| MorphL Platform
The backbone of the platform is the MorphL Orchestrator, that sets up the Big Data techstack required for running pipelines for data ingestion, models training and generating predictions.
|
 | MorphL Integrations
We integrate with various data sources. At the moment, we support Google Analytics, Google Analytics 360, BigQuery, Google Cloud Storage and AWS S3. |
 | MorphL Predictive Models
We're utilizing open-source machine learning algorithms to build predictive models which are then used to develop predictive applications. |
 | MorphL Predictions API
All predictions are available via a REST API, which makes it easier for software developers to incorporate AI capabilities within their digital products or services.
|
The setup guide is available [here](orchestrator/).
# Architecture
The MorphL Platform consists of two main components:
- **[MorphL Platform Orchestrator](orchestrator/)** - This is the backbone of the platform. It sets up the infrastructure required for running pipelines for each model.
- **[MorphL Pipelines](pipelines/)** - Consists of various Python scripts, required for retrieving data from various sources, pre-processing, training a model and generating predictions.
---
The code that you'll find in this repository is a mirror that we use for making releases. If you want to contribute to a pipeline or create a new model, please open a pull request in the corresponding repository from the [MorphL-AI organization](https://github.com/Morphl-AI).
You can read more about MorphL here: https://morphl.io. Follow us on Twitter: https://twitter.com/morphlio. Join our Slack community and chat with other developers: http://bit.ly/morphl-slack
# MorphL Cloud
On-premises, Cloud or Hybrid. For companies that want to AI-enhance their digital products & services without the hassle of dealing with a Big Data & Machine Learning infrastructure, we offer several deployment options that best suits your business needs and budget.
For enterprise sales or partnerships please contact us [here](https://morphl.io/company/contact.html) or at contact [at] morphl.io.
## License
Licensed under the [Apache-2.0 License](https://opensource.org/licenses/Apache2.0).
================================================
FILE: orchestrator/README.md
================================================
# MorphL Platform Orchestrator
The MorphL Orchestrator is the backbone of the MorphL platform. It sets up the infrastructure and software that are necessary for running the MorphL platform. It consists of 3 pipelines:
- **Ingestion Pipeline** - It runs a series of connectors responsible for gathering data from various APIs (Google Analytics, Mixpanel, Google Cloud Storage, etc.) and save it into Cassandra tables.
- **Training Pipeline** - Consists of pre-processors (responsible for cleaning, formatting, deduplicating, normalizing and transforming data) and model training.
- **Prediction Pipeline** - It generates predictions based on the model that was trained. It is triggered at the final step of the ingestion pipeline through a preflight check.
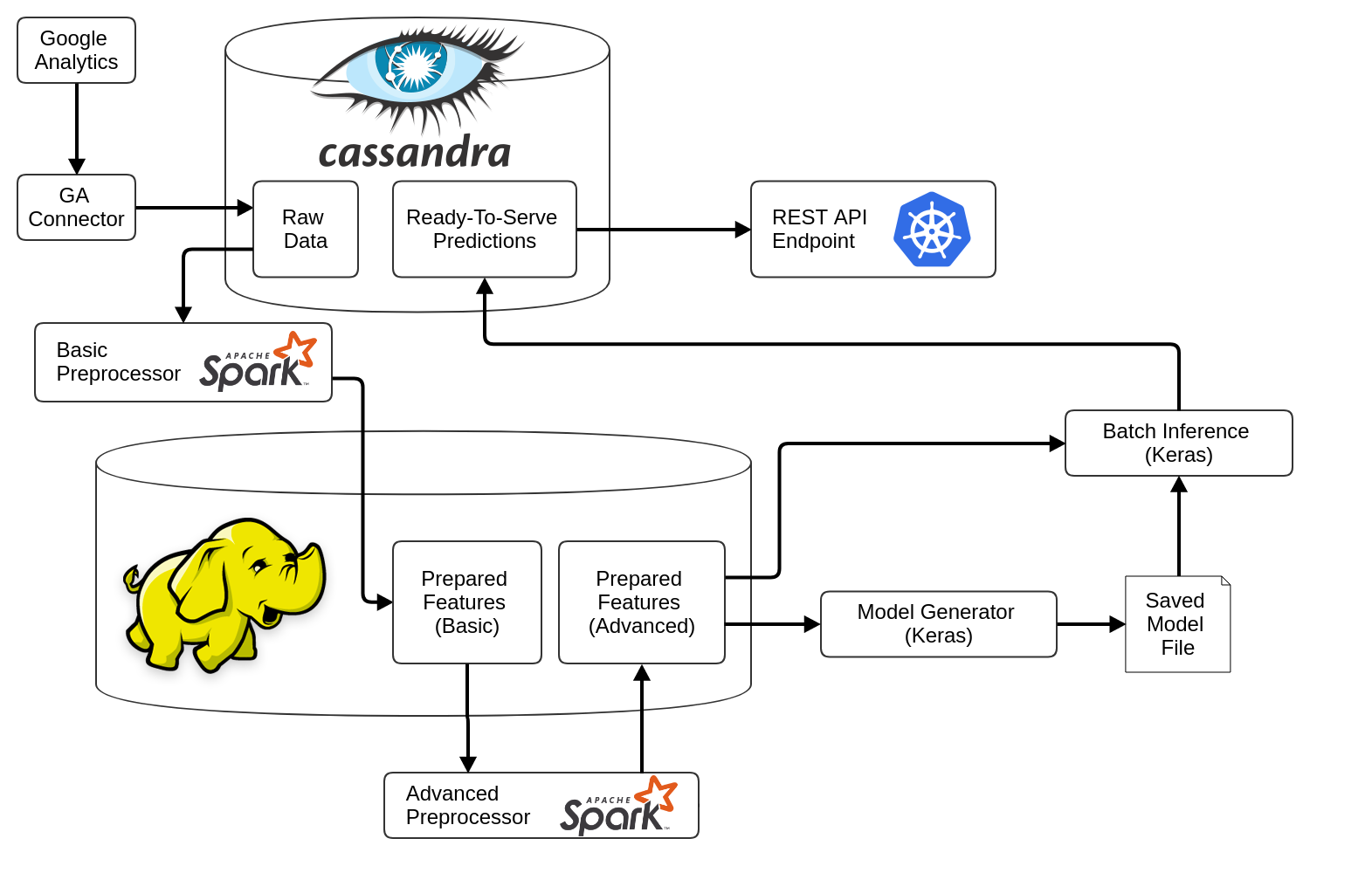
The pipelines are set up using [Apache Airflow](https://github.com/apache/incubator-airflow). Below you can see a diagram of the platform's architecture: