.
For example, if there are 4 classes, the file looks like:
<pre>
+-+-+-+--------------------+
|1|1|1| |
|v|v|v| SVs from class 1 |
|2|3|4| |
+-+-+-+--------------------+
|1|2|2| |
|v|v|v| SVs from class 2 |
|2|3|4| |
+-+-+-+--------------------+
|1|2|3| |
|v|v|v| SVs from class 3 |
|3|3|4| |
+-+-+-+--------------------+
|1|2|3| |
|v|v|v| SVs from class 4 |
|4|4|4| |
+-+-+-+--------------------+
</pre>
See also
<a href="#f804"> an illustration using
MATLAB/OCTAVE.</a>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f403"><b>Q: Should I use float or double to store numbers in the cache ?</b></a>
<br/>
<p>
We have float as the default as you can store more numbers
in the cache.
In general this is good enough but for few difficult
cases (e.g. C very very large) where solutions are huge
numbers, it might be possible that the numerical precision is not
enough using only float.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f404"><b>Q: How do I choose the kernel?</b></a>
<br/>
<p>
In general we suggest you to try the RBF kernel first.
A recent result by Keerthi and Lin
(<a href=http://www.csie.ntu.edu.tw/~cjlin/papers/limit.pdf>
download paper here</a>)
shows that if RBF is used with model selection,
then there is no need to consider the linear kernel.
The kernel matrix using sigmoid may not be positive definite
and in general it's accuracy is not better than RBF.
(see the paper by Lin and Lin
(<a href=http://www.csie.ntu.edu.tw/~cjlin/papers/tanh.pdf>
download paper here</a>).
Polynomial kernels are ok but if a high degree is used,
numerical difficulties tend to happen
(thinking about dth power of (<1) goes to 0
and (>1) goes to infinity).
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f405"><b>Q: Does libsvm have special treatments for linear SVM?</b></a>
<br/>
<p>
No, libsvm solves linear/nonlinear SVMs by the
same way.
Some tricks may save training/testing time if the
linear kernel is used,
so libsvm is <b>NOT</b> particularly efficient for linear SVM,
especially when
C is large and
the number of data is much larger
than the number of attributes.
You can either
<ul>
<li>
Use small C only. We have shown in the following paper
that after C is larger than a certain threshold,
the decision function is the same.
<p>
<a href="http://guppy.mpe.nus.edu.sg/~mpessk/">S. S. Keerthi</a>
and
<B>C.-J. Lin</B>.
<A HREF="papers/limit.pdf">
Asymptotic behaviors of support vector machines with
Gaussian kernel
</A>
.
<I><A HREF="http://mitpress.mit.edu/journal-home.tcl?issn=08997667">Neural Computation</A></I>, 15(2003), 1667-1689.
<li>
Check <a href=http://www.csie.ntu.edu.tw/~cjlin/liblinear>liblinear</a>,
which is designed for large-scale linear classification.
</ul>
<p> Please also see our <a href=../papers/guide/guide.pdf>SVM guide</a>
on the discussion of using RBF and linear
kernels.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f406"><b>Q: The number of free support vectors is large. What should I do?</b></a>
<br/>
<p>
This usually happens when the data are overfitted.
If attributes of your data are in large ranges,
try to scale them. Then the region
of appropriate parameters may be larger.
Note that there is a scale program
in libsvm.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f407"><b>Q: Should I scale training and testing data in a similar way?</b></a>
<br/>
<p>
Yes, you can do the following:
<pre>
> svm-scale -s scaling_parameters train_data > scaled_train_data
> svm-scale -r scaling_parameters test_data > scaled_test_data
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f408"><b>Q: Does it make a big difference if I scale each attribute to [0,1] instead of [-1,1]?</b></a>
<br/>
<p>
For the linear scaling method, if the RBF kernel is
used and parameter selection is conducted, there
is no difference. Assume Mi and mi are
respectively the maximal and minimal values of the
ith attribute. Scaling to [0,1] means
<pre>
x'=(x-mi)/(Mi-mi)
</pre>
For [-1,1],
<pre>
x''=2(x-mi)/(Mi-mi)-1.
</pre>
In the RBF kernel,
<pre>
x'-y'=(x-y)/(Mi-mi), x''-y''=2(x-y)/(Mi-mi).
</pre>
Hence, using (C,g) on the [0,1]-scaled data is the
same as (C,g/2) on the [-1,1]-scaled data.
<p> Though the performance is the same, the computational
time may be different. For data with many zero entries,
[0,1]-scaling keeps the sparsity of input data and hence
may save the time.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f409"><b>Q: The prediction rate is low. How could I improve it?</b></a>
<br/>
<p>
Try to use the model selection tool grid.py in the python
directory find
out good parameters. To see the importance of model selection,
please
see my talk:
<A HREF="http://www.csie.ntu.edu.tw/~cjlin/talks/freiburg.pdf">
A practical guide to support vector
classification
</A>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f410"><b>Q: My data are unbalanced. Could libsvm handle such problems?</b></a>
<br/>
<p>
Yes, there is a -wi options. For example, if you use
<pre>
> svm-train -s 0 -c 10 -w1 1 -w-1 5 data_file
</pre>
<p>
the penalty for class "-1" is larger.
Note that this -w option is for C-SVC only.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f411"><b>Q: What is the difference between nu-SVC and C-SVC?</b></a>
<br/>
<p>
Basically they are the same thing but with different
parameters. The range of C is from zero to infinity
but nu is always between [0,1]. A nice property
of nu is that it is related to the ratio of
support vectors and the ratio of the training
error.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f412"><b>Q: The program keeps running (without showing any output). What should I do?</b></a>
<br/>
<p>
You may want to check your data. Each training/testing
data must be in one line. It cannot be separated.
In addition, you have to remove empty lines.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f413"><b>Q: The program keeps running (with output, i.e. many dots). What should I do?</b></a>
<br/>
<p>
In theory libsvm guarantees to converge.
Therefore, this means you are
handling ill-conditioned situations
(e.g. too large/small parameters) so numerical
difficulties occur.
<p>
You may get better numerical stability by replacing
<pre>
typedef float Qfloat;
</pre>
in svm.cpp with
<pre>
typedef double Qfloat;
</pre>
That is, elements in the kernel cache are stored
in double instead of single. However, this means fewer elements
can be put in the kernel cache.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f414"><b>Q: The training time is too long. What should I do?</b></a>
<br/>
<p>
For large problems, please specify enough cache size (i.e.,
-m).
Slow convergence may happen for some difficult cases (e.g. -c is large).
You can try to use a looser stopping tolerance with -e.
If that still doesn't work, you may train only a subset of the data.
You can use the program subset.py in the directory "tools"
to obtain a random subset.
<p>
If you have extremely large data and face this difficulty, please
contact us. We will be happy to discuss possible solutions.
<p> When using large -e, you may want to check if -h 0 (no shrinking) or -h 1 (shrinking) is faster.
See a related question below.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f4141"><b>Q: Does shrinking always help?</b></a>
<br/>
<p>
If the number of iterations is high, then shrinking
often helps.
However, if the number of iterations is small
(e.g., you specify a large -e), then
probably using -h 0 (no shrinking) is better.
See the
<a href=../papers/libsvm.pdf>implementation document</a> for details.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f415"><b>Q: How do I get the decision value(s)?</b></a>
<br/>
<p>
We print out decision values for regression. For classification,
we solve several binary SVMs for multi-class cases. You
can obtain values by easily calling the subroutine
svm_predict_values. Their corresponding labels
can be obtained from svm_get_labels.
Details are in
README of libsvm package.
<p>
If you are using MATLAB/OCTAVE interface, svmpredict can directly
give you decision values. Please see matlab/README for details.
