Showing preview only (1,068K chars total). Download the full file or copy to clipboard to get everything.
Repository: cjekel/piecewise_linear_fit_py
Branch: master
Commit: 45441d8b4f18
Files: 112
Total size: 1.0 MB
Directory structure:
gitextract_642gn0fy/
├── .github/
│ └── workflows/
│ ├── ci.yml
│ └── cron.yml
├── .gitignore
├── CHANGELOG.md
├── LICENSE
├── MANIFEST.in
├── README.md
├── README.rst
├── convert_README_to_RST.sh
├── docs/
│ ├── .buildinfo
│ ├── .nojekyll
│ ├── _sources/
│ │ ├── about.rst.txt
│ │ ├── examples.rst.txt
│ │ ├── how_it_works.rst.txt
│ │ ├── index.rst.txt
│ │ ├── installation.rst.txt
│ │ ├── license.rst.txt
│ │ ├── modules.rst.txt
│ │ ├── pwlf.rst.txt
│ │ ├── requirements.rst.txt
│ │ └── stubs/
│ │ └── pwlf.PiecewiseLinFit.rst.txt
│ ├── _static/
│ │ ├── alabaster.css
│ │ ├── basic.css
│ │ ├── custom.css
│ │ ├── doctools.js
│ │ ├── documentation_options.js
│ │ ├── language_data.js
│ │ ├── pygments.css
│ │ ├── searchtools.js
│ │ └── sphinx_highlight.js
│ ├── about.html
│ ├── examples.html
│ ├── genindex.html
│ ├── how_it_works.html
│ ├── index.html
│ ├── installation.html
│ ├── license.html
│ ├── modules.html
│ ├── objects.inv
│ ├── pwlf.html
│ ├── requirements.html
│ ├── search.html
│ ├── searchindex.js
│ └── stubs/
│ └── pwlf.PiecewiseLinFit.html
├── doctrees/
│ ├── about.doctree
│ ├── environment.pickle
│ ├── examples.doctree
│ ├── how_it_works.doctree
│ ├── index.doctree
│ ├── installation.doctree
│ ├── license.doctree
│ ├── modules.doctree
│ ├── pwlf.doctree
│ ├── requirements.doctree
│ └── stubs/
│ └── pwlf.PiecewiseLinFit.doctree
├── examples/
│ ├── EGO_integer_only.ipynb
│ ├── README.md
│ ├── README.rst
│ ├── ex_data/
│ │ └── saved_parameters.npy
│ ├── experiment_with_batch_process.py.ipynb
│ ├── fitForSpecifiedNumberOfLineSegments.py
│ ├── fitForSpecifiedNumberOfLineSegments_passDiffEvoKeywords.py
│ ├── fitForSpecifiedNumberOfLineSegments_standard_deviation.py
│ ├── fitWithKnownLineSegmentLocations.py
│ ├── fit_begin_and_end.py
│ ├── min_length_demo.ipynb
│ ├── mixed_degree.py
│ ├── mixed_degree_forcing_slope.py
│ ├── model_persistence.py
│ ├── model_persistence_prediction.py
│ ├── prediction_variance.py
│ ├── prediction_variance_degree2.py
│ ├── robust_regression.py
│ ├── run_opt_to_find_best_number_of_line_segments.py
│ ├── sineWave.py
│ ├── sineWave_custom_opt_bounds.py
│ ├── sineWave_degrees.py
│ ├── sineWave_time_compare.py
│ ├── slope_constraint_demo.ipynb
│ ├── stack_overflow_example.py
│ ├── standard_errrors_and_p-values.py
│ ├── standard_errrors_and_p-values_non-linear.py
│ ├── test0.py
│ ├── test_for_model_significance.py
│ ├── test_if_breaks_exact.py
│ ├── tf/
│ │ ├── fit_begin_and_end.py
│ │ ├── sine_benchmark_fixed_20_break_points.py
│ │ ├── sine_benchmark_six_segments.py
│ │ └── test_fit.py
│ ├── understanding_higher_degrees/
│ │ └── polynomials_in_pwlf.ipynb
│ ├── useCustomOptimizationRoutine.py
│ ├── weighted_least_squares_ex.py
│ └── weighted_least_squares_ex_stats.py
├── pwlf/
│ ├── __init__.py
│ ├── pwlf.py
│ └── version.py
├── setup.py
├── sphinx_build_script.sh
├── sphinxdocs/
│ ├── Makefile
│ ├── make.bat
│ └── source/
│ ├── about.rst
│ ├── conf.py
│ ├── examples.rst
│ ├── how_it_works.rst
│ ├── index.rst
│ ├── installation.rst
│ ├── license.rst
│ ├── modules.rst
│ ├── pwlf.rst
│ └── requirements.rst
└── tests/
├── __init__.py
└── tests.py
================================================
FILE CONTENTS
================================================
================================================
FILE: .github/workflows/ci.yml
================================================
name: pwlf ci
on:
push:
jobs:
build:
runs-on: ubuntu-22.04
strategy:
matrix:
python-version: ['3.10', '3.11', '3.12', '3.13']
steps:
- uses: actions/checkout@v2
- name: Set up Python ${{ matrix.python-version }}
uses: actions/setup-python@v2
with:
python-version: ${{ matrix.python-version }}
- name: Install dependencies
run: |
python -m pip install flake8 coverage pytest pytest-cov
- name: Install pwlf
run: |
python -m pip install . --no-cache-dir
- name: Lint with flake8
run: |
flake8 pwlf
flake8 tests/tests.py
- name: Test with pytest
run: |
pytest --cov=pwlf --cov-report=xml -p no:warnings tests/tests.py
- name: Upload coverage to Codecov
uses: codecov/codecov-action@v1
with:
token: ${{ secrets.CODECOV_TOKEN }}
file: ./coverage.xml
directory: ./coverage/reports/
flags: unittests
env_vars: OS,PYTHON
name: codecov-umbrella
fail_ci_if_error: false
verbose: false
================================================
FILE: .github/workflows/cron.yml
================================================
name: pwlf cron
on:
schedule:
# Also run every 24 hours
- cron: '* */24 * * *'
jobs:
build:
runs-on: ubuntu-22.04
strategy:
matrix:
python-version: ['3.10', '3.11', '3.12', '3.13']
steps:
- uses: actions/checkout@v2
- name: Set up Python ${{ matrix.python-version }}
uses: actions/setup-python@v2
with:
python-version: ${{ matrix.python-version }}
- name: Install dependencies
run: |
python -m pip install flake8 coverage pytest pytest-cov
- name: Install pwlf
run: |
python -m pip install . --no-cache-dir
- name: Lint with flake8
run: |
flake8 pwlf
- name: Test with pytest
run: |
pytest --cov=pwlf --cov-report=xml -p no:warnings tests/tests.py
================================================
FILE: .gitignore
================================================
*.pyc
.coverage
MANIFEST
/build/
setup.cfg
# Setuptools distribution folder.
/dist/
# Python egg metadata, regenerated from source files by setuptools.
/*.egg-info
# pip stuff
/temp/
# vscode folder
.vscode*
# vim temp file
*~
*.swp
*.un~
# temporary documentation folder
tempdocs
tempdocs/*
sphinxdocs/build
sphinxdocs/build/*
================================================
FILE: CHANGELOG.md
================================================
# Changelog
All notable changes to this project will be documented in this file.
The format is based on [Keep a Changelog](https://keepachangelog.com/en/1.0.0/),
and this project adheres to [Semantic Versioning](https://semver.org/spec/v2.0.0.html).
## [2.5.2] - 2025-07-26
### Changed
- Minor typo in documentation. Thanks to [Berend-ASML](https://github.com/Berend-ASML) for the fix!
## [2.5.1] - 2025-02-23
### Changed
- Only test float128 support if numpy allows you to create float128 numbers. This should fix issues on aarch64 builds that do not support float128 or longdouble.
## [2.5.0] - 2025-02-19
### Changed
- remove pyDOE as a dependency in favor of scipy's own latin hypercube sampling. This change will effect results of the `fitfast` method, where previous versions of the `fitfast` results will no longer be reproducible with this version.
## [2.4.0] - 2024-12-31
### Added
- Added support for mixing degree fits. You may now specify `degree=[0,1,0]` to fit a constant, then linear, then constant line. These are discontinuous piecewise linear. Currently this only supports mixing degrees of 0 and 1. Thanks to [wonch002](https://github.com/wonch002) for resurrecting an old branch.
### Changed
- fixed a bug where `seed=0` would net get set. Thanks to [filippobistaffa](https://github.com/filippobistaffa) from https://github.com/cjekel/piecewise_linear_fit_py/pull/118 .
## [2.3.0] - 2024-10-26
### Changed
- You can now force fits of one line segment to go through a particular point `mypwlf.fit(1, x_c, y_c)`. Thanks to [fredrikhellman](https://github.com/fredrikhellman).
- Update docs to sphinx 8 (from 4)
- Update ci version to `python==3.9`, `python==3.10`, `python==3.11`, and `python==3.12`
## [2.2.1] - 2022-05-07
### Added
- You can now perform `fit` and `fitfast` for one line segment!
## [2.2.0] - 2022-05-01
### Added
- Now you can specify a numpy.random.seed to use on init to get reproducible results from the `fit` and `fitfast` methods. Simply specify an integer seed number like so `pwlf.PiecewiseLinFit(x, y, seed=123)`. Note this hijacks your current random seed. By default, not random seed is specified.
- Python 3.9 is now part of the ci
### Changed
- Add flake8 checks to tests
- Two tests were not being checked because the method did not start with `test`
## [2.1.0] - 2022-03-31
### Changed
- All instances of `linalg.inv` now use `linalg.pinv`. All APIs are still the same, but this is potentially a backwards breaking change as previous results may be different from new results. This will mainly affect standard error calculations.
- Previously `calc_slopes` was called after every least squares fit in optimization routines trying to find breakpoints. This would occasionally raise a numpy `RuntimeWarning` if two breakpoints were the same, or if a breakpoint was on the boundary. Now `calc_slopes` is not called during experimental breakpoint calculation, which should no longer raise this warning for most users. Slopes will still be calculated once optimal breakpoints are found!
## [2.0.5] - 2021-12-04
### Added
- conda forge installs are now officially mentioned on readme and in documentation
## [2.0.4] - 2020-08-27
### Deprecated
- Python 2.7 will not be supported in future releases
- Python 3.5 reaches [end-of-life](https://devguide.python.org/#status-of-python-branches) on 2020-09-13, and will not be supported in future releases
### Added
- ```python_requires``` in setup.py
## [2.0.3] - 2020-06-26
### Changed
- version handling does not require any dependencies
### Removed
- importlib requirement
## [2.0.2] - 2020-05-25
### Changed
- Fixed an encoding bug that would not let pwlf install on windows. Thanks to h-vetinari for the [PR](https://github.com/cjekel/piecewise_linear_fit_py/pull/71)!
## [2.0.1] - 2020-05-24
### Changed
- Removed setuptools for importlib single source versioning
### Added
- Requirement for importlib-metadata if Python version is less than 3.8
### Removed
- Requriement for setuptools
## [2.0.0] - 2020-04-02
### Added
- Added supports for pwlf to fit to weighted data sets! Check out [this example](https://github.com/cjekel/piecewise_linear_fit_py/tree/master/examples#weighted-least-squares-fit).
### Changed
- Setup.py now grabs markdown file for long description
### Removed
- Tensorflow support has been removed. It hasn't been updated in a long time. If you still require this object, check out [pwlftf](https://github.com/cjekel/piecewise_linear_fit_py_tf)
## [1.1.7] - 2020-02-05
### Changed
- Minimum SciPy version is now 1.2.0 because of issues with MacOS and the old SciPy versions. See issue https://github.com/cjekel/piecewise_linear_fit_py/issues/40 and thanks to bezineb5 !
## [1.1.6] - 2020-01-22
### Changed
- Single source version now found in setup.py instead of pwlf/VERSION see issue https://github.com/cjekel/piecewise_linear_fit_py/issues/53
- New setuptools requirement to handle new version file
- Fix bug where forcing pwlf through points didn't work with higher degrees. See issue https://github.com/cjekel/piecewise_linear_fit_py/issues/54
## [1.1.5] - 2019-11-21
### Changed
- Fix minor typo in docstring of ```calc_slopes```
- Initialized all attributes in the ```__init__``` funciton
- All attributes are now documented in the ```__init__``` function. To view this docstring, use ```pwlf.PiecewiseLinFit?```.
## [1.1.4] - 2019-10-24
### Changed
- TensorFlow 2.0.0 is not (and most probably will not) be supported. DepreciationWarning is displayed when using the ```PiecewiseLinFitTF``` object. Setup.py checks for this optional requirement. Tests are run on Tensorflow<2.0.0.
- TravisCi now checks Python version 3.7 in addition to 3.6, 3.5, 2.7.
- TravisCi tests should now be run daily.
## [1.1.3] - 2019-09-14
### Changed
- Make .ssr stored with fit_with_break* functions
## [1.1.2] - 2019-08-19
### Changed
- Bug fix in non-linear standard error, predict was calling y instead of x. https://github.com/cjekel/piecewise_linear_fit_py/pull/46 Thanks to @tcanders
## [1.1.1] - 2019-08-18
### Changed
- Raise the correct AttributeError when a fit has not yet been performed
## [1.1.0] - 2019-06-16
### Added
- Now you can calculate standard errors for non-linear regression using the Delta method! Check out this [example](https://github.com/cjekel/piecewise_linear_fit_py/tree/master/examples#non-linear-standard-errors-and-p-values).
## [1.0.1] - 2019-06-15
### Added
- Now you can fit constants and continuous polynomials with pwlf! Just specify the keyword ```degree=``` when initializing the ```PiecewiseLinFit``` object. Note that ```degree=0``` for constants, ```degree==1``` for linear (default), ```degree==2``` for quadratics, etc.
- You can manually specify the optimization bounds for each breakpoint when calling the ```fit``` functions by using the ```bounds=``` keyword. Check out the related example.
### Changed
- n_parameters is now calculated based on the shape of the regression matrix
- assembly of the regression matrix now considers which degree polynomial
- n_segments calculated from break points...
- Greatly reduce teststf.py run time
## [1.0.0] - 2019-05-16
### Changed
- Numpy matrix assembly is now ~100x times faster, which will translate to much faster fits! See this [comment](https://github.com/cjekel/piecewise_linear_fit_py/issues/20#issuecomment-492860953) about the speed up. There should no longer be any performance benefits with using the ```PiecewiseLinFitTF``` (TensorFlow) object, so the only reason to use ```PiecewiseLinFitTF``` is if you want access to TensorFlow's optimizers.
### Removed
- There are no sort or order optional parameters in ```PiecewiseLinFit```. The new matrix assembly method doesn't need sorted data. This may break backwards compatibility with your code.
## [0.5.1] - 2019-05-05
### Changed
- Fixed ```PiecewiseLinFitTF``` for Python 2.
## [0.5.0] - 2019-04-15
### Added
- New ```PiecewiseLinFitTF``` class which uses TensorFlow to accelerate pwlf. This class is nearly identical to ```PiecewiseLinFit```, with the exception of the removed ```sorted_data``` options. If you have TensorFlow installed you'll be able to use ```PiecewiseLinFitTF```. If you do not have TensorFlow installed, importing pwlf will issue a warning that ```PiecewiseLinFitTF``` is not available. See this blog [post](https://jekel.me/2019/Adding-tensorflow-to-pwlf/) for more information and benchmarks. The new class includes an option to use float32 or float64 data types.
- ```lapack_driver``` option to choose between the least squares backend. For more see https://docs.scipy.org/doc/scipy/reference/generated/scipy.linalg.lstsq.html and http://www.netlib.org/lapack/lug/node27.html
### Changed
- Now use scipy.linalg instead of numpy.linalg because scipy is always compiled with lapack
- least squares fit now defaults to scipy instead of numpy
### Removed
- ```rcond``` optional parameter; was not necessary with scipy.linalg
## [0.4.3] - 2019-04-02
### Changed
- You can now manually specify ```rcond``` for the numpy least squares solver. For more see https://github.com/cjekel/piecewise_linear_fit_py/issues/21 .
## [0.4.2] - 2019-03-22
### Changed
- ```assemble_regression_matrix()``` now checks if breaks is a numpy array
## [0.4.1] - 2019-03-18
### Changed
- p_values() now uses the correct degrees of freedom (Thanks to Tong Qiu for pointing this out!)
- p_values() now returns a value that can be compared to alpha; previous values would have been compared to alpha/2
## [0.4.0] - 2019-03-14
### Added
- new ```assemble_regression_matrix()``` function that returns the linear regression matrix ```A``` which can allow you to do some more complicated [fits](https://jekel.me/2019/detect-number-of-line-segments-in-pwlf/)
- test function for the linear regression matrix
- new ```fit_guess()``` function to perform a fit when you have an estimate of the breakpoint locations
### Changed
- consolidated the assembly of the linear regression matrix to a single function (and removed the duplicate code)
## [0.3.5] - 2019-02-25
### Changed
- minor correction to r_squared docstring example
## [0.3.4] - 2019-02-06
### Added
- Uploaded paper and citation information
- Added example of what happens when you have more unknowns than data
### Changed
- Examples now include figures
## [0.3.3] - 2019-01-32
### Added
- Documentation and link to documentation
### Changed
- Minor changes to docstrings (spelling, formatting, attribute fixes)
## [0.3.2] - 2019-01-24
### Added
- y-intercepts are now calculated when running the calc_slopes() function. The Y-intercept for each line is stored in self.intercepts.
## [0.3.1] - 2018-12-09
### Added
- p_values() function to calculate the p-value of each beta parameter (WARNING! you may only want to use this if you specify the break point locations, as this does not account for uncertainty in break point locations)
### Changed
- Now changes stored in CHANGELOG.md
## [0.3.0] - 2018-12-05
### Added
- r_squared() function to calculate the coefficent of determination after a fit has been performed
### Changed
- complete docstring overhaul to match the numpydoc style
- fix issues where sum-of-squares of residuals returned array_like, now should always return float
### Removed
- legacy piecewise_lin_fit object has been removed
## [0.2.10] - 2018-12-04
### Changed
- Minor docstring changes
- Fix spelling mistakes throughout
- Fix README.rst format for PyPI
## [0.0.X - 0.2.9] - from 2017-04-01 to 2018-10-03
- 2018/10/03 Add example of bare minimum model persistance to predict for new data (see examples/model_persistence_prediction.py). Bug fix in predict function for custom parameters. Add new test function to check that predict works with custom parameters.
- 2018/08/11 New function which calculates the predication variance for given array of x locations. The predication variance is the squared version of the standard error (not to be confused with the standard errors of the previous change). New example prediction_variance.py shows how to use the new function.
- 2018/06/16 New function which calculates the standard error for each of the model parameters (Remember model parameters are stored as my_pwlf.beta). Standard errors are calculated by calling se = my_pwlf.standard_errors() after you have performed a fit. For more information about standard errors see [this](https://en.wikipedia.org/wiki/Standard_error). Fix docstrings for all functions.
- 2018/05/11 New sorted_data key which can be used to avoided sorting already ordered data. If your data is already ordered as x[0] < x[1] < ... < x[n-1], you may consider using sorted_data=True for a slight performance increase. Additionally the predict function can take the sorted_data key if the data you want to predict at is already sorted. Thanks to [V-Kh](https://github.com/V-Kh) for the idea and PR.
- 2018/04/15 Now you can find piecewise linear fits that go through specified data points! Read [this post](http://jekel.me/2018/Force-piecwise-linear-fit-through-data/) for the details.
- 2018/04/09 Intelligently converts your x, y, or breaks to be numpy array.
- 2018/04/06 Speed! pwlf just got better and faster! A vast majority of this library has been entirely rewritten! New naming convention. The class piecewise_lin_fit() is being depreciated, now use the class PiecewiseLinFit(). See [this post](http://jekel.me/2018/Continous-piecewise-linear-regression/) for details on the new formulation. New test function that tests predict().
- 2018/03/25 Default now hides optimization results. Use disp_res=True when initializing piecewise_lin_fit to change. The multi-start fitfast() function now defaults to the minimum population of 2.
- 2018/03/11 Added try/except behavior for fitWithBreaks function such that the function could be used in an optimization routine. In general when you have a singular matrix, the function will now return np.inf.
- 2018/02/16 Added new fitfast() function which uses multi-start gradient optimization instead of Differential Evolution. It may be substantially faster for your application. Also it would be a good candidate if you don't need the best solution, but just a reasonable fit. Fixed bug in tests function where assert was checking bound, not SSr. New requirement, pyDOE library. New 0.1.0 Version.
- 2017/11/03 add setup.py, new tests folder and test scripts, new version tracking, initialize break0 breakN in the beginning
- 2017/10/31 bug fix related to the case where break points exactly equal to x data points ( as per issue https://github.com/cjekel/piecewise_linear_fit_py/issues/1 ) and added attributes .sep_data_x, .sep_data_y, .sep_predict_data_x for troubleshooting issues related to the separation of data points to their respective regions
- 2017/10/20 remove determinant calculation and use try-except instead, this will offer a larger performance boost for big problems. Change library name to something more Pythonic. Add version attribute.
- 2017/08/03 gradients (slopes of the line segments) now stored as piecewise_lin_fit.slopes (or myPWLF.slopes) after they have been calculated by performing a fit or predicting
- 2017/04/01 initial release
================================================
FILE: LICENSE
================================================
MIT License
Copyright (c) 2017-2022 Charles Jekel
Permission is hereby granted, free of charge, to any person obtaining a copy
of this software and associated documentation files (the "Software"), to deal
in the Software without restriction, including without limitation the rights
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
copies of the Software, and to permit persons to whom the Software is
furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
================================================
FILE: MANIFEST.in
================================================
include LICENSE
include README.rst
================================================
FILE: README.md
================================================
# About
A library for fitting continuous piecewise linear functions to data. Just specify the number of line segments you desire and provide the data.
  [](https://codecov.io/gh/cjekel/piecewise_linear_fit_py)  [](https://anaconda.org/conda-forge/pwlf)
Check out the [documentation](https://jekel.me/piecewise_linear_fit_py)!
Read the [blog post](http://jekel.me/2017/Fit-a-piecewise-linear-function-to-data/).
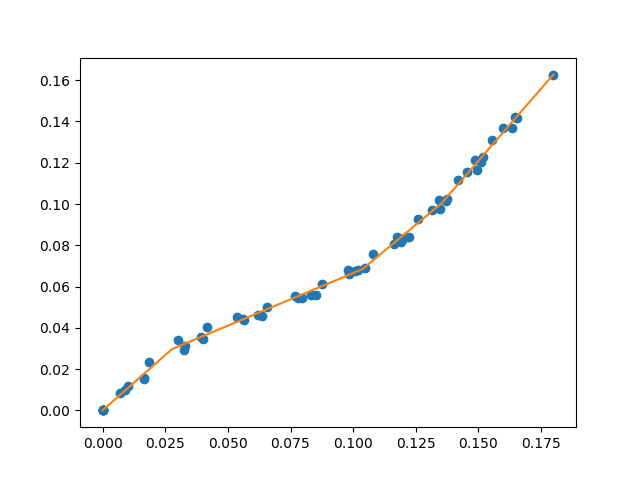
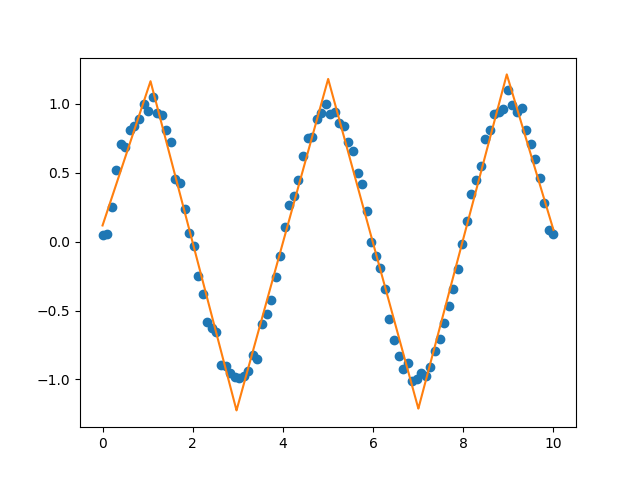
Now you can perform segmented constant fitting and piecewise polynomials!
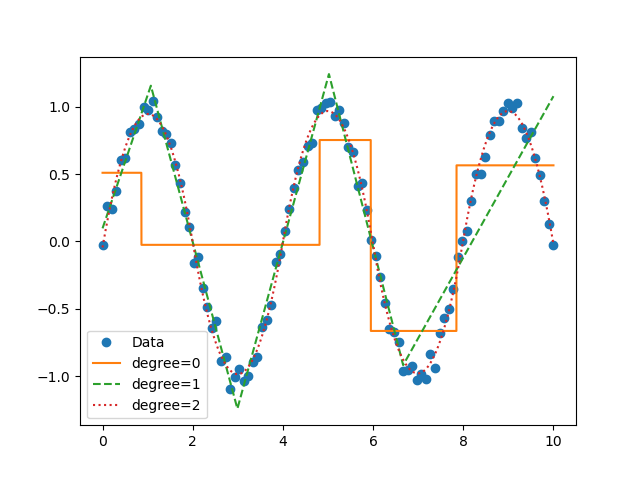
# Features
For a specified number of line segments, you can determine (and predict from) the optimal continuous piecewise linear function f(x). See [this example](https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitForSpecifiedNumberOfLineSegments.py).
You can fit and predict a continuous piecewise linear function f(x) if you know the specific x locations where the line segments terminate. See [this example](https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitWithKnownLineSegmentLocations.py).
If you want to pass different keywords for the SciPy differential evolution algorithm see [this example](https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitForSpecifiedNumberOfLineSegments_passDiffEvoKeywords.py).
You can use a different optimization algorithm to find the optimal location for line segments by using the objective function that minimizes the sum of square of residuals. See [this example](https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/useCustomOptimizationRoutine.py).
Instead of using differential evolution, you can now use a multi-start gradient optimization with fitfast() function. You can specify the number of starting points to use. The default is 2. This means that a latin hyper cube sampling (space filling DOE) of 2 is used to run 2 L-BFGS-B optimizations. See [this example](https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/sineWave_time_compare.py) which runs fit() function, then runs the fitfast() to compare the runtime differences!
# Installation
## Python Package Index (PyPI)
You can now install with pip.
```
python -m pip install pwlf
```
## Conda
If you have conda, you can also install from conda-forge.
```
conda install -c conda-forge pwlf
```
## From source
Or clone the repo
```
git clone https://github.com/cjekel/piecewise_linear_fit_py.git
```
then install with pip
```
python -m pip install ./piecewise_linear_fit_py
```
# How it works
This [paper](https://github.com/cjekel/piecewise_linear_fit_py/raw/master/paper/pwlf_Jekel_Venter_v2.pdf) explains how this library works in detail.
This is based on a formulation of a piecewise linear least squares fit, where the user must specify the location of break points. See [this post](http://jekel.me/2018/Continous-piecewise-linear-regression/) which goes through the derivation of a least squares regression problem if the break point locations are known. Alternatively check out [Golovchenko (2004)](http://golovchenko.org/docs/ContinuousPiecewiseLinearFit.pdf).
Global optimization is used to find the best location for the user defined number of line segments. I specifically use the [differential evolution](https://docs.scipy.org/doc/scipy-0.17.0/reference/generated/scipy.optimize.differential_evolution.html) algorithm in SciPy. I default the differential evolution algorithm to be aggressive, and it is probably overkill for your problem. So feel free to pass your own differential evolution keywords to the library. See [this example](https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitForSpecifiedNumberOfLineSegments_passDiffEvoKeywords.py).
# Changelog
All changes now stored in [CHANGELOG.md](https://github.com/cjekel/piecewise_linear_fit_py/blob/master/CHANGELOG.md)
New ```weights=``` keyword allows you to perform weighted pwlf fits! Removed TensorFlow code which can now be found [here](https://github.com/cjekel/piecewise_linear_fit_py_tf).
# Requirements
NumPy >= 1.14.0
SciPy >= 1.8.0
# License
MIT License
# Citation
```bibtex
@Manual{pwlf,
author = {Jekel, Charles F. and Venter, Gerhard},
title = {{pwlf:} A Python Library for Fitting 1D Continuous Piecewise Linear Functions},
year = {2019},
url = {https://github.com/cjekel/piecewise_linear_fit_py}
}
```
================================================
FILE: README.rst
================================================
About
=====
A library for fitting continuous piecewise linear functions to data.
Just specify the number of line segments you desire and provide the
data.
|Downloads a month| |pwlf ci| |codecov| |PyPI version| |Conda|
Check out the
`documentation <https://jekel.me/piecewise_linear_fit_py>`__!
Read the `blog
post <http://jekel.me/2017/Fit-a-piecewise-linear-function-to-data/>`__.
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/examplePiecewiseFit.png
:alt: Example of a continuous piecewise linear fit to data.
Example of a continuous piecewise linear fit to data.
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/sinWaveFit.png
:alt: Example of a continuous piecewise linear fit to a sine wave.
Example of a continuous piecewise linear fit to a sine wave.
Now you can perform segmented constant fitting and piecewise
polynomials! |Example of multiple degree fits to a sine wave.|
Features
========
For a specified number of line segments, you can determine (and predict
from) the optimal continuous piecewise linear function f(x). See `this
example <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitForSpecifiedNumberOfLineSegments.py>`__.
You can fit and predict a continuous piecewise linear function f(x) if
you know the specific x locations where the line segments terminate. See
`this
example <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitWithKnownLineSegmentLocations.py>`__.
If you want to pass different keywords for the SciPy differential
evolution algorithm see `this
example <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitForSpecifiedNumberOfLineSegments_passDiffEvoKeywords.py>`__.
You can use a different optimization algorithm to find the optimal
location for line segments by using the objective function that
minimizes the sum of square of residuals. See `this
example <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/useCustomOptimizationRoutine.py>`__.
Instead of using differential evolution, you can now use a multi-start
gradient optimization with fitfast() function. You can specify the
number of starting points to use. The default is 2. This means that a
latin hyper cube sampling (space filling DOE) of 2 is used to run 2
L-BFGS-B optimizations. See `this
example <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/sineWave_time_compare.py>`__
which runs fit() function, then runs the fitfast() to compare the
runtime differences!
Installation
============
Python Package Index (PyPI)
---------------------------
You can now install with pip.
::
python -m pip install pwlf
Conda
-----
If you have conda, you can also install from conda-forge.
::
conda install -c conda-forge pwlf
From source
-----------
Or clone the repo
::
git clone https://github.com/cjekel/piecewise_linear_fit_py.git
then install with pip
::
python -m pip install ./piecewise_linear_fit_py
How it works
============
This
`paper <https://github.com/cjekel/piecewise_linear_fit_py/raw/master/paper/pwlf_Jekel_Venter_v2.pdf>`__
explains how this library works in detail.
This is based on a formulation of a piecewise linear least squares fit,
where the user must specify the location of break points. See `this
post <http://jekel.me/2018/Continous-piecewise-linear-regression/>`__
which goes through the derivation of a least squares regression problem
if the break point locations are known. Alternatively check out
`Golovchenko
(2004) <http://golovchenko.org/docs/ContinuousPiecewiseLinearFit.pdf>`__.
Global optimization is used to find the best location for the user
defined number of line segments. I specifically use the `differential
evolution <https://docs.scipy.org/doc/scipy-0.17.0/reference/generated/scipy.optimize.differential_evolution.html>`__
algorithm in SciPy. I default the differential evolution algorithm to be
aggressive, and it is probably overkill for your problem. So feel free
to pass your own differential evolution keywords to the library. See
`this
example <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitForSpecifiedNumberOfLineSegments_passDiffEvoKeywords.py>`__.
Changelog
=========
All changes now stored in
`CHANGELOG.md <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/CHANGELOG.md>`__
New ``weights=`` keyword allows you to perform weighted pwlf fits!
Removed TensorFlow code which can now be found
`here <https://github.com/cjekel/piecewise_linear_fit_py_tf>`__.
Requirements
============
NumPy >= 1.14.0
SciPy >= 1.8.0
License
=======
MIT License
Citation
========
.. code:: bibtex
@Manual{pwlf,
author = {Jekel, Charles F. and Venter, Gerhard},
title = {{pwlf:} A Python Library for Fitting 1D Continuous Piecewise Linear Functions},
year = {2019},
url = {https://github.com/cjekel/piecewise_linear_fit_py}
}
.. |Downloads a month| image:: https://img.shields.io/pypi/dm/pwlf.svg
.. |pwlf ci| image:: https://github.com/cjekel/piecewise_linear_fit_py/workflows/pwlf%20ci/badge.svg
.. |codecov| image:: https://codecov.io/gh/cjekel/piecewise_linear_fit_py/branch/master/graph/badge.svg?token=AgeDFEQXed
:target: https://codecov.io/gh/cjekel/piecewise_linear_fit_py
.. |PyPI version| image:: https://img.shields.io/pypi/v/pwlf
.. |Conda| image:: https://img.shields.io/conda/vn/conda-forge/pwlf
:target: https://anaconda.org/conda-forge/pwlf
.. |Example of multiple degree fits to a sine wave.| image:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/multi_degree.png
================================================
FILE: convert_README_to_RST.sh
================================================
#!/usr/bin/env bash
pandoc --from=markdown --to=rst --output=README.rst README.md
================================================
FILE: docs/.buildinfo
================================================
# Sphinx build info version 1
# This file hashes the configuration used when building these files. When it is not found, a full rebuild will be done.
config: 24f4d9dbb6c120d783c7146d16a6ffbd
tags: 645f666f9bcd5a90fca523b33c5a78b7
================================================
FILE: docs/.nojekyll
================================================
================================================
FILE: docs/_sources/about.rst.txt
================================================
About
============
A library for fitting continuous piecewise linear functions to data. Just specify the number of line segments you desire and provide the data.
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/examplePiecewiseFit.png
:alt: Example of a continuous piecewise linear fit to data.
Example of a continuous piecewise linear fit to data.
All changes now stored in
`CHANGELOG.md <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/CHANGELOG.md>`__
Please cite pwlf in your publications if it helps your research.
.. code:: bibtex
@Manual{pwlf,
author = {Jekel, Charles F. and Venter, Gerhard},
title = {{pwlf:} A Python Library for Fitting 1D Continuous Piecewise Linear Functions},
year = {2019},
url = {https://github.com/cjekel/piecewise_linear_fit_py}
}
================================================
FILE: docs/_sources/examples.rst.txt
================================================
Examples
========
All of these examples will use the following data and imports.
.. code:: python
import numpy as np
import matplotlib.pyplot as plt
import pwlf
# your data
y = np.array([0.00000000e+00, 9.69801700e-03, 2.94350340e-02,
4.39052750e-02, 5.45343950e-02, 6.74104940e-02,
8.34831790e-02, 1.02580042e-01, 1.22767939e-01,
1.42172312e-01, 0.00000000e+00, 8.58600000e-06,
8.31543400e-03, 2.34184100e-02, 3.39709150e-02,
4.03581990e-02, 4.53545600e-02, 5.02345260e-02,
5.55253360e-02, 6.14750770e-02, 6.82125120e-02,
7.55892510e-02, 8.38356810e-02, 9.26413070e-02,
1.02039790e-01, 1.11688258e-01, 1.21390666e-01,
1.31196948e-01, 0.00000000e+00, 1.56706510e-02,
3.54628780e-02, 4.63739040e-02, 5.61442590e-02,
6.78542550e-02, 8.16388310e-02, 9.77756110e-02,
1.16531753e-01, 1.37038283e-01, 0.00000000e+00,
1.16951050e-02, 3.12089850e-02, 4.41776550e-02,
5.42877590e-02, 6.63321350e-02, 8.07655920e-02,
9.70363280e-02, 1.15706975e-01, 1.36687642e-01,
0.00000000e+00, 1.50144640e-02, 3.44519970e-02,
4.55907760e-02, 5.59556700e-02, 6.88450940e-02,
8.41374060e-02, 1.01254006e-01, 1.20605073e-01,
1.41881288e-01, 1.62618058e-01])
x = np.array([0.00000000e+00, 8.82678000e-03, 3.25615100e-02,
5.66106800e-02, 7.95549800e-02, 1.00936330e-01,
1.20351520e-01, 1.37442010e-01, 1.51858250e-01,
1.64433570e-01, 0.00000000e+00, -2.12600000e-05,
7.03872000e-03, 1.85494500e-02, 3.00926700e-02,
4.17617000e-02, 5.37279600e-02, 6.54941000e-02,
7.68092100e-02, 8.76596300e-02, 9.80525800e-02,
1.07961810e-01, 1.17305210e-01, 1.26063930e-01,
1.34180360e-01, 1.41725010e-01, 1.48629710e-01,
1.55374770e-01, 0.00000000e+00, 1.65610200e-02,
3.91016100e-02, 6.18679400e-02, 8.30997400e-02,
1.02132890e-01, 1.19011260e-01, 1.34620080e-01,
1.49429370e-01, 1.63539960e-01, -0.00000000e+00,
1.01980300e-02, 3.28642800e-02, 5.59461900e-02,
7.81388400e-02, 9.84458400e-02, 1.16270210e-01,
1.31279040e-01, 1.45437090e-01, 1.59627540e-01,
0.00000000e+00, 1.63404300e-02, 4.00086000e-02,
6.34390200e-02, 8.51085900e-02, 1.04787860e-01,
1.22120350e-01, 1.36931660e-01, 1.50958760e-01,
1.65299640e-01, 1.79942720e-01])
1. `fit with known breakpoint
locations <#fit-with-known-breakpoint-locations>`__
2. `fit for specified number of line
segments <#fit-for-specified-number-of-line-segments>`__
3. `fitfast for specified number of line
segments <#fitfast-for-specified-number-of-line-segments>`__
4. `force a fit through data
points <#force-a-fit-through-data-points>`__
5. `use custom optimization
routine <#use-custom-optimization-routine>`__
6. `pass differential evolution
keywords <#pass-differential-evolution-keywords>`__
7. `find the best number of line
segments <#find-the-best-number-of-line-segments>`__
8. `model persistence <#model-persistence>`__
9. `bad fits when you have more unknowns than
data <#bad-fits-when-you-have-more-unknowns-than-data>`__
10. `fit with a breakpoint guess <#fit-with-a-breakpoint-guess>`__
11. `get the linear regression
matrix <#get-the-linear-regression-matrix>`__
12. `use of TensorFlow <#use-of-tensorflow>`__
13. `fit constants or polynomials <#fit-constants-or-polynomials>`__
14. `specify breakpoint bounds <#specify-breakpoint-bounds>`__
15. `non-linear standard errors and
p-values <#non-linear-standard-errors-and-p-values>`__
16. `obtain the equations of fitted
pwlf <#obtain-the-equations-of-fitted-pwlf>`__
17. `weighted least squares fit <#weighted-least-squares-fit>`__
18. `reproducible results <#reproducible results>`__
fit with known breakpoint locations
-----------------------------------
You can perform a least squares fit if you know the breakpoint
locations.
.. code:: python
# your desired line segment end locations
x0 = np.array([min(x), 0.039, 0.10, max(x)])
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
# fit the data with the specified break points
# (ie the x locations of where the line segments
# will terminate)
my_pwlf.fit_with_breaks(x0)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
# plot the results
plt.figure()
plt.plot(x, y, 'o')
plt.plot(xHat, yHat, '-')
plt.show()
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/fit_breaks.png
:alt: fit with known breakpoint locations
fit with known breakpoint locations
fit for specified number of line segments
-----------------------------------------
Use a global optimization to find the breakpoint locations that minimize
the sum of squares error. This uses `Differential
Evolution <https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.differential_evolution.html>`__
from scipy.
.. code:: python
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
# fit the data for four line segments
res = my_pwlf.fit(4)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
# plot the results
plt.figure()
plt.plot(x, y, 'o')
plt.plot(xHat, yHat, '-')
plt.show()
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/numberoflines.png
:alt: fit for specified number of line segments
fit for specified number of line segments
fitfast for specified number of line segments
---------------------------------------------
This performs a fit for a specified number of line segments with a
multi-start gradient based optimization. This should be faster than
`Differential
Evolution <https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.differential_evolution.html>`__
for a small number of starting points.
.. code:: python
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
# fit the data for four line segments
# this performs 3 multi-start optimizations
res = my_pwlf.fitfast(4, pop=3)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
# plot the results
plt.figure()
plt.plot(x, y, 'o')
plt.plot(xHat, yHat, '-')
plt.show()
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/fitfast.png
:alt: fitfast for specified number of line segments
fitfast for specified number of line segments
force a fit through data points
-------------------------------
Sometimes it’s necessary to force the piecewise continuous model through
a particular data point, or a set of data points. The following example
finds the best 4 line segments that go through two data points.
.. code:: python
# initialize piecewise linear fit with your x and y data
myPWLF = pwlf.PiecewiseLinFit(x, y)
# fit the function with four line segments
# force the function to go through the data points
# (0.0, 0.0) and (0.19, 0.16)
# where the data points are of the form (x, y)
x_c = [0.0, 0.19]
y_c = [0.0, 0.2]
res = myPWLF.fit(4, x_c, y_c)
# predict for the determined points
xHat = np.linspace(min(x), 0.19, num=10000)
yHat = myPWLF.predict(xHat)
# plot the results
plt.figure()
plt.plot(x, y, 'o')
plt.plot(xHat, yHat, '-')
plt.show()
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/force.png
:alt: force a fit through data points
force a fit through data points
use custom optimization routine
-------------------------------
You can use your favorite optimization routine to find the breakpoint
locations. The following example uses scipy’s
`minimize <https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html>`__
function.
.. code:: python
from scipy.optimize import minimize
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
# initialize custom optimization
number_of_line_segments = 3
my_pwlf.use_custom_opt(number_of_line_segments)
# i have number_of_line_segments - 1 number of variables
# let's guess the correct location of the two unknown variables
# (the program defaults to have end segments at x0= min(x)
# and xn=max(x)
xGuess = np.zeros(number_of_line_segments - 1)
xGuess[0] = 0.02
xGuess[1] = 0.10
res = minimize(my_pwlf.fit_with_breaks_opt, xGuess)
# set up the break point locations
x0 = np.zeros(number_of_line_segments + 1)
x0[0] = np.min(x)
x0[-1] = np.max(x)
x0[1:-1] = res.x
# calculate the parameters based on the optimal break point locations
my_pwlf.fit_with_breaks(x0)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
plt.figure()
plt.plot(x, y, 'o')
plt.plot(xHat, yHat, '-')
plt.show()
pass differential evolution keywords
------------------------------------
You can pass keyword arguments from the ``fit`` function into scipy’s
`Differential
Evolution <https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.differential_evolution.html>`__.
.. code:: python
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
# fit the data for four line segments
# this sets DE to have an absolute tolerance of 0.1
res = my_pwlf.fit(4, atol=0.1)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
# plot the results
plt.figure()
plt.plot(x, y, 'o')
plt.plot(xHat, yHat, '-')
plt.show()
find the best number of line segments
-------------------------------------
This example uses EGO (bayesian optimization) and a penalty function to
find the best number of line segments. This will require careful use of
the penalty parameter ``l``. Use this template to automatically find the
best number of line segments.
.. code:: python
from GPyOpt.methods import BayesianOptimization
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
# define your objective function
def my_obj(x):
# define some penalty parameter l
# you'll have to arbitrarily pick this
# it depends upon the noise in your data,
# and the value of your sum of square of residuals
l = y.mean()*0.001
f = np.zeros(x.shape[0])
for i, j in enumerate(x):
my_pwlf.fit(j[0])
f[i] = my_pwlf.ssr + (l*j[0])
return f
# define the lower and upper bound for the number of line segments
bounds = [{'name': 'var_1', 'type': 'discrete',
'domain': np.arange(2, 40)}]
np.random.seed(12121)
myBopt = BayesianOptimization(my_obj, domain=bounds, model_type='GP',
initial_design_numdata=10,
initial_design_type='latin',
exact_feval=True, verbosity=True,
verbosity_model=False)
max_iter = 30
# perform the bayesian optimization to find the optimum number
# of line segments
myBopt.run_optimization(max_iter=max_iter, verbosity=True)
print('\n \n Opt found \n')
print('Optimum number of line segments:', myBopt.x_opt)
print('Function value:', myBopt.fx_opt)
myBopt.plot_acquisition()
myBopt.plot_convergence()
# perform the fit for the optimum
my_pwlf.fit(myBopt.x_opt)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
# plot the results
plt.figure()
plt.plot(x, y, 'o')
plt.plot(xHat, yHat, '-')
plt.show()
model persistence
-----------------
You can save fitted models with pickle. Alternatively see
`joblib <https://joblib.readthedocs.io/en/latest/>`__.
.. code:: python
# if you use Python 2.x you should import cPickle
# import cPickle as pickle
# if you use Python 3.x you can just use pickle
import pickle
# your desired line segment end locations
x0 = np.array([min(x), 0.039, 0.10, max(x)])
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
# fit the data with the specified break points
my_pwlf.fit_with_breaks(x0)
# save the fitted model
with open('my_fit.pkl', 'wb') as f:
pickle.dump(my_pwlf, f, pickle.HIGHEST_PROTOCOL)
# load the fitted model
with open('my_fit.pkl', 'rb') as f:
my_pwlf = pickle.load(f)
bad fits when you have more unknowns than data
----------------------------------------------
You can get very bad fits with pwlf when you have more unknowns than
data points. The following example will fit 99 line segments to the 59
data points. While this will result in an error of zero, the model will
have very weird predictions within the data. You should not fit more
unknowns than you have data with pwlf!
.. code:: python
break_locations = np.linspace(min(x), max(x), num=100)
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
my_pwlf.fit_with_breaks(break_locations)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
# plot the results
plt.figure()
plt.plot(x, y, 'o')
plt.plot(xHat, yHat, '-')
plt.show()
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/badfit.png
:alt: bad fits when you have more unknowns than data
bad fits when you have more unknowns than data
fit with a breakpoint guess
---------------------------
In this example we see two distinct linear regions, and we believe a
breakpoint occurs at 6.0. We’ll use the fit_guess() function to find the
best breakpoint location starting with this guess. These fits should be
much faster than the ``fit`` or ``fitfast`` function when you have a
reasonable idea where the breakpoints occur.
.. code:: python
import numpy as np
import pwlf
x = np.array([4., 5., 6., 7., 8.])
y = np.array([11., 13., 16., 28.92, 42.81])
my_pwlf = pwlf.PiecewiseLinFit(x, y)
breaks = my_pwlf.fit_guess([6.0])
Note specifying one breakpoint will result in two line segments. If we
wanted three line segments, we’ll have to specify two breakpoints.
.. code:: python
breaks = my_pwlf.fit_guess([5.5, 6.0])
get the linear regression matrix
--------------------------------
In some cases it may be desirable to work with the linear regression
matrix directly. The following example grabs the linear regression
matrix ``A`` for a specific set of breakpoints. In this case we assume
that the breakpoints occur at each of the data points. Please see the
`paper <https://github.com/cjekel/piecewise_linear_fit_py/tree/master/paper>`__
for details about the regression matrix ``A``.
.. code:: python
import numpy as np
import pwlf
# select random seed for reproducibility
np.random.seed(123)
# generate sin wave data
x = np.linspace(0, 10, num=100)
y = np.sin(x * np.pi / 2)
ytrue = y.copy()
# add noise to the data
y = np.random.normal(0, 0.05, 100) + ytrue
my_pwlf_en = pwlf.PiecewiseLinFit(x, y)
# copy the x data to use as break points
breaks = my_pwlf_en.x_data.copy()
# create the linear regression matrix A
A = my_pwlf_en.assemble_regression_matrix(breaks, my_pwlf_en.x_data)
We can perform fits that are more complicated than a least squares fit
when we have the regression matrix. The following uses the Elastic Net
regularizer to perform an interesting fit with the regression matrix.
.. code:: python
from sklearn.linear_model import ElasticNetCV
# set up the elastic net
en_model = ElasticNetCV(cv=5,
l1_ratio=[.1, .5, .7, .9,
.95, .99, 1],
fit_intercept=False,
max_iter=1000000, n_jobs=-1)
# fit the model using the elastic net
en_model.fit(A, my_pwlf_en.y_data)
# predict from the elastic net parameters
xhat = np.linspace(x.min(), x.max(), 1000)
yhat_en = my_pwlf_en.predict(xhat, breaks=breaks,
beta=en_model.coef_)
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/sin_en_net_fit.png
:alt: interesting elastic net fit
interesting elastic net fit
use of tensorflow
-----------------
You need to install
`pwlftf <https://github.com/cjekel/piecewise_linear_fit_py_tf>`__ which
will have the ``PiecewiseLinFitTF`` class. For performance benchmarks
(these benchmarks are outdated! and the regular pwlf may be faster in
many applications) see this blog
`post <https://jekel.me/2019/Adding-tensorflow-to-pwlf/>`__.
The use of the TF class is nearly identical to the original class,
however note the following exceptions. ``PiecewiseLinFitTF`` does:
- not have a ``lapack_driver`` option
- have an optional parameter ``dtype``, so you can choose between the
float64 and float32 data types
- have an optional parameter ``fast`` to switch between Cholesky
decomposition (default ``fast=True``), and orthogonal decomposition
(``fast=False``)
.. code:: python
import pwlftf as pwlf
# your desired line segment end locations
x0 = np.array([min(x), 0.039, 0.10, max(x)])
# initialize TF piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFitTF(x, y, dtype='float32)
# fit the data with the specified break points
# (ie the x locations of where the line segments
# will terminate)
my_pwlf.fit_with_breaks(x0)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
fit constants or polynomials
----------------------------
You can use pwlf to fit segmented constant models, or piecewise
polynomials. The following example fits a segmented constant model,
piecewise linear, and a piecewise quadratic model to a sine wave.
.. code:: python
# generate sin wave data
x = np.linspace(0, 10, num=100)
y = np.sin(x * np.pi / 2)
# add noise to the data
y = np.random.normal(0, 0.05, 100) + y
# initialize piecewise linear fit with your x and y data
# pwlf lets you fit continuous model for many degree polynomials
# degree=0 constant
# degree=1 linear (default)
# degree=2 quadratic
my_pwlf_0 = pwlf.PiecewiseLinFit(x, y, degree=0)
my_pwlf_1 = pwlf.PiecewiseLinFit(x, y, degree=1) # default
my_pwlf_2 = pwlf.PiecewiseLinFit(x, y, degree=2)
# fit the data for four line segments
res0 = my_pwlf_0.fitfast(5, pop=50)
res1 = my_pwlf_1.fitfast(5, pop=50)
res2 = my_pwlf_2.fitfast(5, pop=50)
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat0 = my_pwlf_0.predict(xHat)
yHat1 = my_pwlf_1.predict(xHat)
yHat2 = my_pwlf_2.predict(xHat)
# plot the results
plt.figure()
plt.plot(x, y, 'o', label='Data')
plt.plot(xHat, yHat0, '-', label='degree=0')
plt.plot(xHat, yHat1, '--', label='degree=1')
plt.plot(xHat, yHat2, ':', label='degree=2')
plt.legend()
plt.show()
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/multi_degree.png
:alt: Example of multiple degree fits to a sine wave.
Example of multiple degree fits to a sine wave.
specify breakpoint bounds
-------------------------
You may want extra control over the search space for feasible
breakpoints. One way to do this is to specify the bounds for each
breakpoint location.
.. code:: python
# generate sin wave data
x = np.linspace(0, 10, num=100)
y = np.sin(x * np.pi / 2)
# add noise to the data
y = np.random.normal(0, 0.05, 100) + y
# initialize piecewise linear fit with your x and y data
my_pwlf = pwlf.PiecewiseLinFit(x, y)
# define custom bounds for the interior break points
n_segments = 4
bounds = np.zeros((n_segments-1, 2))
# first breakpoint
bounds[0, 0] = 0.0 # lower bound
bounds[0, 1] = 3.5 # upper bound
# second breakpoint
bounds[1, 0] = 3.0 # lower bound
bounds[1, 1] = 7.0 # upper bound
# third breakpoint
bounds[2, 0] = 6.0 # lower bound
bounds[2, 1] = 10.0 # upper bound
res = my_pwlf.fit(n_segments, bounds=bounds)
non-linear standard errors and p-values
---------------------------------------
You can calculate non-linear standard errors using the Delta method.
This will calculate the standard errors of the piecewise linear
parameters (intercept + slopes) and the breakpoint locations!
First let us generate true piecewise linear data.
.. code:: python
from __future__ import print_function
# generate a true piecewise linear data
np.random.seed(5)
n_data = 100
x = np.linspace(0, 1, num=n_data)
y = np.random.random(n_data)
my_pwlf_gen = pwlf.PiecewiseLinFit(x, y)
true_beta = np.random.normal(size=5)
true_breaks = np.array([0.0, 0.2, 0.5, 0.75, 1.0])
y = my_pwlf_gen.predict(x, beta=true_beta, breaks=true_breaks)
plt.figure()
plt.title('True piecewise linear data')
plt.plot(x, y)
plt.show()
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/true_pwlf.png
:alt: True piecewise linear data.
True piecewise linear data.
Now we can perform a fit, calculate the standard errors, and p-values.
The non-linear method uses a first order taylor series expansion to
linearize the non-linear regression problem. A positive step_size
performs a forward difference, and a negative step_size would perform a
backwards difference.
.. code:: python
my_pwlf = pwlf.PiecewiseLinFit(x, y)
res = my_pwlf.fitfast(4, pop=100)
p = my_pwlf.p_values(method='non-linear', step_size=1e-4)
se = my_pwlf.se # standard errors
The standard errors and p-values correspond to each model parameter.
First the beta parameters (intercept + slopes) and then the breakpoints.
We can assemble the parameters, and print a table of the result with the
following code.
.. code:: python
parameters = np.concatenate((my_pwlf.beta,
my_pwlf.fit_breaks[1:-1]))
header = ['Parameter type', 'Parameter value', 'Standard error', 't',
'P > np.abs(t) (p-value)']
print(*header, sep=' | ')
values = np.zeros((parameters.size, 5), dtype=np.object_)
values[:, 1] = np.around(parameters, decimals=3)
values[:, 2] = np.around(se, decimals=3)
values[:, 3] = np.around(parameters / se, decimals=3)
values[:, 4] = np.around(p, decimals=3)
for i, row in enumerate(values):
if i < my_pwlf.beta.size:
row[0] = 'Beta'
print(*row, sep=' | ')
else:
row[0] = 'Breakpoint'
print(*row, sep=' | ')
============== =============== ============== ============== =======================
Parameter type Parameter value Standard error t P > np.abs(t) (p-value)
============== =============== ============== ============== =======================
Beta 1.821 0.0 1763191476.046 0.0
Beta -0.427 0.0 -46404554.493 0.0
Beta -1.165 0.0 -111181494.162 0.0
Beta -1.397 0.0 -168954500.421 0.0
Beta 0.873 0.0 93753841.242 0.0
Breakpoint 0.2 0.0 166901856.885 0.0
Breakpoint 0.5 0.0 537785803.646 0.0
Breakpoint 0.75 0.0 482311769.159 0.0
============== =============== ============== ============== =======================
obtain the equations of fitted pwlf
-----------------------------------
Sometimes you may want the mathematical equations that represent your
fitted model. This is easy to perform if you don’t mind using sympy.
The following code will fit 5 line segments of degree=2 to a sin wave.
.. code:: python
import numpy as np
import pwlf
# generate sin wave data
x = np.linspace(0, 10, num=100)
y = np.sin(x * np.pi / 2)
# add noise to the data
y = np.random.normal(0, 0.05, 100) + y
my_pwlf_2 = pwlf.PiecewiseLinFit(x, y, degree=2)
res2 = my_pwlf_2.fitfast(5, pop=50)
Given this fit, the following code will print the mathematical equation
for each line segment.
.. code:: python
from sympy import Symbol
from sympy.utilities import lambdify
x = Symbol('x')
def get_symbolic_eqn(pwlf_, segment_number):
if pwlf_.degree < 1:
raise ValueError('Degree must be at least 1')
if segment_number < 1 or segment_number > pwlf_.n_segments:
raise ValueError('segment_number not possible')
# assemble degree = 1 first
for line in range(segment_number):
if line == 0:
my_eqn = pwlf_.beta[0] + (pwlf_.beta[1])*(x-pwlf_.fit_breaks[0])
else:
my_eqn += (pwlf_.beta[line+1])*(x-pwlf_.fit_breaks[line])
# assemble all other degrees
if pwlf_.degree > 1:
for k in range(2, pwlf_.degree + 1):
for line in range(segment_number):
beta_index = pwlf_.n_segments*(k-1) + line + 1
my_eqn += (pwlf_.beta[beta_index])*(x-pwlf_.fit_breaks[line])**k
return my_eqn.simplify()
eqn_list = []
f_list = []
for i in range(my_pwlf_2.n_segments):
eqn_list.append(get_symbolic_eqn(my_pwlf_2, i + 1))
print('Equation number: ', i + 1)
print(eqn_list[-1])
f_list.append(lambdify(x, eqn_list[-1]))
which should print out something like the following:
.. code:: python
Equation number: 1
-0.953964059782599*x**2 + 1.89945177490653*x + 0.00538634182565454
Equation number: 2
0.951561315686298*x**2 - 5.69747505830914*x + 7.5772216545711
Equation number: 3
-0.949735350431857*x**2 + 9.48218236957122*x - 22.720785454735
Equation number: 4
0.926850298824217*x**2 - 12.9824424358344*x + 44.5102742956827
Equation number: 5
-1.03016230425747*x**2 + 18.5306546317065*x - 82.3508513333073
For more information on how this works, see
`this <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/understanding_higher_degrees/polynomials_in_pwlf.ipynb>`__
jupyter notebook.
weighted least squares fit
--------------------------
Sometimes your data will not have a constant variance
(heteroscedasticity), and you need to perform a weighted least squares
fit. The following example will perform a standard and weighted fit so
you can compare the differences. First we need to generate a data set
which will be a good candidate to use for weighted least squares fits.
.. code:: python
# generate data with heteroscedasticity
n = 100
n_data_sets = 100
n_segments = 6
# generate sine data
x = np.linspace(0, 10, n)
y = np.zeros((n_data_sets, n))
sigma_change = np.linspace(0.001, 0.05, 100)
for i in range(n_data_sets):
y[i] = np.sin(x * np.pi / 2)
# add noise to the data
y[i] = np.random.normal(0, sigma_change, 100) + y[i]
X = np.tile(x, n_data_sets)
The individual weights in pwlf are the reciprocal of the standard
deviation for each data point. Here weights[i] corresponds to one over
the standard deviation of the ith data point. The result of this is that
data points with higher variance are less important to the overall pwlf
than data point with small variance. Let’s perform a standard pwlf fit
and a weighted fit.
.. code:: python
# perform an ordinary pwlf fit to the entire data
my_pwlf = pwlf.PiecewiseLinFit(X.flatten(), y.flatten())
my_pwlf.fit(n_segments)
# compute the standard deviation in y
y_std = np.std(y, axis=0)
# set the weights to be one over the standard deviation
weights = 1.0 / y_std
# perform a weighted least squares to the data
my_pwlf_w = pwlf.PiecewiseLinFit(x, y.mean(axis=0), weights=weights)
my_pwlf_w.fit(n_segments)
# compare the fits
xhat = np.linspace(0, 10, 1000)
yhat = my_pwlf.predict(xhat)
yhat_w = my_pwlf_w.predict(xhat)
plt.figure()
plt.plot(X.flatten(), y.flatten(), '.')
plt.plot(xhat, yhat, '-', label='Ordinary LS')
plt.plot(xhat, yhat_w, '-', label='Weighted LS')
plt.legend()
plt.show()
.. figure:: https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/weighted_least_squares_example.png
:alt: Weighted pwlf fit.
Weighted pwlf fit.
We can see that the weighted pwlf fit tries fit data with low variance
better than data with high variance, however the ordinary pwlf fits the
data assuming a uniform variance.
reproducible results
--------------------
The `fit` and `fitfast` methods are stochastic and may not give the same
result every time the program is run. To have reproducible results you can
manually specify a numpy.random.seed on init. Now everytime the following
program is run, the results of the fit(2) should be the same.
.. code:: python
# initialize piecewise linear fit with a random seed
my_pwlf = pwlf.PiecewiseLinFit(x, y, seed=123)
# Now the fit() method will be reproducible
my_pwlf.fit(2)
================================================
FILE: docs/_sources/how_it_works.rst.txt
================================================
How it works
============
This
`paper <https://github.com/cjekel/piecewise_linear_fit_py/raw/master/paper/pwlf_Jekel_Venter_v2.pdf>`__
explains how this library works in detail.
This is based on a formulation of a piecewise linear least squares fit,
where the user must specify the location of break points. See `this
post <http://jekel.me/2018/Continous-piecewise-linear-regression/>`__
which goes through the derivation of a least squares regression problem
if the break point locations are known. Alternatively check out
`Golovchenko
(2004) <http://golovchenko.org/docs/ContinuousPiecewiseLinearFit.pdf>`__.
Global optimization is used to find the best location for the user
defined number of line segments. I specifically use the `differential
evolution <https://docs.scipy.org/doc/scipy-0.17.0/reference/generated/scipy.optimize.differential_evolution.html>`__
algorithm in SciPy. I default the differential evolution algorithm to be
aggressive, and it is probably overkill for your problem. So feel free
to pass your own differential evolution keywords to the library. See
`this
example <https://github.com/cjekel/piecewise_linear_fit_py/blob/master/examples/fitForSpecifiedNumberOfLineSegments_passDiffEvoKeywords.py>`__.
================================================
FILE: docs/_sources/index.rst.txt
================================================
pwlf: piecewise linear fitting
================================
Fit piecewise linear functions to data!
.. toctree::
:maxdepth: 2
installation
how_it_works
examples
pwlf
about
requirements
license
Indices and tables
==================
* :ref:`genindex`
* :ref:`search`
================================================
FILE: docs/_sources/installation.rst.txt
================================================
Installation
============
Python Package Index (PyPI)
---------------------------
You can now install with pip.
::
python -m pip install pwlf
Conda
-----
If you have conda, you can also install from conda-forge.
::
conda install -c conda-forge pwlf
From source
-----------
Or clone the repo
::
git clone https://github.com/cjekel/piecewise_linear_fit_py.git
then install with pip
::
python -m pip install ./piecewise_linear_fit_py
================================================
FILE: docs/_sources/license.rst.txt
================================================
License
=======
MIT License
Copyright (c) 2017-2020 Charles Jekel
Permission is hereby granted, free of charge, to any person obtaining a copy
of this software and associated documentation files (the "Software"), to deal
in the Software without restriction, including without limitation the rights
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
copies of the Software, and to permit persons to whom the Software is
furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
================================================
FILE: docs/_sources/modules.rst.txt
================================================
pwlf
====
.. toctree::
:maxdepth: 4
pwlf
================================================
FILE: docs/_sources/pwlf.rst.txt
================================================
pwlf package contents
============
.. autosummary::
:toctree: stubs
pwlf.PiecewiseLinFit
.. autoclass:: pwlf.PiecewiseLinFit
:members:
:undoc-members:
:show-inheritance:
================================================
FILE: docs/_sources/requirements.rst.txt
================================================
Requirements
============
`NumPy <https://pypi.org/project/numpy/>`__ (>= 1.14.0)
`SciPy <https://pypi.org/project/scipy/>`__ (>= 1.2.0)
`pyDOE <https://pypi.org/project/pyDOE/>`__ ( >= 0.3.8)
================================================
FILE: docs/_sources/stubs/pwlf.PiecewiseLinFit.rst.txt
================================================
pwlf.PiecewiseLinFit
====================
.. currentmodule:: pwlf
.. autoclass:: PiecewiseLinFit
.. automethod:: __init__
.. rubric:: Methods
.. autosummary::
~PiecewiseLinFit.__init__
~PiecewiseLinFit.assemble_regression_matrix
~PiecewiseLinFit.calc_slopes
~PiecewiseLinFit.conlstsq
~PiecewiseLinFit.fit
~PiecewiseLinFit.fit_force_points_opt
~PiecewiseLinFit.fit_guess
~PiecewiseLinFit.fit_with_breaks
~PiecewiseLinFit.fit_with_breaks_force_points
~PiecewiseLinFit.fit_with_breaks_opt
~PiecewiseLinFit.fitfast
~PiecewiseLinFit.lstsq
~PiecewiseLinFit.p_values
~PiecewiseLinFit.predict
~PiecewiseLinFit.prediction_variance
~PiecewiseLinFit.r_squared
~PiecewiseLinFit.standard_errors
~PiecewiseLinFit.use_custom_opt
================================================
FILE: docs/_static/alabaster.css
================================================
/* -- page layout ----------------------------------------------------------- */
body {
font-family: Georgia, serif;
font-size: 17px;
background-color: #fff;
color: #000;
margin: 0;
padding: 0;
}
div.document {
width: 940px;
margin: 30px auto 0 auto;
}
div.documentwrapper {
float: left;
width: 100%;
}
div.bodywrapper {
margin: 0 0 0 220px;
}
div.sphinxsidebar {
width: 220px;
font-size: 14px;
line-height: 1.5;
}
hr {
border: 1px solid #B1B4B6;
}
div.body {
background-color: #fff;
color: #3E4349;
padding: 0 30px 0 30px;
}
div.body > .section {
text-align: left;
}
div.footer {
width: 940px;
margin: 20px auto 30px auto;
font-size: 14px;
color: #888;
text-align: right;
}
div.footer a {
color: #888;
}
p.caption {
font-family: inherit;
font-size: inherit;
}
div.relations {
display: none;
}
div.sphinxsidebar {
max-height: 100%;
overflow-y: auto;
}
div.sphinxsidebar a {
color: #444;
text-decoration: none;
border-bottom: 1px dotted #999;
}
div.sphinxsidebar a:hover {
border-bottom: 1px solid #999;
}
div.sphinxsidebarwrapper {
padding: 18px 10px;
}
div.sphinxsidebarwrapper p.logo {
padding: 0;
margin: -10px 0 0 0px;
text-align: center;
}
div.sphinxsidebarwrapper h1.logo {
margin-top: -10px;
text-align: center;
margin-bottom: 5px;
text-align: left;
}
div.sphinxsidebarwrapper h1.logo-name {
margin-top: 0px;
}
div.sphinxsidebarwrapper p.blurb {
margin-top: 0;
font-style: normal;
}
div.sphinxsidebar h3,
div.sphinxsidebar h4 {
font-family: Georgia, serif;
color: #444;
font-size: 24px;
font-weight: normal;
margin: 0 0 5px 0;
padding: 0;
}
div.sphinxsidebar h4 {
font-size: 20px;
}
div.sphinxsidebar h3 a {
color: #444;
}
div.sphinxsidebar p.logo a,
div.sphinxsidebar h3 a,
div.sphinxsidebar p.logo a:hover,
div.sphinxsidebar h3 a:hover {
border: none;
}
div.sphinxsidebar p {
color: #555;
margin: 10px 0;
}
div.sphinxsidebar ul {
margin: 10px 0;
padding: 0;
color: #000;
}
div.sphinxsidebar ul li.toctree-l1 > a {
font-size: 120%;
}
div.sphinxsidebar ul li.toctree-l2 > a {
font-size: 110%;
}
div.sphinxsidebar input {
border: 1px solid #CCC;
font-family: Georgia, serif;
font-size: 1em;
}
div.sphinxsidebar #searchbox {
margin: 1em 0;
}
div.sphinxsidebar .search > div {
display: table-cell;
}
div.sphinxsidebar hr {
border: none;
height: 1px;
color: #AAA;
background: #AAA;
text-align: left;
margin-left: 0;
width: 50%;
}
div.sphinxsidebar .badge {
border-bottom: none;
}
div.sphinxsidebar .badge:hover {
border-bottom: none;
}
/* To address an issue with donation coming after search */
div.sphinxsidebar h3.donation {
margin-top: 10px;
}
/* -- body styles ----------------------------------------------------------- */
a {
color: #004B6B;
text-decoration: underline;
}
a:hover {
color: #6D4100;
text-decoration: underline;
}
div.body h1,
div.body h2,
div.body h3,
div.body h4,
div.body h5,
div.body h6 {
font-family: Georgia, serif;
font-weight: normal;
margin: 30px 0px 10px 0px;
padding: 0;
}
div.body h1 { margin-top: 0; padding-top: 0; font-size: 240%; }
div.body h2 { font-size: 180%; }
div.body h3 { font-size: 150%; }
div.body h4 { font-size: 130%; }
div.body h5 { font-size: 100%; }
div.body h6 { font-size: 100%; }
a.headerlink {
color: #DDD;
padding: 0 4px;
text-decoration: none;
}
a.headerlink:hover {
color: #444;
background: #EAEAEA;
}
div.body p, div.body dd, div.body li {
line-height: 1.4em;
}
div.admonition {
margin: 20px 0px;
padding: 10px 30px;
background-color: #EEE;
border: 1px solid #CCC;
}
div.admonition tt.xref, div.admonition code.xref, div.admonition a tt {
background-color: #FBFBFB;
border-bottom: 1px solid #fafafa;
}
div.admonition p.admonition-title {
font-family: Georgia, serif;
font-weight: normal;
font-size: 24px;
margin: 0 0 10px 0;
padding: 0;
line-height: 1;
}
div.admonition p.last {
margin-bottom: 0;
}
dt:target, .highlight {
background: #FAF3E8;
}
div.warning {
background-color: #FCC;
border: 1px solid #FAA;
}
div.danger {
background-color: #FCC;
border: 1px solid #FAA;
-moz-box-shadow: 2px 2px 4px #D52C2C;
-webkit-box-shadow: 2px 2px 4px #D52C2C;
box-shadow: 2px 2px 4px #D52C2C;
}
div.error {
background-color: #FCC;
border: 1px solid #FAA;
-moz-box-shadow: 2px 2px 4px #D52C2C;
-webkit-box-shadow: 2px 2px 4px #D52C2C;
box-shadow: 2px 2px 4px #D52C2C;
}
div.caution {
background-color: #FCC;
border: 1px solid #FAA;
}
div.attention {
background-color: #FCC;
border: 1px solid #FAA;
}
div.important {
background-color: #EEE;
border: 1px solid #CCC;
}
div.note {
background-color: #EEE;
border: 1px solid #CCC;
}
div.tip {
background-color: #EEE;
border: 1px solid #CCC;
}
div.hint {
background-color: #EEE;
border: 1px solid #CCC;
}
div.seealso {
background-color: #EEE;
border: 1px solid #CCC;
}
div.topic {
background-color: #EEE;
}
p.admonition-title {
display: inline;
}
p.admonition-title:after {
content: ":";
}
pre, tt, code {
font-family: 'Consolas', 'Menlo', 'DejaVu Sans Mono', 'Bitstream Vera Sans Mono', monospace;
font-size: 0.9em;
}
.hll {
background-color: #FFC;
margin: 0 -12px;
padding: 0 12px;
display: block;
}
img.screenshot {
}
tt.descname, tt.descclassname, code.descname, code.descclassname {
font-size: 0.95em;
}
tt.descname, code.descname {
padding-right: 0.08em;
}
img.screenshot {
-moz-box-shadow: 2px 2px 4px #EEE;
-webkit-box-shadow: 2px 2px 4px #EEE;
box-shadow: 2px 2px 4px #EEE;
}
table.docutils {
border: 1px solid #888;
-moz-box-shadow: 2px 2px 4px #EEE;
-webkit-box-shadow: 2px 2px 4px #EEE;
box-shadow: 2px 2px 4px #EEE;
}
table.docutils td, table.docutils th {
border: 1px solid #888;
padding: 0.25em 0.7em;
}
table.field-list, table.footnote {
border: none;
-moz-box-shadow: none;
-webkit-box-shadow: none;
box-shadow: none;
}
table.footnote {
margin: 15px 0;
width: 100%;
border: 1px solid #EEE;
background: #FDFDFD;
font-size: 0.9em;
}