<p>
We do not recommend the following. But if you would
like to get values for
TWO-class classification with labels +1 and -1
(note: +1 and -1 but not things like 5 and 10)
in the easiest way, simply add
<pre>
printf("%f\n", dec_values[0]*model->label[0]);
</pre>
after the line
<pre>
svm_predict_values(model, x, dec_values);
</pre>
of the file svm.cpp.
Positive (negative)
decision values correspond to data predicted as +1 (-1).
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f4151"><b>Q: How do I get the distance between a point and the hyperplane?</b></a>
<br/>
<p>
The distance is |decision_value| / |w|.
We have |w|^2 = w^Tw = alpha^T Q alpha = 2*(dual_obj + sum alpha_i).
Thus in svm.cpp please find the place
where we calculate the dual objective value
(i.e., the subroutine Solve())
and add a statement to print w^Tw.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f416"><b>Q: On 32-bit machines, if I use a large cache (i.e. large -m) on a linux machine, why sometimes I get "segmentation fault ?"</b></a>
<br/>
<p>
On 32-bit machines, the maximum addressable
memory is 4GB. The Linux kernel uses 3:1
split which means user space is 3G and
kernel space is 1G. Although there are
3G user space, the maximum dynamic allocation
memory is 2G. So, if you specify -m near 2G,
the memory will be exhausted. And svm-train
will fail when it asks more memory.
For more details, please read
<a href=http://groups.google.com/groups?hl=en&lr=&ie=UTF-8&selm=3BA164F6.BAFA4FB%40daimi.au.dk>
this article</a>.
<p>
The easiest solution is to switch to a
64-bit machine.
Otherwise, there are two ways to solve this. If your
machine supports Intel's PAE (Physical Address
Extension), you can turn on the option HIGHMEM64G
in Linux kernel which uses 4G:4G split for
kernel and user space. If you don't, you can
try a software `tub' which can eliminate the 2G
boundary for dynamic allocated memory. The `tub'
is available at
<a href=http://www.bitwagon.com/tub.html>http://www.bitwagon.com/tub.html</a>.
<!--
This may happen only when the cache is large, but each cached row is
not large enough. <b>Note:</b> This problem is specific to
gnu C library which is used in linux.
The solution is as follows:
<p>
In our program we have malloc() which uses two methods
to allocate memory from kernel. One is
sbrk() and another is mmap(). sbrk is faster, but mmap
has a larger address
space. So malloc uses mmap only if the wanted memory size is larger
than some threshold (default 128k).
In the case where each row is not large enough (#elements < 128k/sizeof(float)) but we need a large cache ,
the address space for sbrk can be exhausted. The solution is to
lower the threshold to force malloc to use mmap
and increase the maximum number of chunks to allocate
with mmap.
<p>
Therefore, in the main program (i.e. svm-train.c) you want
to have
<pre>
#include <malloc.h>
</pre>
and then in main():
<pre>
mallopt(M_MMAP_THRESHOLD, 32768);
mallopt(M_MMAP_MAX,1000000);
</pre>
You can also set the environment variables instead
of writing them in the program:
<pre>
$ M_MMAP_MAX=1000000 M_MMAP_THRESHOLD=32768 ./svm-train .....
</pre>
More information can be found by
<pre>
$ info libc "Malloc Tunable Parameters"
</pre>
-->
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f417"><b>Q: How do I disable screen output of svm-train?</b></a>
<br/>
<p>
For commend-line users, use the -q option:
<pre>
> ./svm-train -q heart_scale
</pre>
<p>
For library users, set the global variable
<pre>
extern void (*svm_print_string) (const char *);
</pre>
to specify the output format. You can disable the output by the following steps:
<ol>
<li>
Declare a function to output nothing:
<pre>
void print_null(const char *s) {}
</pre>
</li>
<li>
Assign the output function of libsvm by
<pre>
svm_print_string = &print_null;
</pre>
</li>
</ol>
Finally, a way used in earlier libsvm
is by updating svm.cpp from
<pre>
#if 1
void info(const char *fmt,...)
</pre>
to
<pre>
#if 0
void info(const char *fmt,...)
</pre>
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f418"><b>Q: I would like to use my own kernel. Any example? In svm.cpp, there are two subroutines for kernel evaluations: k_function() and kernel_function(). Which one should I modify ?</b></a>
<br/>
<p>
An example is "LIBSVM for string data" in LIBSVM Tools.
<p>
The reason why we have two functions is as follows.
For the RBF kernel exp(-g |xi - xj|^2), if we calculate
xi - xj first and then the norm square, there are 3n operations.
Thus we consider exp(-g (|xi|^2 - 2dot(xi,xj) +|xj|^2))
and by calculating all |xi|^2 in the beginning,
the number of operations is reduced to 2n.
This is for the training. For prediction we cannot
do this so a regular subroutine using that 3n operations is
needed.
The easiest way to have your own kernel is
to put the same code in these two
subroutines by replacing any kernel.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f419"><b>Q: What method does libsvm use for multi-class SVM ? Why don't you use the "1-against-the rest" method?</b></a>
<br/>
<p>
It is one-against-one. We chose it after doing the following
comparison:
C.-W. Hsu and C.-J. Lin.
<A HREF="http://www.csie.ntu.edu.tw/~cjlin/papers/multisvm.pdf">
A comparison of methods
for multi-class support vector machines
</A>,
<I>IEEE Transactions on Neural Networks</A></I>, 13(2002), 415-425.
<p>
"1-against-the rest" is a good method whose performance
is comparable to "1-against-1." We do the latter
simply because its training time is shorter.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f4191"><b>Q: How does LIBSVM perform parameter selection for multi-class problems? </b></a>
<br/>
<p>
LIBSVM implements "one-against-one" multi-class method, so there are
k(k-1)/2 binary models, where k is the number of classes.
<p>
We can consider two ways to conduct parameter selection.
<ol>
<li>
For any two classes of data, a parameter selection procedure is conducted. Finally,
each decision function has its own optimal parameters.
</li>
<li>
The same parameters are used for all k(k-1)/2 binary classification problems.
We select parameters that achieve the highest overall performance.
</li>
</ol>
Each has its own advantages. A
single parameter set may not be uniformly good for all k(k-1)/2 decision functions.
However, as the overall accuracy is the final consideration, one parameter set
for one decision function may lead to over-fitting. In the paper
<p>
Chen, Lin, and Schölkopf,
<A HREF="../papers/nusvmtutorial.pdf">
A tutorial on nu-support vector machines.
</A>
Applied Stochastic Models in Business and Industry, 21(2005), 111-136,
<p>
they have experimentally
shown that the two methods give similar performance.
Therefore, currently the parameter selection in LIBSVM
takes the second approach by considering the same parameters for
all k(k-1)/2 models.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f420"><b>Q: After doing cross validation, why there is no model file outputted ?</b></a>
<br/>
<p>
Cross validation is used for selecting good parameters.
After finding them, you want to re-train the whole
data without the -v option.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f4201"><b>Q: Why my cross-validation results are different from those in the Practical Guide?</b></a>
<br/>
<p>
Due to random partitions of
the data, on different systems CV accuracy values
may be different.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f421"><b>Q: On some systems CV accuracy is the same in several runs. How could I use different data partitions? In other words, how do I set random seed in LIBSVM?</b></a>
<br/>
<p>
If you use GNU C library,
the default seed 1 is considered. Thus you always
get the same result of running svm-train -v.
To have different seeds, you can add the following code
in svm-train.c:
<pre>
#include <time.h>
</pre>
and in the beginning of main(),
<pre>
srand(time(0));
</pre>
Alternatively, if you are not using GNU C library
and would like to use a fixed seed, you can have
<pre>
srand(1);
</pre>
<p>
For Java, the random number generator
is initialized using the time information.