table.footnote + table.footnote {
margin-top: -15px;
border-top: none;
}
table.field-list th {
padding: 0 0.8em 0 0;
}
table.field-list td {
padding: 0;
}
table.field-list p {
margin-bottom: 0.8em;
}
/* Cloned from
* https://github.com/sphinx-doc/sphinx/commit/ef60dbfce09286b20b7385333d63a60321784e68
*/
.field-name {
-moz-hyphens: manual;
-ms-hyphens: manual;
-webkit-hyphens: manual;
hyphens: manual;
}
table.footnote td.label {
width: .1px;
padding: 0.3em 0 0.3em 0.5em;
}
table.footnote td {
padding: 0.3em 0.5em;
}
dl {
margin-left: 0;
margin-right: 0;
margin-top: 0;
padding: 0;
}
dl dd {
margin-left: 30px;
}
blockquote {
margin: 0 0 0 30px;
padding: 0;
}
ul, ol {
/* Matches the 30px from the narrow-screen "li > ul" selector below */
margin: 10px 0 10px 30px;
padding: 0;
}
pre {
background: unset;
padding: 7px 30px;
margin: 15px 0px;
line-height: 1.3em;
}
div.viewcode-block:target {
background: #ffd;
}
dl pre, blockquote pre, li pre {
margin-left: 0;
padding-left: 30px;
}
tt, code {
background-color: #ecf0f3;
color: #222;
/* padding: 1px 2px; */
}
tt.xref, code.xref, a tt {
background-color: #FBFBFB;
border-bottom: 1px solid #fff;
}
a.reference {
text-decoration: none;
border-bottom: 1px dotted #004B6B;
}
a.reference:hover {
border-bottom: 1px solid #6D4100;
}
/* Don't put an underline on images */
a.image-reference, a.image-reference:hover {
border-bottom: none;
}
a.footnote-reference {
text-decoration: none;
font-size: 0.7em;
vertical-align: top;
border-bottom: 1px dotted #004B6B;
}
a.footnote-reference:hover {
border-bottom: 1px solid #6D4100;
}
a:hover tt, a:hover code {
background: #EEE;
}
@media screen and (max-width: 940px) {
body {
margin: 0;
padding: 20px 30px;
}
div.documentwrapper {
float: none;
background: #fff;
margin-left: 0;
margin-top: 0;
margin-right: 0;
margin-bottom: 0;
}
div.sphinxsidebar {
display: block;
float: none;
width: unset;
margin: 50px -30px -20px -30px;
padding: 10px 20px;
background: #333;
color: #FFF;
}
div.sphinxsidebar h3, div.sphinxsidebar h4, div.sphinxsidebar p,
div.sphinxsidebar h3 a {
color: #fff;
}
div.sphinxsidebar a {
color: #AAA;
}
div.sphinxsidebar p.logo {
display: none;
}
div.document {
width: 100%;
margin: 0;
}
div.footer {
display: none;
}
div.bodywrapper {
margin: 0;
}
div.body {
min-height: 0;
min-width: auto; /* fixes width on small screens, breaks .hll */
padding: 0;
}
.hll {
/* "fixes" the breakage */
width: max-content;
}
.rtd_doc_footer {
display: none;
}
.document {
width: auto;
}
.footer {
width: auto;
}
.github {
display: none;
}
ul {
margin-left: 0;
}
li > ul {
/* Matches the 30px from the "ul, ol" selector above */
margin-left: 30px;
}
}
/* misc. */
.revsys-inline {
display: none!important;
}
/* Hide ugly table cell borders in ..bibliography:: directive output */
table.docutils.citation, table.docutils.citation td, table.docutils.citation th {
border: none;
/* Below needed in some edge cases; if not applied, bottom shadows appear */
-moz-box-shadow: none;
-webkit-box-shadow: none;
box-shadow: none;
}
/* relbar */
.related {
line-height: 30px;
width: 100%;
font-size: 0.9rem;
}
.related.top {
border-bottom: 1px solid #EEE;
margin-bottom: 20px;
}
.related.bottom {
border-top: 1px solid #EEE;
}
.related ul {
padding: 0;
margin: 0;
list-style: none;
}
.related li {
display: inline;
}
nav#rellinks {
float: right;
}
nav#rellinks li+li:before {
content: "|";
}
nav#breadcrumbs li+li:before {
content: "\00BB";
}
/* Hide certain items when printing */
@media print {
div.related {
display: none;
}
}
img.github {
position: absolute;
top: 0;
border: 0;
right: 0;
}
================================================
FILE: docs/_static/basic.css
================================================
/*
* Sphinx stylesheet -- basic theme.
*/
/* -- main layout ----------------------------------------------------------- */
div.clearer {
clear: both;
}
div.section::after {
display: block;
content: '';
clear: left;
}
/* -- relbar ---------------------------------------------------------------- */
div.related {
width: 100%;
font-size: 90%;
}
div.related h3 {
display: none;
}
div.related ul {
margin: 0;
padding: 0 0 0 10px;
list-style: none;
}
div.related li {
display: inline;
}
div.related li.right {
float: right;
margin-right: 5px;
}
/* -- sidebar --------------------------------------------------------------- */
div.sphinxsidebarwrapper {
padding: 10px 5px 0 10px;
}
div.sphinxsidebar {
float: left;
width: 230px;
margin-left: -100%;
font-size: 90%;
word-wrap: break-word;
overflow-wrap : break-word;
}
div.sphinxsidebar ul {
list-style: none;
}
div.sphinxsidebar ul ul,
div.sphinxsidebar ul.want-points {
margin-left: 20px;
list-style: square;
}
div.sphinxsidebar ul ul {
margin-top: 0;
margin-bottom: 0;
}
div.sphinxsidebar form {
margin-top: 10px;
}
div.sphinxsidebar input {
border: 1px solid #98dbcc;
font-family: sans-serif;
font-size: 1em;
}
div.sphinxsidebar #searchbox form.search {
overflow: hidden;
}
div.sphinxsidebar #searchbox input[type="text"] {
float: left;
width: 80%;
padding: 0.25em;
box-sizing: border-box;
}
div.sphinxsidebar #searchbox input[type="submit"] {
float: left;
width: 20%;
border-left: none;
padding: 0.25em;
box-sizing: border-box;
}
img {
border: 0;
max-width: 100%;
}
/* -- search page ----------------------------------------------------------- */
ul.search {
margin-top: 10px;
}
ul.search li {
padding: 5px 0;
}
ul.search li a {
font-weight: bold;
}
ul.search li p.context {
color: #888;
margin: 2px 0 0 30px;
text-align: left;
}
ul.keywordmatches li.goodmatch a {
font-weight: bold;
}
/* -- index page ------------------------------------------------------------ */
table.contentstable {
width: 90%;
margin-left: auto;
margin-right: auto;
}
table.contentstable p.biglink {
line-height: 150%;
}
a.biglink {
font-size: 1.3em;
}
span.linkdescr {
font-style: italic;
padding-top: 5px;
font-size: 90%;
}
/* -- general index --------------------------------------------------------- */
table.indextable {
width: 100%;
}
table.indextable td {
text-align: left;
vertical-align: top;
}
table.indextable ul {
margin-top: 0;
margin-bottom: 0;
list-style-type: none;
}
table.indextable > tbody > tr > td > ul {
padding-left: 0em;
}
table.indextable tr.pcap {
height: 10px;
}
table.indextable tr.cap {
margin-top: 10px;
background-color: #f2f2f2;
}
img.toggler {
margin-right: 3px;
margin-top: 3px;
cursor: pointer;
}
div.modindex-jumpbox {
border-top: 1px solid #ddd;
border-bottom: 1px solid #ddd;
margin: 1em 0 1em 0;
padding: 0.4em;
}
div.genindex-jumpbox {
border-top: 1px solid #ddd;
border-bottom: 1px solid #ddd;
margin: 1em 0 1em 0;
padding: 0.4em;
}
/* -- domain module index --------------------------------------------------- */
table.modindextable td {
padding: 2px;
border-collapse: collapse;
}
/* -- general body styles --------------------------------------------------- */
div.body {
min-width: inherit;
max-width: 800px;
}
div.body p, div.body dd, div.body li, div.body blockquote {
-moz-hyphens: auto;
-ms-hyphens: auto;
-webkit-hyphens: auto;
hyphens: auto;
}
a.headerlink {
visibility: hidden;
}
a:visited {
color: #551A8B;
}
h1:hover > a.headerlink,
h2:hover > a.headerlink,
h3:hover > a.headerlink,
h4:hover > a.headerlink,
h5:hover > a.headerlink,
h6:hover > a.headerlink,
dt:hover > a.headerlink,
caption:hover > a.headerlink,
p.caption:hover > a.headerlink,
div.code-block-caption:hover > a.headerlink {
visibility: visible;
}
div.body p.caption {
text-align: inherit;
}
div.body td {
text-align: left;
}
.first {
margin-top: 0 !important;
}
p.rubric {
margin-top: 30px;
font-weight: bold;
}
img.align-left, figure.align-left, .figure.align-left, object.align-left {
clear: left;
float: left;
margin-right: 1em;
}
img.align-right, figure.align-right, .figure.align-right, object.align-right {
clear: right;
float: right;
margin-left: 1em;
}
img.align-center, figure.align-center, .figure.align-center, object.align-center {
display: block;
margin-left: auto;
margin-right: auto;
}
img.align-default, figure.align-default, .figure.align-default {
display: block;
margin-left: auto;
margin-right: auto;
}
.align-left {
text-align: left;
}
.align-center {
text-align: center;
}
.align-default {
text-align: center;
}
.align-right {
text-align: right;
}
/* -- sidebars -------------------------------------------------------------- */
div.sidebar,
aside.sidebar {
margin: 0 0 0.5em 1em;
border: 1px solid #ddb;
padding: 7px;
background-color: #ffe;
width: 40%;
float: right;
clear: right;
overflow-x: auto;
}
p.sidebar-title {
font-weight: bold;
}
nav.contents,
aside.topic,
div.admonition, div.topic, blockquote {
clear: left;
}
/* -- topics ---------------------------------------------------------------- */
nav.contents,
aside.topic,
div.topic {
border: 1px solid #ccc;
padding: 7px;
margin: 10px 0 10px 0;
}
p.topic-title {
font-size: 1.1em;
font-weight: bold;
margin-top: 10px;
}
/* -- admonitions ----------------------------------------------------------- */
div.admonition {
margin-top: 10px;
margin-bottom: 10px;
padding: 7px;
}
div.admonition dt {
font-weight: bold;
}
p.admonition-title {
margin: 0px 10px 5px 0px;
font-weight: bold;
}
div.body p.centered {
text-align: center;
margin-top: 25px;
}
/* -- content of sidebars/topics/admonitions -------------------------------- */
div.sidebar > :last-child,
aside.sidebar > :last-child,
nav.contents > :last-child,
aside.topic > :last-child,
div.topic > :last-child,
div.admonition > :last-child {
margin-bottom: 0;
}
div.sidebar::after,
aside.sidebar::after,
nav.contents::after,
aside.topic::after,
div.topic::after,
div.admonition::after,
blockquote::after {
display: block;
content: '';
clear: both;
}
/* -- tables ---------------------------------------------------------------- */
table.docutils {
margin-top: 10px;
margin-bottom: 10px;
border: 0;
border-collapse: collapse;
}
table.align-center {
margin-left: auto;
margin-right: auto;
}
table.align-default {
margin-left: auto;
margin-right: auto;
}
table caption span.caption-number {
font-style: italic;
}
table caption span.caption-text {
}
table.docutils td, table.docutils th {
padding: 1px 8px 1px 5px;
border-top: 0;
border-left: 0;
border-right: 0;
border-bottom: 1px solid #aaa;
}
th {
text-align: left;
padding-right: 5px;
}
table.citation {
border-left: solid 1px gray;
margin-left: 1px;
}
table.citation td {
border-bottom: none;
}
th > :first-child,
td > :first-child {
margin-top: 0px;
}
th > :last-child,
td > :last-child {
margin-bottom: 0px;
}
/* -- figures --------------------------------------------------------------- */
div.figure, figure {
margin: 0.5em;
padding: 0.5em;
}
div.figure p.caption, figcaption {
padding: 0.3em;
}
div.figure p.caption span.caption-number,
figcaption span.caption-number {
font-style: italic;
}
div.figure p.caption span.caption-text,
figcaption span.caption-text {
}
/* -- field list styles ----------------------------------------------------- */
table.field-list td, table.field-list th {
border: 0 !important;
}
.field-list ul {
margin: 0;
padding-left: 1em;
}
.field-list p {
margin: 0;
}
.field-name {
-moz-hyphens: manual;
-ms-hyphens: manual;
-webkit-hyphens: manual;
hyphens: manual;
}
/* -- hlist styles ---------------------------------------------------------- */
table.hlist {
margin: 1em 0;
}
table.hlist td {
vertical-align: top;
}
/* -- object description styles --------------------------------------------- */
.sig {
font-family: 'Consolas', 'Menlo', 'DejaVu Sans Mono', 'Bitstream Vera Sans Mono', monospace;
}
.sig-name, code.descname {
background-color: transparent;
font-weight: bold;
}
.sig-name {
font-size: 1.1em;
}
code.descname {
font-size: 1.2em;
}
.sig-prename, code.descclassname {
background-color: transparent;
}
.optional {
font-size: 1.3em;
}
.sig-paren {
font-size: larger;
}
.sig-param.n {
font-style: italic;
}
/* C++ specific styling */
.sig-inline.c-texpr,
.sig-inline.cpp-texpr {
font-family: unset;
}
.sig.c .k, .sig.c .kt,
.sig.cpp .k, .sig.cpp .kt {
color: #0033B3;
}
.sig.c .m,
.sig.cpp .m {
color: #1750EB;
}
.sig.c .s, .sig.c .sc,
.sig.cpp .s, .sig.cpp .sc {
color: #067D17;
}
/* -- other body styles ----------------------------------------------------- */
ol.arabic {
list-style: decimal;
}
ol.loweralpha {
list-style: lower-alpha;
}
ol.upperalpha {
list-style: upper-alpha;
}
ol.lowerroman {
list-style: lower-roman;
}
ol.upperroman {
list-style: upper-roman;
}
:not(li) > ol > li:first-child > :first-child,
:not(li) > ul > li:first-child > :first-child {
margin-top: 0px;
}
:not(li) > ol > li:last-child > :last-child,
:not(li) > ul > li:last-child > :last-child {
margin-bottom: 0px;
}
ol.simple ol p,
ol.simple ul p,
ul.simple ol p,
ul.simple ul p {
margin-top: 0;
}
ol.simple > li:not(:first-child) > p,
ul.simple > li:not(:first-child) > p {
margin-top: 0;
}
ol.simple p,
ul.simple p {
margin-bottom: 0;
}
aside.footnote > span,
div.citation > span {
float: left;
}
aside.footnote > span:last-of-type,
div.citation > span:last-of-type {
padding-right: 0.5em;
}
aside.footnote > p {
margin-left: 2em;
}
div.citation > p {
margin-left: 4em;
}
aside.footnote > p:last-of-type,
div.citation > p:last-of-type {
margin-bottom: 0em;
}
aside.footnote > p:last-of-type:after,
div.citation > p:last-of-type:after {
content: "";
clear: both;
}
dl.field-list {
display: grid;
grid-template-columns: fit-content(30%) auto;
}
dl.field-list > dt {
font-weight: bold;
word-break: break-word;
padding-left: 0.5em;
padding-right: 5px;
}
dl.field-list > dd {
padding-left: 0.5em;
margin-top: 0em;
margin-left: 0em;
margin-bottom: 0em;
}
dl {
margin-bottom: 15px;
}
dd > :first-child {
margin-top: 0px;
}
dd ul, dd table {
margin-bottom: 10px;
}
dd {
margin-top: 3px;
margin-bottom: 10px;
margin-left: 30px;
}
.sig dd {
margin-top: 0px;
margin-bottom: 0px;
}
.sig dl {
margin-top: 0px;
margin-bottom: 0px;
}
dl > dd:last-child,
dl > dd:last-child > :last-child {
margin-bottom: 0;
}
dt:target, span.highlighted {
background-color: #fbe54e;
}
rect.highlighted {
fill: #fbe54e;
}
dl.glossary dt {
font-weight: bold;
font-size: 1.1em;
}
.versionmodified {
font-style: italic;
}
.system-message {
background-color: #fda;
padding: 5px;
border: 3px solid red;
}
.footnote:target {
background-color: #ffa;
}
.line-block {
display: block;
margin-top: 1em;
margin-bottom: 1em;
}
.line-block .line-block {
margin-top: 0;
margin-bottom: 0;
margin-left: 1.5em;
}
.guilabel, .menuselection {
font-family: sans-serif;
}
.accelerator {
text-decoration: underline;
}
.classifier {
font-style: oblique;
}
.classifier:before {
font-style: normal;
margin: 0 0.5em;
content: ":";
display: inline-block;
}
abbr, acronym {
border-bottom: dotted 1px;
cursor: help;
}
.translated {
background-color: rgba(207, 255, 207, 0.2)
}
.untranslated {
background-color: rgba(255, 207, 207, 0.2)
}
/* -- code displays --------------------------------------------------------- */
pre {
overflow: auto;
overflow-y: hidden; /* fixes display issues on Chrome browsers */
}
pre, div[class*="highlight-"] {
clear: both;
}
span.pre {
-moz-hyphens: none;
-ms-hyphens: none;
-webkit-hyphens: none;
hyphens: none;
white-space: nowrap;
}
div[class*="highlight-"] {
margin: 1em 0;
}
td.linenos pre {
border: 0;
background-color: transparent;
color: #aaa;
}
table.highlighttable {
display: block;
}
table.highlighttable tbody {
display: block;
}
table.highlighttable tr {
display: flex;
}
table.highlighttable td {
margin: 0;
padding: 0;
}
table.highlighttable td.linenos {
padding-right: 0.5em;
}
table.highlighttable td.code {
flex: 1;
overflow: hidden;
}
.highlight .hll {
display: block;
}
div.highlight pre,
table.highlighttable pre {
margin: 0;
}
div.code-block-caption + div {
margin-top: 0;
}
div.code-block-caption {
margin-top: 1em;
padding: 2px 5px;
font-size: small;
}
div.code-block-caption code {
background-color: transparent;
}
table.highlighttable td.linenos,
span.linenos,
div.highlight span.gp { /* gp: Generic.Prompt */
user-select: none;
-webkit-user-select: text; /* Safari fallback only */
-webkit-user-select: none; /* Chrome/Safari */
-moz-user-select: none; /* Firefox */
-ms-user-select: none; /* IE10+ */
}
div.code-block-caption span.caption-number {
padding: 0.1em 0.3em;
font-style: italic;
}
div.code-block-caption span.caption-text {
}
div.literal-block-wrapper {
margin: 1em 0;
}
code.xref, a code {
background-color: transparent;
font-weight: bold;
}
h1 code, h2 code, h3 code, h4 code, h5 code, h6 code {
background-color: transparent;
}
.viewcode-link {
float: right;
}
.viewcode-back {
float: right;
font-family: sans-serif;
}
div.viewcode-block:target {
margin: -1px -10px;
padding: 0 10px;
}
/* -- math display ---------------------------------------------------------- */
img.math {
vertical-align: middle;
}
div.body div.math p {
text-align: center;
}
span.eqno {
float: right;
}
span.eqno a.headerlink {
position: absolute;
z-index: 1;
}
div.math:hover a.headerlink {
visibility: visible;
}
/* -- printout stylesheet --------------------------------------------------- */
@media print {
div.document,
div.documentwrapper,
div.bodywrapper {
margin: 0 !important;
width: 100%;
}
div.sphinxsidebar,
div.related,
div.footer,
#top-link {
display: none;
}
}
================================================
FILE: docs/_static/custom.css
================================================
/* This file intentionally left blank. */
================================================
FILE: docs/_static/doctools.js
================================================
/*
* Base JavaScript utilities for all Sphinx HTML documentation.