So results of two CV runs are different.
To fix the seed, after version 3.1 (released
in mid 2011), you can add
<pre>
svm.rand.setSeed(0);
</pre>
in the main() function of svm_train.java.
<p>
If you use CV to select parameters, it is recommended to use identical folds
under different parameters. In this case, you can consider fixing the seed.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f422"><b>Q: I would like to solve L2-loss SVM (i.e., error term is quadratic). How should I modify the code ?</b></a>
<br/>
<p>
It is extremely easy. Taking c-svc for example, to solve
<p>
min_w w^Tw/2 + C \sum max(0, 1- (y_i w^Tx_i+b))^2,
<p>
only two
places of svm.cpp have to be changed.
First, modify the following line of
solve_c_svc from
<pre>
s.Solve(l, SVC_Q(*prob,*param,y), minus_ones, y,
alpha, Cp, Cn, param->eps, si, param->shrinking);
</pre>
to
<pre>
s.Solve(l, SVC_Q(*prob,*param,y), minus_ones, y,
alpha, INF, INF, param->eps, si, param->shrinking);
</pre>
Second, in the class of SVC_Q, declare C as
a private variable:
<pre>
double C;
</pre>
In the constructor replace
<pre>
for(int i=0;i<prob.l;i++)
QD[i]= (Qfloat)(this->*kernel_function)(i,i);
</pre>
with
<pre>
this->C = param.C;
for(int i=0;i<prob.l;i++)
QD[i]= (Qfloat)(this->*kernel_function)(i,i)+0.5/C;
</pre>
Then in the subroutine get_Q, after the for loop, add
<pre>
if(i >= start && i < len)
data[i] += 0.5/C;
</pre>
<p>
For one-class svm, the modification is exactly the same. For SVR, you don't need an if statement like the above. Instead, you only need a simple assignment:
<pre>
data[real_i] += 0.5/C;
</pre>
<p>
For large linear L2-loss SVM, please use
<a href=../liblinear>LIBLINEAR</a>.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f424"><b>Q: How do I choose parameters for one-class svm as training data are in only one class?</b></a>
<br/>
<p>
You have pre-specified true positive rate in mind and then search for
parameters which achieve similar cross-validation accuracy.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f427"><b>Q: Why the code gives NaN (not a number) results?</b></a>
<br/>
<p>
This rarely happens, but few users reported the problem.
It seems that their
computers for training libsvm have the VPN client
running. The VPN software has some bugs and causes this
problem. Please try to close or disconnect the VPN client.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f428"><b>Q: Why on windows sometimes grid.py fails?</b></a>
<br/>
<p>
This problem shouldn't happen after version
2.85. If you are using earlier versions,
please download the latest one.
<!--
<p>
If you are using earlier
versions, the error message is probably
<pre>
Traceback (most recent call last):
File "grid.py", line 349, in ?
main()
File "grid.py", line 344, in main
redraw(db)
File "grid.py", line 132, in redraw
gnuplot.write("set term windows\n")
IOError: [Errno 22] Invalid argument
</pre>
<p>Please try to close gnuplot windows and rerun.
If the problem still occurs, comment the following
two lines in grid.py by inserting "#" in the beginning:
<pre>
redraw(db)
redraw(db,1)
</pre>
Then you get accuracy only but not cross validation contours.
-->
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f429"><b>Q: Why grid.py/easy.py sometimes generates the following warning message?</b></a>
<br/>
<pre>
Warning: empty z range [62.5:62.5], adjusting to [61.875:63.125]
Notice: cannot contour non grid data!
</pre>
<p>Nothing is wrong and please disregard the
message. It is from gnuplot when drawing
the contour.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f430"><b>Q: Why the sign of predicted labels and decision values are sometimes reversed?</b></a>
<br/>
<p>Nothing is wrong. Very likely you have two labels +1/-1 and the first instance in your data
has -1.
Think about the case of labels +5/+10. Since
SVM needs to use +1/-1, internally
we map +5/+10 to +1/-1 according to which
label appears first.
Hence a positive decision value implies
that we should predict the "internal" +1,
which may not be the +1 in the input file.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f431"><b>Q: I don't know class labels of test data. What should I put in the first column of the test file?</b></a>
<br/>
<p>Any value is ok. In this situation, what you will use is the output file of svm-predict, which gives predicted class labels.
<p align="right">
<a href="#_TOP">[Go Top]</a>
<hr/>
<a name="/Q4:_Training_and_prediction"></a>
<a name="f432"><b>Q: How can I use OpenMP to parallelize LIBSVM on a multicore/shared-memory computer?</b></a>
<br/>
<p>It is very easy if you are using GCC 4.2
or after.
<p> In Makefile, add -fopenmp to CFLAGS.
<p> In class SVC_Q of svm.cpp, modify the for loop
of get_Q to:
<pre>
#pragma omp parallel for private(j)
for(j=start;j<len;j++)
</pre>
<p> In the subroutine svm_predict_values of svm.cpp, add one line to the for loop:
<pre>
#pragma omp parallel for private(i)
for(i=0;i<l;i++)
kvalue[i] = Kernel::k_function(x,model->SV[i],model->param);
</pre>
For regression, you need to modify
class SVR_Q instead. The loop in svm_predict_values
is also different because you need
a reduction clause for the variable sum:
<pre>
#pragma omp parallel for private(i) reduction(+:sum)
for(i=0;i<model->l;i++)
sum += sv_coef[i] * Kernel::k_function(x,model->SV[i],model->param);
</pre>
<p> Then rebuild the package. Kernel evaluations in training/testing will be parallelized. An example of running this modification on
an 8-core machine using the data set
<a href=../libsvmtools/datasets/binary/ijcnn1.bz2>ijcnn1</a>:
<p> 8 cores:
<pre>
%setenv OMP_NUM_THREADS 8
%time svm-train -c 16 -g 4 -m 400 ijcnn1
27.1sec
</pre>
1 core:
<pre>
%setenv OMP_NUM_THREADS 1
%time svm-train -c 16 -g 4 -m 400 ijcnn1
79.8sec
</pre>
For this data, kernel evaluations take 80% of training time. In the above example, we assume you use csh. For bash, use
<pre>
export OMP_NUM_THREADS=8
</pre>
instead.