*/
"use strict";
const BLACKLISTED_KEY_CONTROL_ELEMENTS = new Set([
"TEXTAREA",
"INPUT",
"SELECT",
"BUTTON",
]);
const _ready = (callback) => {
if (document.readyState !== "loading") {
callback();
} else {
document.addEventListener("DOMContentLoaded", callback);
}
};
/**
* Small JavaScript module for the documentation.
*/
const Documentation = {
init: () => {
Documentation.initDomainIndexTable();
Documentation.initOnKeyListeners();
},
/**
* i18n support
*/
TRANSLATIONS: {},
PLURAL_EXPR: (n) => (n === 1 ? 0 : 1),
LOCALE: "unknown",
// gettext and ngettext don't access this so that the functions
// can safely bound to a different name (_ = Documentation.gettext)
gettext: (string) => {
const translated = Documentation.TRANSLATIONS[string];
switch (typeof translated) {
case "undefined":
return string; // no translation
case "string":
return translated; // translation exists
default:
return translated[0]; // (singular, plural) translation tuple exists
}
},
ngettext: (singular, plural, n) => {
const translated = Documentation.TRANSLATIONS[singular];
if (typeof translated !== "undefined")
return translated[Documentation.PLURAL_EXPR(n)];
return n === 1 ? singular : plural;
},
addTranslations: (catalog) => {
Object.assign(Documentation.TRANSLATIONS, catalog.messages);
Documentation.PLURAL_EXPR = new Function(
"n",
`return (${catalog.plural_expr})`
);
Documentation.LOCALE = catalog.locale;
},
/**
* helper function to focus on search bar
*/
focusSearchBar: () => {
document.querySelectorAll("input[name=q]")[0]?.focus();
},
/**
* Initialise the domain index toggle buttons
*/
initDomainIndexTable: () => {
const toggler = (el) => {
const idNumber = el.id.substr(7);
const toggledRows = document.querySelectorAll(`tr.cg-${idNumber}`);
if (el.src.substr(-9) === "minus.png") {
el.src = `${el.src.substr(0, el.src.length - 9)}plus.png`;
toggledRows.forEach((el) => (el.style.display = "none"));
} else {
el.src = `${el.src.substr(0, el.src.length - 8)}minus.png`;
toggledRows.forEach((el) => (el.style.display = ""));
}
};
const togglerElements = document.querySelectorAll("img.toggler");
togglerElements.forEach((el) =>
el.addEventListener("click", (event) => toggler(event.currentTarget))
);
togglerElements.forEach((el) => (el.style.display = ""));
if (DOCUMENTATION_OPTIONS.COLLAPSE_INDEX) togglerElements.forEach(toggler);
},
initOnKeyListeners: () => {
// only install a listener if it is really needed
if (
!DOCUMENTATION_OPTIONS.NAVIGATION_WITH_KEYS &&
!DOCUMENTATION_OPTIONS.ENABLE_SEARCH_SHORTCUTS
)
return;
document.addEventListener("keydown", (event) => {
// bail for input elements
if (BLACKLISTED_KEY_CONTROL_ELEMENTS.has(document.activeElement.tagName)) return;
// bail with special keys
if (event.altKey || event.ctrlKey || event.metaKey) return;
if (!event.shiftKey) {
switch (event.key) {
case "ArrowLeft":
if (!DOCUMENTATION_OPTIONS.NAVIGATION_WITH_KEYS) break;
const prevLink = document.querySelector('link[rel="prev"]');
if (prevLink && prevLink.href) {
window.location.href = prevLink.href;
event.preventDefault();
}
break;
case "ArrowRight":
if (!DOCUMENTATION_OPTIONS.NAVIGATION_WITH_KEYS) break;
const nextLink = document.querySelector('link[rel="next"]');
if (nextLink && nextLink.href) {
window.location.href = nextLink.href;
event.preventDefault();
}
break;
}
}
// some keyboard layouts may need Shift to get /
switch (event.key) {
case "/":
if (!DOCUMENTATION_OPTIONS.ENABLE_SEARCH_SHORTCUTS) break;
Documentation.focusSearchBar();
event.preventDefault();
}
});
},
};
// quick alias for translations
const _ = Documentation.gettext;
_ready(Documentation.init);
================================================
FILE: docs/_static/documentation_options.js
================================================
const DOCUMENTATION_OPTIONS = {
VERSION: '2.5.2',
LANGUAGE: 'en',
COLLAPSE_INDEX: false,
BUILDER: 'html',
FILE_SUFFIX: '.html',
LINK_SUFFIX: '.html',
HAS_SOURCE: true,
SOURCELINK_SUFFIX: '.txt',
NAVIGATION_WITH_KEYS: false,
SHOW_SEARCH_SUMMARY: true,
ENABLE_SEARCH_SHORTCUTS: true,
};
================================================
FILE: docs/_static/language_data.js
================================================
/*
* This script contains the language-specific data used by searchtools.js,
* namely the list of stopwords, stemmer, scorer and splitter.
*/
var stopwords = ["a", "and", "are", "as", "at", "be", "but", "by", "for", "if", "in", "into", "is", "it", "near", "no", "not", "of", "on", "or", "such", "that", "the", "their", "then", "there", "these", "they", "this", "to", "was", "will", "with"];
/* Non-minified version is copied as a separate JS file, if available */
/**
* Porter Stemmer
*/
var Stemmer = function() {
var step2list = {
ational: 'ate',
tional: 'tion',
enci: 'ence',
anci: 'ance',
izer: 'ize',
bli: 'ble',
alli: 'al',
entli: 'ent',
eli: 'e',
ousli: 'ous',
ization: 'ize',
ation: 'ate',
ator: 'ate',
alism: 'al',
iveness: 'ive',
fulness: 'ful',
ousness: 'ous',
aliti: 'al',
iviti: 'ive',
biliti: 'ble',
logi: 'log'
};
var step3list = {
icate: 'ic',
ative: '',
alize: 'al',
iciti: 'ic',
ical: 'ic',
ful: '',
ness: ''
};
var c = "[^aeiou]"; // consonant
var v = "[aeiouy]"; // vowel
var C = c + "[^aeiouy]*"; // consonant sequence
var V = v + "[aeiou]*"; // vowel sequence
var mgr0 = "^(" + C + ")?" + V + C; // [C]VC... is m>0
var meq1 = "^(" + C + ")?" + V + C + "(" + V + ")?$"; // [C]VC[V] is m=1
var mgr1 = "^(" + C + ")?" + V + C + V + C; // [C]VCVC... is m>1
var s_v = "^(" + C + ")?" + v; // vowel in stem
this.stemWord = function (w) {
var stem;
var suffix;
var firstch;
var origword = w;
if (w.length < 3)
return w;
var re;
var re2;
var re3;
var re4;
firstch = w.substr(0,1);
if (firstch == "y")
w = firstch.toUpperCase() + w.substr(1);
// Step 1a
re = /^(.+?)(ss|i)es$/;
re2 = /^(.+?)([^s])s$/;
if (re.test(w))
w = w.replace(re,"$1$2");
else if (re2.test(w))
w = w.replace(re2,"$1$2");
// Step 1b
re = /^(.+?)eed$/;
re2 = /^(.+?)(ed|ing)$/;
if (re.test(w)) {
var fp = re.exec(w);
re = new RegExp(mgr0);
if (re.test(fp[1])) {
re = /.$/;
w = w.replace(re,"");
}
}
else if (re2.test(w)) {
var fp = re2.exec(w);
stem = fp[1];
re2 = new RegExp(s_v);
if (re2.test(stem)) {
w = stem;
re2 = /(at|bl|iz)$/;
re3 = new RegExp("([^aeiouylsz])\\1$");
re4 = new RegExp("^" + C + v + "[^aeiouwxy]$");
if (re2.test(w))
w = w + "e";
else if (re3.test(w)) {
re = /.$/;
w = w.replace(re,"");
}
else if (re4.test(w))
w = w + "e";
}
}
// Step 1c
re = /^(.+?)y$/;
if (re.test(w)) {
var fp = re.exec(w);
stem = fp[1];
re = new RegExp(s_v);
if (re.test(stem))
w = stem + "i";
}
// Step 2
re = /^(.+?)(ational|tional|enci|anci|izer|bli|alli|entli|eli|ousli|ization|ation|ator|alism|iveness|fulness|ousness|aliti|iviti|biliti|logi)$/;
if (re.test(w)) {
var fp = re.exec(w);
stem = fp[1];
suffix = fp[2];
re = new RegExp(mgr0);
if (re.test(stem))
w = stem + step2list[suffix];
}
// Step 3
re = /^(.+?)(icate|ative|alize|iciti|ical|ful|ness)$/;
if (re.test(w)) {
var fp = re.exec(w);
stem = fp[1];
suffix = fp[2];
re = new RegExp(mgr0);
if (re.test(stem))
w = stem + step3list[suffix];
}
// Step 4
re = /^(.+?)(al|ance|ence|er|ic|able|ible|ant|ement|ment|ent|ou|ism|ate|iti|ous|ive|ize)$/;
re2 = /^(.+?)(s|t)(ion)$/;
if (re.test(w)) {
var fp = re.exec(w);
stem = fp[1];
re = new RegExp(mgr1);
if (re.test(stem))
w = stem;
}
else if (re2.test(w)) {
var fp = re2.exec(w);
stem = fp[1] + fp[2];
re2 = new RegExp(mgr1);
if (re2.test(stem))
w = stem;
}
// Step 5
re = /^(.+?)e$/;
if (re.test(w)) {
var fp = re.exec(w);
stem = fp[1];
re = new RegExp(mgr1);
re2 = new RegExp(meq1);
re3 = new RegExp("^" + C + v + "[^aeiouwxy]$");
if (re.test(stem) || (re2.test(stem) && !(re3.test(stem))))
w = stem;
}
re = /ll$/;
re2 = new RegExp(mgr1);
if (re.test(w) && re2.test(w)) {
re = /.$/;
w = w.replace(re,"");
}
// and turn initial Y back to y
if (firstch == "y")
w = firstch.toLowerCase() + w.substr(1);
return w;
}
}
================================================
FILE: docs/_static/pygments.css
================================================
pre { line-height: 125%; }
td.linenos .normal { color: inherit; background-color: transparent; padding-left: 5px; padding-right: 5px; }
span.linenos { color: inherit; background-color: transparent; padding-left: 5px; padding-right: 5px; }
td.linenos .special { color: #000000; background-color: #ffffc0; padding-left: 5px; padding-right: 5px; }
span.linenos.special { color: #000000; background-color: #ffffc0; padding-left: 5px; padding-right: 5px; }
.highlight .hll { background-color: #ffffcc }
.highlight { background: #f8f8f8; }
.highlight .c { color: #8f5902; font-style: italic } /* Comment */
.highlight .err { color: #a40000; border: 1px solid #ef2929 } /* Error */
.highlight .g { color: #000000 } /* Generic */
.highlight .k { color: #004461; font-weight: bold } /* Keyword */
.highlight .l { color: #000000 } /* Literal */
.highlight .n { color: #000000 } /* Name */
.highlight .o { color: #582800 } /* Operator */
.highlight .x { color: #000000 } /* Other */
.highlight .p { color: #000000; font-weight: bold } /* Punctuation */
.highlight .ch { color: #8f5902; font-style: italic } /* Comment.Hashbang */
.highlight .cm { color: #8f5902; font-style: italic } /* Comment.Multiline */
.highlight .cp { color: #8f5902 } /* Comment.Preproc */
.highlight .cpf { color: #8f5902; font-style: italic } /* Comment.PreprocFile */
.highlight .c1 { color: #8f5902; font-style: italic } /* Comment.Single */
.highlight .cs { color: #8f5902; font-style: italic } /* Comment.Special */
.highlight .gd { color: #a40000 } /* Generic.Deleted */
.highlight .ge { color: #000000; font-style: italic } /* Generic.Emph */
.highlight .ges { color: #000000 } /* Generic.EmphStrong */
.highlight .gr { color: #ef2929 } /* Generic.Error */
.highlight .gh { color: #000080; font-weight: bold } /* Generic.Heading */
.highlight .gi { color: #00A000 } /* Generic.Inserted */
.highlight .go { color: #888888 } /* Generic.Output */
.highlight .gp { color: #745334 } /* Generic.Prompt */
.highlight .gs { color: #000000; font-weight: bold } /* Generic.Strong */
.highlight .gu { color: #800080; font-weight: bold } /* Generic.Subheading */
.highlight .gt { color: #a40000; font-weight: bold } /* Generic.Traceback */
.highlight .kc { color: #004461; font-weight: bold } /* Keyword.Constant */
.highlight .kd { color: #004461; font-weight: bold } /* Keyword.Declaration */
.highlight .kn { color: #004461; font-weight: bold } /* Keyword.Namespace */
.highlight .kp { color: #004461; font-weight: bold } /* Keyword.Pseudo */
.highlight .kr { color: #004461; font-weight: bold } /* Keyword.Reserved */
.highlight .kt { color: #004461; font-weight: bold } /* Keyword.Type */
.highlight .ld { color: #000000 } /* Literal.Date */
.highlight .m { color: #990000 } /* Literal.Number */
.highlight .s { color: #4e9a06 } /* Literal.String */
.highlight .na { color: #c4a000 } /* Name.Attribute */
.highlight .nb { color: #004461 } /* Name.Builtin */
.highlight .nc { color: #000000 } /* Name.Class */
.highlight .no { color: #000000 } /* Name.Constant */
.highlight .nd { color: #888888 } /* Name.Decorator */
.highlight .ni { color: #ce5c00 } /* Name.Entity */
.highlight .ne { color: #cc0000; font-weight: bold } /* Name.Exception */
.highlight .nf { color: #000000 } /* Name.Function */
.highlight .nl { color: #f57900 } /* Name.Label */
.highlight .nn { color: #000000 } /* Name.Namespace */
.highlight .nx { color: #000000 } /* Name.Other */
.highlight .py { color: #000000 } /* Name.Property */
.highlight .nt { color: #004461; font-weight: bold } /* Name.Tag */
.highlight .nv { color: #000000 } /* Name.Variable */
.highlight .ow { color: #004461; font-weight: bold } /* Operator.Word */
.highlight .pm { color: #000000; font-weight: bold } /* Punctuation.Marker */
.highlight .w { color: #f8f8f8 } /* Text.Whitespace */
.highlight .mb { color: #990000 } /* Literal.Number.Bin */
.highlight .mf { color: #990000 } /* Literal.Number.Float */
.highlight .mh { color: #990000 } /* Literal.Number.Hex */
.highlight .mi { color: #990000 } /* Literal.Number.Integer */
.highlight .mo { color: #990000 } /* Literal.Number.Oct */
.highlight .sa { color: #4e9a06 } /* Literal.String.Affix */
.highlight .sb { color: #4e9a06 } /* Literal.String.Backtick */
.highlight .sc { color: #4e9a06 } /* Literal.String.Char */
.highlight .dl { color: #4e9a06 } /* Literal.String.Delimiter */
.highlight .sd { color: #8f5902; font-style: italic } /* Literal.String.Doc */
.highlight .s2 { color: #4e9a06 } /* Literal.String.Double */
.highlight .se { color: #4e9a06 } /* Literal.String.Escape */
.highlight .sh { color: #4e9a06 } /* Literal.String.Heredoc */
.highlight .si { color: #4e9a06 } /* Literal.String.Interpol */
.highlight .sx { color: #4e9a06 } /* Literal.String.Other */
.highlight .sr { color: #4e9a06 } /* Literal.String.Regex */
.highlight .s1 { color: #4e9a06 } /* Literal.String.Single */
.highlight .ss { color: #4e9a06 } /* Literal.String.Symbol */
.highlight .bp { color: #3465a4 } /* Name.Builtin.Pseudo */
.highlight .fm { color: #000000 } /* Name.Function.Magic */
.highlight .vc { color: #000000 } /* Name.Variable.Class */
.highlight .vg { color: #000000 } /* Name.Variable.Global */
.highlight .vi { color: #000000 } /* Name.Variable.Instance */
.highlight .vm { color: #000000 } /* Name.Variable.Magic */
.highlight .il { color: #990000 } /* Literal.Number.Integer.Long */
================================================
FILE: docs/_static/searchtools.js
================================================
/*
* Sphinx JavaScript utilities for the full-text search.