<p> For Python interface, you need to add the -lgomp link option:
<pre>
$(CXX) -lgomp -shared -dynamiclib svm.o -o
gitextract_hg6xup5t/
├── README.md
├── binaries/
│ ├── linux/
│ │ ├── COPYRIGHT
│ │ ├── FAQ.html
│ │ ├── README
│ │ ├── README-GPU
│ │ ├── svm-predict
│ │ ├── svm-scale
│ │ ├── svm-train
│ │ ├── svm-train-gpu
│ │ ├── tools/
│ │ │ ├── README
│ │ │ ├── checkdata.py
│ │ │ ├── easy.py
│ │ │ ├── grid.py
│ │ │ └── subset.py
│ │ └── train_set
│ └── windows/
│ ├── x64/
│ │ ├── COPYRIGHT
│ │ ├── FAQ.html
│ │ ├── README
│ │ ├── README-GPU
│ │ ├── tools/
│ │ │ ├── README
│ │ │ ├── checkdata.py
│ │ │ ├── easy.py
│ │ │ ├── grid.py
│ │ │ └── subset.py
│ │ └── train_set
│ └── x86/
│ ├── COPYRIGHT
│ ├── FAQ.html
│ ├── README
│ ├── README-GPU
│ ├── tools/
│ │ ├── README
│ │ ├── checkdata.py
│ │ ├── easy.py
│ │ ├── grid.py
│ │ └── subset.py
│ └── train_set
└── src/
├── linux/
│ ├── COPYRIGHT
│ ├── Makefile
│ ├── README
│ ├── README-GPU
│ ├── cross_validation_with_matrix_precomputation.c
│ ├── findcudalib.mk
│ ├── kernel_matrix_calculation.c
│ ├── readme.txt
│ ├── svm-train.c
│ ├── svm.cpp
│ └── svm.h
└── windows/
├── README-GPU
├── libsvm_train_dense_gpu/
│ ├── cross_validation_with_matrix_precomputation.c
│ ├── kernel_matrix_calculation.c
│ ├── libsvm_train_dense_gpu.vcxproj
│ ├── libsvm_train_dense_gpu.vcxproj.filters
│ ├── libsvm_train_dense_gpu.vcxproj.user
│ ├── svm-train.c
│ ├── svm.cpp
│ └── svm.h
├── libsvm_train_dense_gpu.ncb
├── libsvm_train_dense_gpu.sdf
├── libsvm_train_dense_gpu.sln
└── libsvm_train_dense_gpu.suo
SYMBOL INDEX (329 symbols across 19 files)
FILE: binaries/linux/tools/checkdata.py
function err (line 18) | def err(line_no, msg):
function my_float (line 22) | def my_float(x):
function main (line 28) | def main():
FILE: binaries/linux/tools/grid.py
class GridOption (line 17) | class GridOption:
method __init__ (line 18) | def __init__(self, dataset_pathname, options):
method parse_options (line 40) | def parse_options(self, options):
function redraw (line 105) | def redraw(db,best_param,gnuplot,options,tofile=False):
function calculate_jobs (line 159) | def calculate_jobs(options):
class WorkerStopToken (line 241) | class WorkerStopToken: # used to notify the worker to stop or if a work...
class Worker (line 244) | class Worker(Thread):
method __init__ (line 245) | def __init__(self,name,job_queue,result_queue,options):
method run (line 252) | def run(self):
method get_cmd (line 278) | def get_cmd(self,c,g):
class LocalWorker (line 289) | class LocalWorker(Worker):
method run_one (line 290) | def run_one(self,c,g):
class SSHWorker (line 297) | class SSHWorker(Worker):
method __init__ (line 298) | def __init__(self,name,job_queue,result_queue,host,options):
method run_one (line 302) | def run_one(self,c,g):
class TelnetWorker (line 310) | class TelnetWorker(Worker):
method __init__ (line 311) | def __init__(self,name,job_queue,result_queue,host,username,password,o...
method run (line 316) | def run(self):
method run_one (line 331) | def run_one(self,c,g):
function find_parameters (line 339) | def find_parameters(dataset_pathname, options=''):
function exit_with_help (line 465) | def exit_with_help():
FILE: binaries/linux/tools/subset.py
function exit_with_help (line 9) | def exit_with_help(argv):
function process_options (line 25) | def process_options(argv):
function random_selection (line 56) | def random_selection(dataset, subset_size):
function stratified_selection (line 60) | def stratified_selection(dataset, subset_size):
function main (line 89) | def main(argv=sys.argv):
FILE: binaries/windows/x64/tools/checkdata.py
function err (line 18) | def err(line_no, msg):
function my_float (line 22) | def my_float(x):
function main (line 28) | def main():
FILE: binaries/windows/x64/tools/grid.py
class GridOption (line 17) | class GridOption:
method __init__ (line 18) | def __init__(self, dataset_pathname, options):
method parse_options (line 40) | def parse_options(self, options):
function redraw (line 105) | def redraw(db,best_param,gnuplot,options,tofile=False):
function calculate_jobs (line 159) | def calculate_jobs(options):
class WorkerStopToken (line 241) | class WorkerStopToken: # used to notify the worker to stop or if a work...
class Worker (line 244) | class Worker(Thread):
method __init__ (line 245) | def __init__(self,name,job_queue,result_queue,options):
method run (line 252) | def run(self):
method get_cmd (line 278) | def get_cmd(self,c,g):
class LocalWorker (line 289) | class LocalWorker(Worker):
method run_one (line 290) | def run_one(self,c,g):
class SSHWorker (line 297) | class SSHWorker(Worker):
method __init__ (line 298) | def __init__(self,name,job_queue,result_queue,host,options):
method run_one (line 302) | def run_one(self,c,g):
class TelnetWorker (line 310) | class TelnetWorker(Worker):
method __init__ (line 311) | def __init__(self,name,job_queue,result_queue,host,username,password,o...
method run (line 316) | def run(self):
method run_one (line 331) | def run_one(self,c,g):
function find_parameters (line 339) | def find_parameters(dataset_pathname, options=''):
function exit_with_help (line 465) | def exit_with_help():
FILE: binaries/windows/x64/tools/subset.py
function exit_with_help (line 9) | def exit_with_help(argv):
function process_options (line 25) | def process_options(argv):
function random_selection (line 56) | def random_selection(dataset, subset_size):
function stratified_selection (line 60) | def stratified_selection(dataset, subset_size):
function main (line 89) | def main(argv=sys.argv):
FILE: binaries/windows/x86/tools/checkdata.py
function err (line 18) | def err(line_no, msg):
function my_float (line 22) | def my_float(x):
function main (line 28) | def main():
FILE: binaries/windows/x86/tools/grid.py
class GridOption (line 17) | class GridOption:
method __init__ (line 18) | def __init__(self, dataset_pathname, options):
method parse_options (line 40) | def parse_options(self, options):
function redraw (line 105) | def redraw(db,best_param,gnuplot,options,tofile=False):
function calculate_jobs (line 159) | def calculate_jobs(options):
class WorkerStopToken (line 241) | class WorkerStopToken: # used to notify the worker to stop or if a work...
class Worker (line 244) | class Worker(Thread):
method __init__ (line 245) | def __init__(self,name,job_queue,result_queue,options):
method run (line 252) | def run(self):
method get_cmd (line 278) | def get_cmd(self,c,g):
class LocalWorker (line 289) | class LocalWorker(Worker):
method run_one (line 290) | def run_one(self,c,g):
class SSHWorker (line 297) | class SSHWorker(Worker):
method __init__ (line 298) | def __init__(self,name,job_queue,result_queue,host,options):
method run_one (line 302) | def run_one(self,c,g):
class TelnetWorker (line 310) | class TelnetWorker(Worker):
method __init__ (line 311) | def __init__(self,name,job_queue,result_queue,host,username,password,o...
method run (line 316) | def run(self):
method run_one (line 331) | def run_one(self,c,g):
function find_parameters (line 339) | def find_parameters(dataset_pathname, options=''):
function exit_with_help (line 465) | def exit_with_help():
FILE: binaries/windows/x86/tools/subset.py
function exit_with_help (line 9) | def exit_with_help(argv):
function process_options (line 25) | def process_options(argv):
function random_selection (line 56) | def random_selection(dataset, subset_size):
function stratified_selection (line 60) | def stratified_selection(dataset, subset_size):
function main (line 89) | def main(argv=sys.argv):
FILE: src/linux/cross_validation_with_matrix_precomputation.c
function setup_pkm (line 1) | void setup_pkm(struct svm_problem *p_km)
function free_pkm (line 19) | void free_pkm(struct svm_problem *p_km)
function do_crossvalidation (line 33) | double do_crossvalidation(struct svm_problem * p_km)
function run_pair (line 83) | void run_pair(struct svm_problem * p_km)
function do_cross_validation_with_KM_precalculated (line 100) | void do_cross_validation_with_KM_precalculated( )
FILE: src/linux/kernel_matrix_calculation.c
function ckm (line 8) | void ckm( struct svm_problem *prob, struct svm_problem *pecm, float *gam...
function cal_km (line 129) | void cal_km( struct svm_problem * p_km)
FILE: src/linux/svm-train.c
function print_null (line 9) | void print_null(const char *s) {}
function exit_with_help (line 11) | void exit_with_help()
function exit_input_error (line 45) | void exit_input_error(int line_num)
type svm_parameter (line 55) | struct svm_parameter
type svm_problem (line 56) | struct svm_problem
type svm_model (line 57) | struct svm_model
type svm_node (line 58) | struct svm_node
function main (line 86) | int main(int argc, char **argv)
function do_cross_validation (line 133) | void do_cross_validation()
function parse_command_line (line 172) | void parse_command_line(int argc, char **argv, char *input_file_name, ch...
function read_problem (line 289) | void read_problem(const char *filename)
FILE: src/linux/svm.cpp
function T (line 16) | static inline T min(T x,T y) { return (x<y)?x:y; }
function T (line 19) | static inline T max(T x,T y) { return (x>y)?x:y; }
function swap (line 21) | static inline void swap(T& x, T& y) { T t=x; x=y; y=t; }
function clone (line 22) | static inline void clone(T*& dst, S* src, int n)
function powi (line 27) | static inline double powi(double base, int times)
function print_string_stdout (line 42) | static void print_string_stdout(const char *s)
function info (line 49) | static void info(const char *fmt,...)
function info (line 59) | static void info(const char *fmt,...) {}
class Cache (line 68) | class Cache
type head_t (line 82) | struct head_t
class QMatrix (line 195) | class QMatrix {
class Kernel (line 203) | class Kernel: public QMatrix {
method swap_index (line 216) | virtual void swap_index(int i, int j) const // no so const...
method kernel_linear (line 244) | double kernel_linear(int i, int j) const
method kernel_poly (line 248) | double kernel_poly(int i, int j) const
method kernel_rbf (line 252) | double kernel_rbf(int i, int j) const
method kernel_sigmoid (line 256) | double kernel_sigmoid(int i, int j) const
method kernel_precomputed (line 260) | double kernel_precomputed(int i, int j) const
class Solver (line 452) | class Solver {
method Solver (line 454) | Solver() {}
type SolutionInfo (line 457) | struct SolutionInfo {
method get_C (line 485) | double get_C(int i)
method update_alpha_status (line 489) | void update_alpha_status(int i)
method is_upper_bound (line 497) | bool is_upper_bound(int i) { return alpha_status[i] == UPPER_BOUND; }
method is_lower_bound (line 498) | bool is_lower_bound(int i) { return alpha_status[i] == LOWER_BOUND; }
method is_free (line 499) | bool is_free(int i) { return alpha_status[i] == FREE; }
class Solver_NU (line 1068) | class Solver_NU: public Solver
method Solver_NU (line 1071) | Solver_NU() {}
method Solve (line 1072) | void Solve(int l, const QMatrix& Q, const double *p, const schar *y,
class SVC_Q (line 1325) | class SVC_Q: public Kernel
method SVC_Q (line 1328) | SVC_Q(const svm_problem& prob, const svm_parameter& param, const schar...
method Qfloat (line 1338) | Qfloat *get_Q(int i, int len) const
method swap_index (line 1355) | void swap_index(int i, int j) const
class ONE_CLASS_Q (line 1375) | class ONE_CLASS_Q: public Kernel
method ONE_CLASS_Q (line 1378) | ONE_CLASS_Q(const svm_problem& prob, const svm_parameter& param)
method Qfloat (line 1387) | Qfloat *get_Q(int i, int len) const
method swap_index (line 1404) | void swap_index(int i, int j) const
class SVR_Q (line 1421) | class SVR_Q: public Kernel
method SVR_Q (line 1424) | SVR_Q(const svm_problem& prob, const svm_parameter& param)
method swap_index (line 1446) | void swap_index(int i, int j) const
method Qfloat (line 1453) | Qfloat *get_Q(int i, int len) const
function solve_c_svc (line 1499) | static void solve_c_svc(
function solve_nu_svc (line 1534) | static void solve_nu_svc(
function solve_one_class (line 1589) | static void solve_one_class(
function solve_epsilon_svr (line 1621) | static void solve_epsilon_svr(
function solve_nu_svr (line 1659) | static void solve_nu_svr(
type decision_function (line 1700) | struct decision_function
function decision_function (line 1706) | static decision_function svm_train_one(
function sigmoid_train (line 1764) | static void sigmoid_train(
function sigmoid_predict (line 1877) | static double sigmoid_predict(double decision_value, double A, double B)
function multiclass_probability (line 1888) | static void multiclass_probability(int k, double **r, double *p)
function svm_binary_svc_probability (line 1952) | static void svm_binary_svc_probability(
function svm_svr_probability (line 2047) | static double svm_svr_probability(
function svm_group_classes (line 2081) | static void svm_group_classes(const svm_problem *prob, int *nr_class_ret...
function svm_model (line 2159) | svm_model *svm_train(const svm_problem *prob, const svm_parameter *param)
function svm_cross_validation (line 2419) | void svm_cross_validation(const svm_problem *prob, const svm_parameter *...
function svm_get_svm_type (line 2552) | int svm_get_svm_type(const svm_model *model)
function svm_get_nr_class (line 2557) | int svm_get_nr_class(const svm_model *model)
function svm_get_labels (line 2562) | void svm_get_labels(const svm_model *model, int* label)
function svm_get_sv_indices (line 2569) | void svm_get_sv_indices(const svm_model *model, int* indices)
function svm_get_nr_sv (line 2576) | int svm_get_nr_sv(const svm_model *model)
function svm_get_svr_probability (line 2581) | double svm_get_svr_probability(const svm_model *model)
function svm_predict_values (line 2593) | double svm_predict_values(const svm_model *model, const svm_node *x, dou...
function svm_predict (line 2679) | double svm_predict(const svm_model *model, const svm_node *x)
function svm_predict_probability (line 2694) | double svm_predict_probability(
function svm_save_model (line 2743) | int svm_save_model(const char *model_file_name, const svm_model *model)
function svm_model (line 2873) | svm_model *svm_load_model(const char *model_file_name)
function svm_free_model_content (line 3143) | void svm_free_model_content(svm_model* model_ptr)
function svm_free_and_destroy_model (line 3182) | void svm_free_and_destroy_model(svm_model** model_ptr_ptr)
function svm_destroy_param (line 3192) | void svm_destroy_param(svm_parameter* param)
function svm_check_probability_model (line 3319) | int svm_check_probability_model(const svm_model *model)
function svm_set_print_string_function (line 3327) | void svm_set_print_string_function(void (*print_func)(const char *))
FILE: src/linux/svm.h
type svm_node (line 13) | struct svm_node
type svm_problem (line 19) | struct svm_problem
type svm_node (line 27) | struct svm_node
type svm_problem (line 33) | struct svm_problem
type svm_parameter (line 44) | struct svm_parameter
type svm_model (line 68) | struct svm_model
type svm_model (line 94) | struct svm_model
type svm_problem (line 94) | struct svm_problem
type svm_parameter (line 94) | struct svm_parameter
type svm_problem (line 95) | struct svm_problem
type svm_parameter (line 95) | struct svm_parameter
type svm_model (line 97) | struct svm_model
type svm_model (line 98) | struct svm_model
type svm_model (line 100) | struct svm_model
type svm_model (line 101) | struct svm_model
type svm_model (line 102) | struct svm_model
type svm_model (line 103) | struct svm_model
type svm_model (line 104) | struct svm_model
type svm_model (line 105) | struct svm_model
type svm_model (line 107) | struct svm_model
type svm_node (line 107) | struct svm_node
type svm_model (line 108) | struct svm_model
type svm_node (line 108) | struct svm_node
type svm_model (line 109) | struct svm_model
type svm_node (line 109) | struct svm_node
type svm_model (line 111) | struct svm_model
type svm_model (line 112) | struct svm_model
type svm_parameter (line 113) | struct svm_parameter
type svm_problem (line 115) | struct svm_problem
type svm_parameter (line 115) | struct svm_parameter
type svm_model (line 116) | struct svm_model
FILE: src/windows/libsvm_train_dense_gpu/cross_validation_with_matrix_precomputation.c
function setup_pkm (line 1) | void setup_pkm(struct svm_problem *p_km)
function free_pkm (line 19) | void free_pkm(struct svm_problem *p_km)
function do_crossvalidation (line 33) | double do_crossvalidation(struct svm_problem * p_km)
function run_pair (line 83) | void run_pair(struct svm_problem * p_km)
function do_cross_validation_with_KM_precalculated (line 100) | void do_cross_validation_with_KM_precalculated( )
FILE: src/windows/libsvm_train_dense_gpu/kernel_matrix_calculation.c
function ckm (line 8) | void ckm( struct svm_problem *prob, struct svm_problem *pecm, float *gam...