*/
"use strict";
/**
* Simple result scoring code.
*/
if (typeof Scorer === "undefined") {
var Scorer = {
// Implement the following function to further tweak the score for each result
// The function takes a result array [docname, title, anchor, descr, score, filename]
// and returns the new score.
/*
score: result => {
const [docname, title, anchor, descr, score, filename, kind] = result
return score
},
*/
// query matches the full name of an object
objNameMatch: 11,
// or matches in the last dotted part of the object name
objPartialMatch: 6,
// Additive scores depending on the priority of the object
objPrio: {
0: 15, // used to be importantResults
1: 5, // used to be objectResults
2: -5, // used to be unimportantResults
},
// Used when the priority is not in the mapping.
objPrioDefault: 0,
// query found in title
title: 15,
partialTitle: 7,
// query found in terms
term: 5,
partialTerm: 2,
};
}
// Global search result kind enum, used by themes to style search results.
class SearchResultKind {
static get index() { return "index"; }
static get object() { return "object"; }
static get text() { return "text"; }
static get title() { return "title"; }
}
const _removeChildren = (element) => {
while (element && element.lastChild) element.removeChild(element.lastChild);
};
/**
* See https://developer.mozilla.org/en-US/docs/Web/JavaScript/Guide/Regular_Expressions#escaping
*/
const _escapeRegExp = (string) =>
string.replace(/[.*+\-?^${}()|[\]\\]/g, "\\$&"); // $& means the whole matched string
const _displayItem = (item, searchTerms, highlightTerms) => {
const docBuilder = DOCUMENTATION_OPTIONS.BUILDER;
const docFileSuffix = DOCUMENTATION_OPTIONS.FILE_SUFFIX;
const docLinkSuffix = DOCUMENTATION_OPTIONS.LINK_SUFFIX;
const showSearchSummary = DOCUMENTATION_OPTIONS.SHOW_SEARCH_SUMMARY;
const contentRoot = document.documentElement.dataset.content_root;
const [docName, title, anchor, descr, score, _filename, kind] = item;
let listItem = document.createElement("li");
// Add a class representing the item's type:
// can be used by a theme's CSS selector for styling
// See SearchResultKind for the class names.
listItem.classList.add(`kind-${kind}`);
let requestUrl;
let linkUrl;
if (docBuilder === "dirhtml") {
// dirhtml builder
let dirname = docName + "/";
if (dirname.match(/\/index\/$/))
dirname = dirname.substring(0, dirname.length - 6);
else if (dirname === "index/") dirname = "";
requestUrl = contentRoot + dirname;
linkUrl = requestUrl;
} else {
// normal html builders
requestUrl = contentRoot + docName + docFileSuffix;
linkUrl = docName + docLinkSuffix;
}
let linkEl = listItem.appendChild(document.createElement("a"));
linkEl.href = linkUrl + anchor;
linkEl.dataset.score = score;
linkEl.innerHTML = title;
if (descr) {
listItem.appendChild(document.createElement("span")).innerHTML =
" (" + descr + ")";
// highlight search terms in the description
if (SPHINX_HIGHLIGHT_ENABLED) // set in sphinx_highlight.js
highlightTerms.forEach((term) => _highlightText(listItem, term, "highlighted"));
}
else if (showSearchSummary)
fetch(requestUrl)
.then((responseData) => responseData.text())
.then((data) => {
if (data)
listItem.appendChild(
Search.makeSearchSummary(data, searchTerms, anchor)
);
// highlight search terms in the summary
if (SPHINX_HIGHLIGHT_ENABLED) // set in sphinx_highlight.js
highlightTerms.forEach((term) => _highlightText(listItem, term, "highlighted"));
});
Search.output.appendChild(listItem);
};
const _finishSearch = (resultCount) => {
Search.stopPulse();
Search.title.innerText = _("Search Results");
if (!resultCount)
Search.status.innerText = Documentation.gettext(
"Your search did not match any documents. Please make sure that all words are spelled correctly and that you've selected enough categories."
);
else
Search.status.innerText = Documentation.ngettext(
"Search finished, found one page matching the search query.",
"Search finished, found ${resultCount} pages matching the search query.",
resultCount,
).replace('${resultCount}', resultCount);
};
const _displayNextItem = (
results,
resultCount,
searchTerms,
highlightTerms,
) => {
// results left, load the summary and display it
// this is intended to be dynamic (don't sub resultsCount)
if (results.length) {
_displayItem(results.pop(), searchTerms, highlightTerms);
setTimeout(
() => _displayNextItem(results, resultCount, searchTerms, highlightTerms),
5
);
}
// search finished, update title and status message
else _finishSearch(resultCount);
};
// Helper function used by query() to order search results.
// Each input is an array of [docname, title, anchor, descr, score, filename, kind].
// Order the results by score (in opposite order of appearance, since the
// `_displayNextItem` function uses pop() to retrieve items) and then alphabetically.
const _orderResultsByScoreThenName = (a, b) => {
const leftScore = a[4];
const rightScore = b[4];
if (leftScore === rightScore) {
// same score: sort alphabetically
const leftTitle = a[1].toLowerCase();
const rightTitle = b[1].toLowerCase();
if (leftTitle === rightTitle) return 0;
return leftTitle > rightTitle ? -1 : 1; // inverted is intentional
}
return leftScore > rightScore ? 1 : -1;
};
/**
* Default splitQuery function. Can be overridden in ``sphinx.search`` with a
* custom function per language.
*
* The regular expression works by splitting the string on consecutive characters
* that are not Unicode letters, numbers, underscores, or emoji characters.
* This is the same as ``\W+`` in Python, preserving the surrogate pair area.
*/
if (typeof splitQuery === "undefined") {
var splitQuery = (query) => query
.split(/[^\p{Letter}\p{Number}_\p{Emoji_Presentation}]+/gu)
.filter(term => term) // remove remaining empty strings
}
/**
* Search Module
*/
const Search = {
_index: null,
_queued_query: null,
_pulse_status: -1,
htmlToText: (htmlString, anchor) => {
const htmlElement = new DOMParser().parseFromString(htmlString, 'text/html');
for (const removalQuery of [".headerlink", "script", "style"]) {
htmlElement.querySelectorAll(removalQuery).forEach((el) => { el.remove() });
}
if (anchor) {
const anchorContent = htmlElement.querySelector(`[role="main"] ${anchor}`);
if (anchorContent) return anchorContent.textContent;
console.warn(
`Anchored content block not found. Sphinx search tries to obtain it via DOM query '[role=main] ${anchor}'. Check your theme or template.`
);
}
// if anchor not specified or not found, fall back to main content
const docContent = htmlElement.querySelector('[role="main"]');
if (docContent) return docContent.textContent;
console.warn(
"Content block not found. Sphinx search tries to obtain it via DOM query '[role=main]'. Check your theme or template."
);
return "";
},
init: () => {
const query = new URLSearchParams(window.location.search).get("q");
document
.querySelectorAll('input[name="q"]')
.forEach((el) => (el.value = query));
if (query) Search.performSearch(query);
},
loadIndex: (url) =>
(document.body.appendChild(document.createElement("script")).src = url),
setIndex: (index) => {
Search._index = index;
if (Search._queued_query !== null) {
const query = Search._queued_query;
Search._queued_query = null;
Search.query(query);
}
},
hasIndex: () => Search._index !== null,
deferQuery: (query) => (Search._queued_query = query),
stopPulse: () => (Search._pulse_status = -1),
startPulse: () => {
if (Search._pulse_status >= 0) return;
const pulse = () => {
Search._pulse_status = (Search._pulse_status + 1) % 4;
Search.dots.innerText = ".".repeat(Search._pulse_status);
if (Search._pulse_status >= 0) window.setTimeout(pulse, 500);
};
pulse();
},
/**
* perform a search for something (or wait until index is loaded)
*/
performSearch: (query) => {
// create the required interface elements
const searchText = document.createElement("h2");
searchText.textContent = _("Searching");
const searchSummary = document.createElement("p");
searchSummary.classList.add("search-summary");
searchSummary.innerText = "";
const searchList = document.createElement("ul");
searchList.setAttribute("role", "list");
searchList.classList.add("search");
const out = document.getElementById("search-results");
Search.title = out.appendChild(searchText);
Search.dots = Search.title.appendChild(document.createElement("span"));
Search.status = out.appendChild(searchSummary);
Search.output = out.appendChild(searchList);
const searchProgress = document.getElementById("search-progress");
// Some themes don't use the search progress node
if (searchProgress) {
searchProgress.innerText = _("Preparing search...");
}
Search.startPulse();
// index already loaded, the browser was quick!
if (Search.hasIndex()) Search.query(query);
else Search.deferQuery(query);
},
_parseQuery: (query) => {
// stem the search terms and add them to the correct list
const stemmer = new Stemmer();
const searchTerms = new Set();
const excludedTerms = new Set();
const highlightTerms = new Set();
const objectTerms = new Set(splitQuery(query.toLowerCase().trim()));
splitQuery(query.trim()).forEach((queryTerm) => {
const queryTermLower = queryTerm.toLowerCase();
// maybe skip this "word"
// stopwords array is from language_data.js
if (
stopwords.indexOf(queryTermLower) !== -1 ||
queryTerm.match(/^\d+$/)
)
return;
// stem the word
let word = stemmer.stemWord(queryTermLower);
// select the correct list
if (word[0] === "-") excludedTerms.add(word.substr(1));
else {
searchTerms.add(word);
highlightTerms.add(queryTermLower);
}
});
if (SPHINX_HIGHLIGHT_ENABLED) { // set in sphinx_highlight.js
localStorage.setItem("sphinx_highlight_terms", [...highlightTerms].join(" "))
}
// console.debug("SEARCH: searching for:");
// console.info("required: ", [...searchTerms]);
// console.info("excluded: ", [...excludedTerms]);
return [query, searchTerms, excludedTerms, highlightTerms, objectTerms];
},
/**
* execute search (requires search index to be loaded)
*/
_performSearch: (query, searchTerms, excludedTerms, highlightTerms, objectTerms) => {
const filenames = Search._index.filenames;
const docNames = Search._index.docnames;
const titles = Search._index.titles;
const allTitles = Search._index.alltitles;
const indexEntries = Search._index.indexentries;
// Collect multiple result groups to be sorted separately and then ordered.
// Each is an array of [docname, title, anchor, descr, score, filename, kind].
const normalResults = [];
const nonMainIndexResults = [];
_removeChildren(document.getElementById("search-progress"));
const queryLower = query.toLowerCase().trim();
for (const [title, foundTitles] of Object.entries(allTitles)) {
if (title.toLowerCase().trim().includes(queryLower) && (queryLower.length >= title.length/2)) {
for (const [file, id] of foundTitles) {
const score = Math.round(Scorer.title * queryLower.length / title.length);
const boost = titles[file] === title ? 1 : 0; // add a boost for document titles
normalResults.push([
docNames[file],
titles[file] !== title ? `${titles[file]} > ${title}` : title,
id !== null ? "#" + id : "",
null,
score + boost,
filenames[file],
SearchResultKind.title,
]);
}
}
}
// search for explicit entries in index directives
for (const [entry, foundEntries] of Object.entries(indexEntries)) {
if (entry.includes(queryLower) && (queryLower.length >= entry.length/2)) {
for (const [file, id, isMain] of foundEntries) {
const score = Math.round(100 * queryLower.length / entry.length);
const result = [
docNames[file],
titles[file],
id ? "#" + id : "",
null,
score,
filenames[file],
SearchResultKind.index,
];
if (isMain) {
normalResults.push(result);
} else {
nonMainIndexResults.push(result);
}
}
}
}
// lookup as object
objectTerms.forEach((term) =>
normalResults.push(...Search.performObjectSearch(term, objectTerms))
);
// lookup as search terms in fulltext
normalResults.push(...Search.performTermsSearch(searchTerms, excludedTerms));
// let the scorer override scores with a custom scoring function
if (Scorer.score) {
normalResults.forEach((item) => (item[4] = Scorer.score(item)));
nonMainIndexResults.forEach((item) => (item[4] = Scorer.score(item)));
}
// Sort each group of results by score and then alphabetically by name.
normalResults.sort(_orderResultsByScoreThenName);
nonMainIndexResults.sort(_orderResultsByScoreThenName);
// Combine the result groups in (reverse) order.
// Non-main index entries are typically arbitrary cross-references,
// so display them after other results.
let results = [...nonMainIndexResults, ...normalResults];
// remove duplicate search results
// note the reversing of results, so that in the case of duplicates, the highest-scoring entry is kept
let seen = new Set();
results = results.reverse().reduce((acc, result) => {
let resultStr = result.slice(0, 4).concat([result[5]]).map(v => String(v)).join(',');
if (!seen.has(resultStr)) {
acc.push(result);
seen.add(resultStr);
}
return acc;
}, []);
return results.reverse();
},
query: (query) => {
const [searchQuery, searchTerms, excludedTerms, highlightTerms, objectTerms] = Search._parseQuery(query);
const results = Search._performSearch(searchQuery, searchTerms, excludedTerms, highlightTerms, objectTerms);
// for debugging
//Search.lastresults = results.slice(); // a copy
// console.info("search results:", Search.lastresults);
// print the results
_displayNextItem(results, results.length, searchTerms, highlightTerms);
},
/**
* search for object names
*/
performObjectSearch: (object, objectTerms) => {
const filenames = Search._index.filenames;
const docNames = Search._index.docnames;
const objects = Search._index.objects;
const objNames = Search._index.objnames;
const titles = Search._index.titles;
const results = [];
const objectSearchCallback = (prefix, match) => {
const name = match[4]
const fullname = (prefix ? prefix + "." : "") + name;
const fullnameLower = fullname.toLowerCase();
if (fullnameLower.indexOf(object) < 0) return;
let score = 0;
const parts = fullnameLower.split(".");
// check for different match types: exact matches of full name or
// "last name" (i.e. last dotted part)
if (fullnameLower === object || parts.slice(-1)[0] === object)
score += Scorer.objNameMatch;
else if (parts.slice(-1)[0].indexOf(object) > -1)
score += Scorer.objPartialMatch; // matches in last name
const objName = objNames[match[1]][2];
const title = titles[match[0]];
// If more than one term searched for, we require other words to be
// found in the name/title/description
const otherTerms = new Set(objectTerms);
otherTerms.delete(object);
if (otherTerms.size > 0) {
const haystack = `${prefix} ${name} ${objName} ${title}`.toLowerCase();
if (
[...otherTerms].some((otherTerm) => haystack.indexOf(otherTerm) < 0)
)
return;
}
let anchor = match[3];
if (anchor === "") anchor = fullname;
else if (anchor === "-") anchor = objNames[match[1]][1] + "-" + fullname;
const descr = objName + _(", in ") + title;
// add custom score for some objects according to scorer
if (Scorer.objPrio.hasOwnProperty(match[2]))
score += Scorer.objPrio[match[2]];
else score += Scorer.objPrioDefault;
results.push([
docNames[match[0]],
fullname,
"#" + anchor,
descr,
score,
filenames[match[0]],
SearchResultKind.object,
]);
};
Object.keys(objects).forEach((prefix) =>
objects[prefix].forEach((array) =>
objectSearchCallback(prefix, array)
)
);
return results;
},
/**
* search for full-text terms in the index
*/
performTermsSearch: (searchTerms, excludedTerms) => {
// prepare search
const terms = Search._index.terms;
const titleTerms = Search._index.titleterms;
const filenames = Search._index.filenames;
const docNames = Search._index.docnames;
const titles = Search._index.titles;
const scoreMap = new Map();
const fileMap = new Map();
// perform the search on the required terms
searchTerms.forEach((word) => {
const files = [];
const arr = [
{ files: terms[word], score: Scorer.term },
{ files: titleTerms[word], score: Scorer.title },
];
// add support for partial matches
if (word.length > 2) {
const escapedWord = _escapeRegExp(word);
if (!terms.hasOwnProperty(word)) {
Object.keys(terms).forEach((term) => {
if (term.match(escapedWord))
arr.push({ files: terms[term], score: Scorer.partialTerm });
});
}
if (!titleTerms.hasOwnProperty(word)) {
Object.keys(titleTerms).forEach((term) => {
if (term.match(escapedWord))
arr.push({ files: titleTerms[term], score: Scorer.partialTitle });
});
}
}
// no match but word was a required one
if (arr.every((record) => record.files === undefined)) return;
// found search word in contents
arr.forEach((record) => {
if (record.files === undefined) return;
let recordFiles = record.files;
if (recordFiles.length === undefined) recordFiles = [recordFiles];
files.push(...recordFiles);
// set score for the word in each file
recordFiles.forEach((file) => {
if (!scoreMap.has(file)) scoreMap.set(file, {});
scoreMap.get(file)[word] = record.score;
});
});
// create the mapping
files.forEach((file) => {
if (!fileMap.has(file)) fileMap.set(file, [word]);
else if (fileMap.get(file).indexOf(word) === -1) fileMap.get(file).push(word);
});
});
// now check if the files don't contain excluded terms
const results = [];
for (const [file, wordList] of fileMap) {
// check if all requirements are matched
// as search terms with length < 3 are discarded
const filteredTermCount = [...searchTerms].filter(
(term) => term.length > 2
).length;
if (
wordList.length !== searchTerms.size &&
wordList.length !== filteredTermCount
)
continue;
// ensure that none of the excluded terms is in the search result
if (
[...excludedTerms].some(
(term) =>
terms[term] === file ||
titleTerms[term] === file ||
(terms[term] || []).includes(file) ||
(titleTerms[term] || []).includes(file)
)
)
break;
// select one (max) score for the file.
const score = Math.max(...wordList.map((w) => scoreMap.get(file)[w]));
// add result to the result list
results.push([
docNames[file],
titles[file],
"",
null,
score,
filenames[file],
SearchResultKind.text,
]);
}
return results;
},
/**
* helper function to return a node containing the
* search summary for a given text. keywords is a list
* of stemmed words.