function cal_km (line 129) | void cal_km( struct svm_problem * p_km)
FILE: src/windows/libsvm_train_dense_gpu/svm-train.c
function print_null (line 9) | void print_null(const char *s) {}
function exit_with_help (line 11) | void exit_with_help()
function exit_input_error (line 45) | void exit_input_error(int line_num)
type svm_parameter (line 55) | struct svm_parameter
type svm_problem (line 56) | struct svm_problem
type svm_model (line 57) | struct svm_model
type svm_node (line 58) | struct svm_node
function main (line 86) | int main(int argc, char **argv)
function do_cross_validation (line 133) | void do_cross_validation()
function parse_command_line (line 172) | void parse_command_line(int argc, char **argv, char *input_file_name, ch...
function read_problem (line 289) | void read_problem(const char *filename)
FILE: src/windows/libsvm_train_dense_gpu/svm.cpp
function T (line 16) | static inline T min(T x,T y) { return (x<y)?x:y; }
function T (line 19) | static inline T max(T x,T y) { return (x>y)?x:y; }
function swap (line 21) | static inline void swap(T& x, T& y) { T t=x; x=y; y=t; }
function clone (line 22) | static inline void clone(T*& dst, S* src, int n)
function powi (line 27) | static inline double powi(double base, int times)
function print_string_stdout (line 42) | static void print_string_stdout(const char *s)
function info (line 49) | static void info(const char *fmt,...)
function info (line 59) | static void info(const char *fmt,...) {}
class Cache (line 68) | class Cache
type head_t (line 82) | struct head_t
class QMatrix (line 195) | class QMatrix {
class Kernel (line 203) | class Kernel: public QMatrix {
method swap_index (line 216) | virtual void swap_index(int i, int j) const // no so const...
method kernel_linear (line 244) | double kernel_linear(int i, int j) const
method kernel_poly (line 248) | double kernel_poly(int i, int j) const
method kernel_rbf (line 252) | double kernel_rbf(int i, int j) const
method kernel_sigmoid (line 256) | double kernel_sigmoid(int i, int j) const
method kernel_precomputed (line 260) | double kernel_precomputed(int i, int j) const
class Solver (line 452) | class Solver {
method Solver (line 454) | Solver() {}
type SolutionInfo (line 457) | struct SolutionInfo {
method get_C (line 485) | double get_C(int i)
method update_alpha_status (line 489) | void update_alpha_status(int i)
method is_upper_bound (line 497) | bool is_upper_bound(int i) { return alpha_status[i] == UPPER_BOUND; }
method is_lower_bound (line 498) | bool is_lower_bound(int i) { return alpha_status[i] == LOWER_BOUND; }
method is_free (line 499) | bool is_free(int i) { return alpha_status[i] == FREE; }
class Solver_NU (line 1068) | class Solver_NU: public Solver
method Solver_NU (line 1071) | Solver_NU() {}
method Solve (line 1072) | void Solve(int l, const QMatrix& Q, const double *p, const schar *y,
class SVC_Q (line 1325) | class SVC_Q: public Kernel
method SVC_Q (line 1328) | SVC_Q(const svm_problem& prob, const svm_parameter& param, const schar...
method Qfloat (line 1338) | Qfloat *get_Q(int i, int len) const
method swap_index (line 1355) | void swap_index(int i, int j) const
class ONE_CLASS_Q (line 1375) | class ONE_CLASS_Q: public Kernel
method ONE_CLASS_Q (line 1378) | ONE_CLASS_Q(const svm_problem& prob, const svm_parameter& param)
method Qfloat (line 1387) | Qfloat *get_Q(int i, int len) const
method swap_index (line 1404) | void swap_index(int i, int j) const
class SVR_Q (line 1421) | class SVR_Q: public Kernel
method SVR_Q (line 1424) | SVR_Q(const svm_problem& prob, const svm_parameter& param)
method swap_index (line 1446) | void swap_index(int i, int j) const
method Qfloat (line 1453) | Qfloat *get_Q(int i, int len) const
function solve_c_svc (line 1499) | static void solve_c_svc(
function solve_nu_svc (line 1534) | static void solve_nu_svc(
function solve_one_class (line 1589) | static void solve_one_class(
function solve_epsilon_svr (line 1621) | static void solve_epsilon_svr(
function solve_nu_svr (line 1659) | static void solve_nu_svr(
type decision_function (line 1700) | struct decision_function
function decision_function (line 1706) | static decision_function svm_train_one(
function sigmoid_train (line 1764) | static void sigmoid_train(
function sigmoid_predict (line 1877) | static double sigmoid_predict(double decision_value, double A, double B)
function multiclass_probability (line 1888) | static void multiclass_probability(int k, double **r, double *p)
function svm_binary_svc_probability (line 1952) | static void svm_binary_svc_probability(
function svm_svr_probability (line 2047) | static double svm_svr_probability(
function svm_group_classes (line 2081) | static void svm_group_classes(const svm_problem *prob, int *nr_class_ret...
function svm_model (line 2159) | svm_model *svm_train(const svm_problem *prob, const svm_parameter *param)
function svm_cross_validation (line 2419) | void svm_cross_validation(const svm_problem *prob, const svm_parameter *...
function svm_get_svm_type (line 2552) | int svm_get_svm_type(const svm_model *model)
function svm_get_nr_class (line 2557) | int svm_get_nr_class(const svm_model *model)
function svm_get_labels (line 2562) | void svm_get_labels(const svm_model *model, int* label)
function svm_get_sv_indices (line 2569) | void svm_get_sv_indices(const svm_model *model, int* indices)
function svm_get_nr_sv (line 2576) | int svm_get_nr_sv(const svm_model *model)
function svm_get_svr_probability (line 2581) | double svm_get_svr_probability(const svm_model *model)
function svm_predict_values (line 2593) | double svm_predict_values(const svm_model *model, const svm_node *x, dou...
function svm_predict (line 2679) | double svm_predict(const svm_model *model, const svm_node *x)
function svm_predict_probability (line 2694) | double svm_predict_probability(
function svm_save_model (line 2743) | int svm_save_model(const char *model_file_name, const svm_model *model)
function svm_model (line 2873) | svm_model *svm_load_model(const char *model_file_name)
function svm_free_model_content (line 3143) | void svm_free_model_content(svm_model* model_ptr)
function svm_free_and_destroy_model (line 3182) | void svm_free_and_destroy_model(svm_model** model_ptr_ptr)
function svm_destroy_param (line 3192) | void svm_destroy_param(svm_parameter* param)
function svm_check_probability_model (line 3319) | int svm_check_probability_model(const svm_model *model)
function svm_set_print_string_function (line 3327) | void svm_set_print_string_function(void (*print_func)(const char *))
FILE: src/windows/libsvm_train_dense_gpu/svm.h
type svm_node (line 13) | struct svm_node
type svm_problem (line 19) | struct svm_problem
type svm_node (line 27) | struct svm_node
type svm_problem (line 33) | struct svm_problem
type svm_parameter (line 44) | struct svm_parameter
type svm_model (line 68) | struct svm_model
type svm_model (line 94) | struct svm_model
type svm_problem (line 94) | struct svm_problem
type svm_parameter (line 94) | struct svm_parameter
type svm_problem (line 95) | struct svm_problem
type svm_parameter (line 95) | struct svm_parameter
type svm_model (line 97) | struct svm_model
type svm_model (line 98) | struct svm_model
type svm_model (line 100) | struct svm_model
type svm_model (line 101) | struct svm_model
type svm_model (line 102) | struct svm_model
type svm_model (line 103) | struct svm_model
type svm_model (line 104) | struct svm_model
type svm_model (line 105) | struct svm_model
type svm_model (line 107) | struct svm_model
type svm_node (line 107) | struct svm_node
type svm_model (line 108) | struct svm_model
type svm_node (line 108) | struct svm_node
type svm_model (line 109) | struct svm_model
type svm_node (line 109) | struct svm_node
type svm_model (line 111) | struct svm_model
type svm_model (line 112) | struct svm_model
type svm_parameter (line 113) | struct svm_parameter
type svm_problem (line 115) | struct svm_problem
type svm_parameter (line 115) | struct svm_parameter
type svm_model (line 116) | struct svm_model
Condensed preview — 59 files, each showing path, character count, and a content snippet. Download the .json file or copy for the full structured content (699K chars).
[
{
"path": "README.md",
"chars": 5029,
"preview": "CUDA: GPU-accelerated LIBSVM\n====\n**LIBSVM Accelerated with GPU using the CUDA Framework**\n\nGPU-accelerated LIBSVM is a "
},
{
"path": "binaries/linux/COPYRIGHT",
"chars": 1497,
"preview": "\nCopyright (c) 2000-2013 Chih-Chung Chang and Chih-Jen Lin\nAll rights reserved.\n\nRedistribution and use in source and bi"
},
{
"path": "binaries/linux/FAQ.html",
"chars": 73848,
"preview": "\n\n<html>\n<head>\n<title>LIBSVM FAQ</title>\n</head>\n<body bgcolor=\"#ffffcc\">\n\n<a name=\"_TOP\"><b><h1><a\nhref=http://www.csi"
},
{
"path": "binaries/linux/README",
"chars": 28271,
"preview": "Libsvm is a simple, easy-to-use, and efficient software for SVM\nclassification and regression. It solves C-SVM classific"
},
{
"path": "binaries/linux/README-GPU",
"chars": 1305,
"preview": "GPU-Accelerated LIBSVM is exploiting the GPU, using the CUDA interface, to\nspeed-up the training process. This package c"
},
{
"path": "binaries/linux/tools/README",
"chars": 7033,
"preview": "This directory includes some useful codes:\n\n1. subset selection tools.\n2. parameter selection tools.\n3. LIBSVM format ch"
},
{
"path": "binaries/linux/tools/checkdata.py",
"chars": 2479,
"preview": "#!/usr/bin/env python\n\n#\n# A format checker for LIBSVM\n#\n\n#\n# Copyright (c) 2007, Rong-En Fan\n#\n# All rights reserved.\n#"
},
{
"path": "binaries/linux/tools/easy.py",
"chars": 2707,
"preview": "#!/usr/bin/env python\n\nimport sys\nimport os\nfrom subprocess import *\n\nif len(sys.argv) <= 1:\n\tprint('Usage: {0} training"
},
{
"path": "binaries/linux/tools/grid.py",
"chars": 15316,
"preview": "#!/usr/bin/env python\n__all__ = ['find_parameters']\n\nimport os, sys, traceback, getpass, time, re\nfrom threading import "
},
{
"path": "binaries/linux/tools/subset.py",
"chars": 3202,
"preview": "#!/usr/bin/env python\n\nimport os, sys, math, random\nfrom collections import defaultdict\n\nif sys.version_info[0] >= 3:\n\tx"
},
{
"path": "binaries/windows/x64/COPYRIGHT",
"chars": 1497,
"preview": "\nCopyright (c) 2000-2013 Chih-Chung Chang and Chih-Jen Lin\nAll rights reserved.\n\nRedistribution and use in source and bi"
},
{
"path": "binaries/windows/x64/FAQ.html",
"chars": 73848,
"preview": "\n\n<html>\n<head>\n<title>LIBSVM FAQ</title>\n</head>\n<body bgcolor=\"#ffffcc\">\n\n<a name=\"_TOP\"><b><h1><a\nhref=http://www.csi"
},
{
"path": "binaries/windows/x64/README",
"chars": 28271,
"preview": "Libsvm is a simple, easy-to-use, and efficient software for SVM\nclassification and regression. It solves C-SVM classific"
},
{
"path": "binaries/windows/x64/README-GPU",
"chars": 1313,
"preview": "GPU-Accelerated LIBSVM is exploiting the GPU, using the CUDA interface, to\nspeed-up the training process. This package c"
},
{
"path": "binaries/windows/x64/tools/README",
"chars": 7033,
"preview": "This directory includes some useful codes:\n\n1. subset selection tools.\n2. parameter selection tools.\n3. LIBSVM format ch"
},
{
"path": "binaries/windows/x64/tools/checkdata.py",
"chars": 2479,
"preview": "#!/usr/bin/env python\n\n#\n# A format checker for LIBSVM\n#\n\n#\n# Copyright (c) 2007, Rong-En Fan\n#\n# All rights reserved.\n#"
},
{
"path": "binaries/windows/x64/tools/easy.py",
"chars": 2724,
"preview": "#!/usr/bin/env python\n\nimport sys\nimport os\nfrom subprocess import *\n\nif len(sys.argv) <= 1:\n\tprint('Usage: {0} training"
},
{
"path": "binaries/windows/x64/tools/grid.py",
"chars": 15317,
"preview": "#!/usr/bin/env python\n__all__ = ['find_parameters']\n\nimport os, sys, traceback, getpass, time, re\nfrom threading import "
},
{
"path": "binaries/windows/x64/tools/subset.py",
"chars": 3202,
"preview": "#!/usr/bin/env python\n\nimport os, sys, math, random\nfrom collections import defaultdict\n\nif sys.version_info[0] >= 3:\n\tx"
},
{
"path": "binaries/windows/x86/COPYRIGHT",
"chars": 1497,
"preview": "\nCopyright (c) 2000-2013 Chih-Chung Chang and Chih-Jen Lin\nAll rights reserved.\n\nRedistribution and use in source and bi"
},
{
"path": "binaries/windows/x86/FAQ.html",
"chars": 73848,
"preview": "\n\n<html>\n<head>\n<title>LIBSVM FAQ</title>\n</head>\n<body bgcolor=\"#ffffcc\">\n\n<a name=\"_TOP\"><b><h1><a\nhref=http://www.csi"
},
{
"path": "binaries/windows/x86/README",
"chars": 28271,
"preview": "Libsvm is a simple, easy-to-use, and efficient software for SVM\nclassification and regression. It solves C-SVM classific"
},
{
"path": "binaries/windows/x86/README-GPU",
"chars": 1313,
"preview": "GPU-Accelerated LIBSVM is exploiting the GPU, using the CUDA interface, to\nspeed-up the training process. This package c"
},
{
"path": "binaries/windows/x86/tools/README",
"chars": 7033,
"preview": "This directory includes some useful codes:\n\n1. subset selection tools.\n2. parameter selection tools.\n3. LIBSVM format ch"
},
{
"path": "binaries/windows/x86/tools/checkdata.py",
"chars": 2479,
"preview": "#!/usr/bin/env python\n\n#\n# A format checker for LIBSVM\n#\n\n#\n# Copyright (c) 2007, Rong-En Fan\n#\n# All rights reserved.\n#"
},
{
"path": "binaries/windows/x86/tools/easy.py",
"chars": 2724,
"preview": "#!/usr/bin/env python\n\nimport sys\nimport os\nfrom subprocess import *\n\nif len(sys.argv) <= 1:\n\tprint('Usage: {0} training"
},
{
"path": "binaries/windows/x86/tools/grid.py",
"chars": 15317,
"preview": "#!/usr/bin/env python\n__all__ = ['find_parameters']\n\nimport os, sys, traceback, getpass, time, re\nfrom threading import "
},
{
"path": "binaries/windows/x86/tools/subset.py",
"chars": 3202,
"preview": "#!/usr/bin/env python\n\nimport os, sys, math, random\nfrom collections import defaultdict\n\nif sys.version_info[0] >= 3:\n\tx"
},
{
"path": "src/linux/COPYRIGHT",
"chars": 1497,
"preview": "\nCopyright (c) 2000-2010 Chih-Chung Chang and Chih-Jen Lin\nAll rights reserved.\n\nRedistribution and use in source and bi"
},
{
"path": "src/linux/Makefile",
"chars": 4628,
"preview": "################################################################################\n#\n# Copyright 1993-2013 NVIDIA Corporat"
},
{
"path": "src/linux/README",
"chars": 26529,
"preview": "Libsvm is a simple, easy-to-use, and efficient software for SVM\nclassification and regression. It solves C-SVM classific"
},
{
"path": "src/linux/README-GPU",
"chars": 2365,
"preview": "GPU-Accelerated LIBSVM is exploiting the GPU, using the CUDA interface, to\nspeed-up the training process. This package c"
},
{
"path": "src/linux/cross_validation_with_matrix_precomputation.c",
"chars": 1856,
"preview": "void setup_pkm(struct svm_problem *p_km)\n{\n\n\tint i;\n\n\tp_km->l = prob.l;\n\tp_km->x = Malloc(struct svm_node,p_km->l);\n\tp_k"
},
{
"path": "src/linux/findcudalib.mk",
"chars": 7799,
"preview": "################################################################################\n#\n# Copyright 1993-2013 NVIDIA Corporat"
},
{
"path": "src/linux/kernel_matrix_calculation.c",
"chars": 2879,
"preview": "#include \"/usr/local/cuda-5.5/include/cuda_runtime.h\"\n#include \"/usr/local/cuda-5.5/include/cublas_v2.h\"\n\n// Scalars\ncon"
},
{
"path": "src/linux/readme.txt",
"chars": 1310,
"preview": "Instructions to compile Linux GPU-Accelerated LIBSVM\n\n1. Install the NVIDIA drivers, CUDA toolkit and GPU Computing SDK "
},
{
"path": "src/linux/svm-train.c",
"chars": 11021,
"preview": "#include <stdio.h>\n#include <stdlib.h>\n#include <string.h>\n#include <ctype.h>\n#include <errno.h>\n#include \"svm.h\"\n#defin"
},
{
"path": "src/linux/svm.cpp",
"chars": 68116,
"preview": "#include <math.h>\n#include <stdio.h>\n#include <stdlib.h>\n#include <ctype.h>\n#include <float.h>\n#include <string.h>\n#incl"
},
{
"path": "src/linux/svm.h",
"chars": 3619,
"preview": "#ifndef _LIBSVM_H\n#define _LIBSVM_H\n#define _DENSE_REP\n#define LIBSVM_VERSION 317\n\n#ifdef __cplusplus\nextern \"C\" {\n#endi"
},
{
"path": "src/windows/README-GPU",
"chars": 1461,
"preview": "GPU-Accelerated LIBSVM is exploiting the GPU, using the CUDA interface, to\nspeed-up the training process. This package c"
},
{
"path": "src/windows/libsvm_train_dense_gpu/cross_validation_with_matrix_precomputation.c",
"chars": 1856,
"preview": "void setup_pkm(struct svm_problem *p_km)\n{\n\n\tint i;\n\n\tp_km->l = prob.l;\n\tp_km->x = Malloc(struct svm_node,p_km->l);\n\tp_k"
},
{
"path": "src/windows/libsvm_train_dense_gpu/kernel_matrix_calculation.c",
"chars": 2822,
"preview": "#include <cuda_runtime.h>\n#include \"cublas_v2.h\"\n\n// Scalars\nconst float alpha = 1;\nconst float beta = 0;\n\nvoid ckm( str"
},
{
"path": "src/windows/libsvm_train_dense_gpu/libsvm_train_dense_gpu.vcxproj",
"chars": 13544,
"preview": "<?xml version=\"1.0\" encoding=\"utf-8\"?>\n<Project DefaultTargets=\"Build\" ToolsVersion=\"4.0\" xmlns=\"http://schemas.microso"
},
{
"path": "src/windows/libsvm_train_dense_gpu/libsvm_train_dense_gpu.vcxproj.filters",
"chars": 1441,
"preview": "<?xml version=\"1.0\" encoding=\"utf-8\"?>\n<Project ToolsVersion=\"4.0\" xmlns=\"http://schemas.microsoft.com/developer/msbuil"
},
{
"path": "src/windows/libsvm_train_dense_gpu/libsvm_train_dense_gpu.vcxproj.user",
"chars": 139,
"preview": "<?xml version=\"1.0\" encoding=\"utf-8\"?>\n<Project ToolsVersion=\"4.0\" xmlns=\"http://schemas.microsoft.com/developer/msbuil"
},
{
"path": "src/windows/libsvm_train_dense_gpu/svm-train.c",
"chars": 11021,
"preview": "#include <stdio.h>\n#include <stdlib.h>\n#include <string.h>\n#include <ctype.h>\n#include <errno.h>\n#include \"svm.h\"\n#defin"
},
{
"path": "src/windows/libsvm_train_dense_gpu/svm.cpp",
"chars": 68116,
"preview": "#include <math.h>\n#include <stdio.h>\n#include <stdlib.h>\n#include <ctype.h>\n#include <float.h>\n#include <string.h>\n#incl"
},
{
"path": "src/windows/libsvm_train_dense_gpu/svm.h",
"chars": 3619,
"preview": "#ifndef _LIBSVM_H\n#define _LIBSVM_H\n#define _DENSE_REP\n#define LIBSVM_VERSION 317\n\n#ifdef __cplusplus\nextern \"C\" {\n#endi"
},
{
"path": "src/windows/libsvm_train_dense_gpu.sln",
"chars": 1259,
"preview": "\nMicrosoft Visual Studio Solution File, Format Version 11.00\n# Visual Studio 2010\nProject(\"{8BC9CEB8-8B4A-11D0-8D11-00A"
}
]
// ... and 10 more files (download for full content)
About this extraction
This page contains the full source code of the MKLab-ITI/CUDA GitHub repository, extracted and formatted as plain text for AI agents and large language models (LLMs). The extraction includes 59 files (33.1 MB), approximately 191.0k tokens, and a symbol index with 329 extracted functions, classes, methods, constants, and types. Use this with OpenClaw, Claude, ChatGPT, Cursor, Windsurf, or any other AI tool that accepts text input. You can copy the full output to your clipboard or download it as a .txt file.
Extracted by GitExtract — free GitHub repo to text converter for AI. Built by Nikandr Surkov.