*/
makeSearchSummary: (htmlText, keywords, anchor) => {
const text = Search.htmlToText(htmlText, anchor);
if (text === "") return null;
const textLower = text.toLowerCase();
const actualStartPosition = [...keywords]
.map((k) => textLower.indexOf(k.toLowerCase()))
.filter((i) => i > -1)
.slice(-1)[0];
const startWithContext = Math.max(actualStartPosition - 120, 0);
const top = startWithContext === 0 ? "" : "...";
const tail = startWithContext + 240 < text.length ? "..." : "";
let summary = document.createElement("p");
summary.classList.add("context");
summary.textContent = top + text.substr(startWithContext, 240).trim() + tail;
return summary;
},
};
_ready(Search.init);
================================================
FILE: docs/_static/sphinx_highlight.js
================================================
/* Highlighting utilities for Sphinx HTML documentation. */
"use strict";
const SPHINX_HIGHLIGHT_ENABLED = true
/**
* highlight a given string on a node by wrapping it in
* span elements with the given class name.
*/
const _highlight = (node, addItems, text, className) => {
if (node.nodeType === Node.TEXT_NODE) {
const val = node.nodeValue;
const parent = node.parentNode;
const pos = val.toLowerCase().indexOf(text);
if (
pos >= 0 &&
!parent.classList.contains(className) &&
!parent.classList.contains("nohighlight")
) {
let span;
const closestNode = parent.closest("body, svg, foreignObject");
const isInSVG = closestNode && closestNode.matches("svg");
if (isInSVG) {
span = document.createElementNS("http://www.w3.org/2000/svg", "tspan");
} else {
span = document.createElement("span");
span.classList.add(className);
}
span.appendChild(document.createTextNode(val.substr(pos, text.length)));
const rest = document.createTextNode(val.substr(pos + text.length));
parent.insertBefore(
span,
parent.insertBefore(
rest,
node.nextSibling
)
);
node.nodeValue = val.substr(0, pos);
/* There may be more occurrences of search term in this node. So call this
* function recursively on the remaining fragment.
*/
_highlight(rest, addItems, text, className);
if (isInSVG) {
const rect = document.createElementNS(
"http://www.w3.org/2000/svg",
"rect"
);
const bbox = parent.getBBox();
rect.x.baseVal.value = bbox.x;
rect.y.baseVal.value = bbox.y;
rect.width.baseVal.value = bbox.width;
rect.height.baseVal.value = bbox.height;
rect.setAttribute("class", className);
addItems.push({ parent: parent, target: rect });
}
}
} else if (node.matches && !node.matches("button, select, textarea")) {
node.childNodes.forEach((el) => _highlight(el, addItems, text, className));
}
};
const _highlightText = (thisNode, text, className) => {
let addItems = [];
_highlight(thisNode, addItems, text, className);
addItems.forEach((obj) =>
obj.parent.insertAdjacentElement("beforebegin", obj.target)
);
};
/**
* Small JavaScript module for the documentation.
*/
const SphinxHighlight = {
/**
* highlight the search words provided in localstorage in the text
*/
highlightSearchWords: () => {
if (!SPHINX_HIGHLIGHT_ENABLED) return; // bail if no highlight
// get and clear terms from localstorage
const url = new URL(window.location);
const highlight =
localStorage.getItem("sphinx_highlight_terms")
|| url.searchParams.get("highlight")
|| "";
localStorage.removeItem("sphinx_highlight_terms")
url.searchParams.delete("highlight");
window.history.replaceState({}, "", url);
// get individual terms from highlight string
const terms = highlight.toLowerCase().split(/\s+/).filter(x => x);
if (terms.length === 0) return; // nothing to do
// There should never be more than one element matching "div.body"
const divBody = document.querySelectorAll("div.body");
const body = divBody.length ? divBody[0] : document.querySelector("body");
window.setTimeout(() => {
terms.forEach((term) => _highlightText(body, term, "highlighted"));
}, 10);
const searchBox = document.getElementById("searchbox");
if (searchBox === null) return;
searchBox.appendChild(
document
.createRange()
.createContextualFragment(
'<p class="highlight-link">' +
'<a href="javascript:SphinxHighlight.hideSearchWords()">' +
_("Hide Search Matches") +
"</a></p>"
)
);
},
/**
* helper function to hide the search marks again
*/
hideSearchWords: () => {
document
.querySelectorAll("#searchbox .highlight-link")
.forEach((el) => el.remove());
document
.querySelectorAll("span.highlighted")
.forEach((el) => el.classList.remove("highlighted"));
localStorage.removeItem("sphinx_highlight_terms")
},
initEscapeListener: () => {
// only install a listener if it is really needed
if (!DOCUMENTATION_OPTIONS.ENABLE_SEARCH_SHORTCUTS) return;
document.addEventListener("keydown", (event) => {
// bail for input elements
if (BLACKLISTED_KEY_CONTROL_ELEMENTS.has(document.activeElement.tagName)) return;
// bail with special keys
if (event.shiftKey || event.altKey || event.ctrlKey || event.metaKey) return;
if (DOCUMENTATION_OPTIONS.ENABLE_SEARCH_SHORTCUTS && (event.key === "Escape")) {
SphinxHighlight.hideSearchWords();
event.preventDefault();
}
});
},
};
_ready(() => {
/* Do not call highlightSearchWords() when we are on the search page.
* It will highlight words from the *previous* search query.
*/
if (typeof Search === "undefined") SphinxHighlight.highlightSearchWords();
SphinxHighlight.initEscapeListener();
});
================================================
FILE: docs/about.html
================================================
<!DOCTYPE html>
<html lang="en" data-content_root="./">
<head>
<meta charset="utf-8" />
<meta name="viewport" content="width=device-width, initial-scale=1.0" /><meta name="viewport" content="width=device-width, initial-scale=1" />
<title>About — pwlf 2.5.2 documentation</title>
<link rel="stylesheet" type="text/css" href="_static/pygments.css?v=d1102ebc" />
<link rel="stylesheet" type="text/css" href="_static/basic.css?v=686e5160" />
<link rel="stylesheet" type="text/css" href="_static/alabaster.css?v=27fed22d" />
<script src="_static/documentation_options.js?v=afc61bbc"></script>
<script src="_static/doctools.js?v=9bcbadda"></script>
<script src="_static/sphinx_highlight.js?v=dc90522c"></script>
<link rel="icon" href="_static/favicon.ico"/>
<link rel="author" title="About these documents" href="#" />
<link rel="index" title="Index" href="genindex.html" />
<link rel="search" title="Search" href="search.html" />
<link rel="next" title="Requirements" href="requirements.html" />
<link rel="prev" title="pwlf.PiecewiseLinFit" href="stubs/pwlf.PiecewiseLinFit.html" />
<link rel="stylesheet" href="_static/custom.css" type="text/css" />
</head><body>
<div class="document">
<div class="documentwrapper">
<div class="bodywrapper">
<div class="body" role="main">
<section id="about">
<h1>About<a class="headerlink" href="#about" title="Link to this heading">¶</a></h1>
<p>A library for fitting continuous piecewise linear functions to data. Just specify the number of line segments you desire and provide the data.</p>
<figure class="align-default">
<img alt="Example of a continuous piecewise linear fit to data." src="https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/examplePiecewiseFit.png" />
</figure>
<p>Example of a continuous piecewise linear fit to data.</p>
<p>All changes now stored in
<a class="reference external" href="https://github.com/cjekel/piecewise_linear_fit_py/blob/master/CHANGELOG.md">CHANGELOG.md</a></p>
<p>Please cite pwlf in your publications if it helps your research.</p>
<div class="highlight-bibtex notranslate"><div class="highlight"><pre><span></span><span class="nc">@Manual</span><span class="p">{</span><span class="nl">pwlf</span><span class="p">,</span>
<span class="w"> </span><span class="na">author</span><span class="w"> </span><span class="p">=</span><span class="w"> </span><span class="s">{Jekel, Charles F. and Venter, Gerhard}</span><span class="p">,</span>
<span class="w"> </span><span class="na">title</span><span class="w"> </span><span class="p">=</span><span class="w"> </span><span class="s">{{pwlf:} A Python Library for Fitting 1D Continuous Piecewise Linear Functions}</span><span class="p">,</span>
<span class="w"> </span><span class="na">year</span><span class="w"> </span><span class="p">=</span><span class="w"> </span><span class="s">{2019}</span><span class="p">,</span>
<span class="w"> </span><span class="na">url</span><span class="w"> </span><span class="p">=</span><span class="w"> </span><span class="s">{https://github.com/cjekel/piecewise_linear_fit_py}</span>
<span class="p">}</span>
</pre></div>
</div>
</section>
</div>
</div>
</div>
<div class="sphinxsidebar" role="navigation" aria-label="Main">
<div class="sphinxsidebarwrapper">
<h1 class="logo"><a href="index.html">pwlf</a></h1>
<search id="searchbox" style="display: none" role="search">
<div class="searchformwrapper">
<form class="search" action="search.html" method="get">
<input type="text" name="q" aria-labelledby="searchlabel" autocomplete="off" autocorrect="off" autocapitalize="off" spellcheck="false" placeholder="Search"/>
<input type="submit" value="Go" />
</form>
</div>
</search>
<script>document.getElementById('searchbox').style.display = "block"</script><h3>Navigation</h3>
<ul class="current">
<li class="toctree-l1"><a class="reference internal" href="installation.html">Installation</a></li>
<li class="toctree-l1"><a class="reference internal" href="how_it_works.html">How it works</a></li>
<li class="toctree-l1"><a class="reference internal" href="examples.html">Examples</a></li>
<li class="toctree-l1"><a class="reference internal" href="pwlf.html">pwlf package contents</a></li>
<li class="toctree-l1 current"><a class="current reference internal" href="#">About</a></li>
<li class="toctree-l1"><a class="reference internal" href="requirements.html">Requirements</a></li>
<li class="toctree-l1"><a class="reference internal" href="license.html">License</a></li>
</ul>
<div class="relations">
<h3>Related Topics</h3>
<ul>
<li><a href="index.html">Documentation overview</a><ul>
<li>Previous: <a href="stubs/pwlf.PiecewiseLinFit.html" title="previous chapter">pwlf.PiecewiseLinFit</a></li>
<li>Next: <a href="requirements.html" title="next chapter">Requirements</a></li>
</ul></li>
</ul>
</div>
</div>
</div>
<div class="clearer"></div>
</div>
<div class="footer">
©2024, Charles Jekel.
|
Powered by <a href="https://www.sphinx-doc.org/">Sphinx 8.1.3</a>
& <a href="https://alabaster.readthedocs.io">Alabaster 1.0.0</a>
|
<a href="_sources/about.rst.txt"
rel="nofollow">Page source</a>
</div>
<script>
var _gaq = _gaq || [];
_gaq.push(['_setAccount', 'G-QKPGZSZ8CD']);
_gaq.push(['_setDomainName', 'none']);
_gaq.push(['_setAllowLinker', true]);
_gaq.push(['_trackPageview']);
(function() {
var ga = document.createElement('script'); ga.async = true;
ga.src = ('https:' == document.location.protocol ? 'https://ssl' : 'https://www') + '.google-analytics.com/ga.js';
var s = document.getElementsByTagName('script')[0]; s.parentNode.insertBefore(ga, s);
})();
</script>
</body>
</html>
================================================
FILE: docs/examples.html
================================================
<!DOCTYPE html>
<html lang="en" data-content_root="./">
<head>
<meta charset="utf-8" />
<meta name="viewport" content="width=device-width, initial-scale=1.0" /><meta name="viewport" content="width=device-width, initial-scale=1" />
<title>Examples — pwlf 2.5.2 documentation</title>
<link rel="stylesheet" type="text/css" href="_static/pygments.css?v=d1102ebc" />
<link rel="stylesheet" type="text/css" href="_static/basic.css?v=686e5160" />
<link rel="stylesheet" type="text/css" href="_static/alabaster.css?v=27fed22d" />
<script src="_static/documentation_options.js?v=afc61bbc"></script>
<script src="_static/doctools.js?v=9bcbadda"></script>
<script src="_static/sphinx_highlight.js?v=dc90522c"></script>
<link rel="icon" href="_static/favicon.ico"/>
<link rel="author" title="About these documents" href="about.html" />
<link rel="index" title="Index" href="genindex.html" />
<link rel="search" title="Search" href="search.html" />
<link rel="next" title="pwlf package contents" href="pwlf.html" />
<link rel="prev" title="How it works" href="how_it_works.html" />
<link rel="stylesheet" href="_static/custom.css" type="text/css" />
</head><body>
<div class="document">
<div class="documentwrapper">
<div class="bodywrapper">
<div class="body" role="main">
<section id="examples">
<h1>Examples<a class="headerlink" href="#examples" title="Link to this heading">¶</a></h1>
<p>All of these examples will use the following data and imports.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="kn">import</span> <span class="nn">numpy</span> <span class="k">as</span> <span class="nn">np</span>
<span class="kn">import</span> <span class="nn">matplotlib.pyplot</span> <span class="k">as</span> <span class="nn">plt</span>
<span class="kn">import</span> <span class="nn">pwlf</span>
<span class="c1"># your data</span>
<span class="n">y</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">array</span><span class="p">([</span><span class="mf">0.00000000e+00</span><span class="p">,</span> <span class="mf">9.69801700e-03</span><span class="p">,</span> <span class="mf">2.94350340e-02</span><span class="p">,</span>
<span class="mf">4.39052750e-02</span><span class="p">,</span> <span class="mf">5.45343950e-02</span><span class="p">,</span> <span class="mf">6.74104940e-02</span><span class="p">,</span>
<span class="mf">8.34831790e-02</span><span class="p">,</span> <span class="mf">1.02580042e-01</span><span class="p">,</span> <span class="mf">1.22767939e-01</span><span class="p">,</span>
<span class="mf">1.42172312e-01</span><span class="p">,</span> <span class="mf">0.00000000e+00</span><span class="p">,</span> <span class="mf">8.58600000e-06</span><span class="p">,</span>
<span class="mf">8.31543400e-03</span><span class="p">,</span> <span class="mf">2.34184100e-02</span><span class="p">,</span> <span class="mf">3.39709150e-02</span><span class="p">,</span>
<span class="mf">4.03581990e-02</span><span class="p">,</span> <span class="mf">4.53545600e-02</span><span class="p">,</span> <span class="mf">5.02345260e-02</span><span class="p">,</span>
<span class="mf">5.55253360e-02</span><span class="p">,</span> <span class="mf">6.14750770e-02</span><span class="p">,</span> <span class="mf">6.82125120e-02</span><span class="p">,</span>
<span class="mf">7.55892510e-02</span><span class="p">,</span> <span class="mf">8.38356810e-02</span><span class="p">,</span> <span class="mf">9.26413070e-02</span><span class="p">,</span>
<span class="mf">1.02039790e-01</span><span class="p">,</span> <span class="mf">1.11688258e-01</span><span class="p">,</span> <span class="mf">1.21390666e-01</span><span class="p">,</span>
<span class="mf">1.31196948e-01</span><span class="p">,</span> <span class="mf">0.00000000e+00</span><span class="p">,</span> <span class="mf">1.56706510e-02</span><span class="p">,</span>
<span class="mf">3.54628780e-02</span><span class="p">,</span> <span class="mf">4.63739040e-02</span><span class="p">,</span> <span class="mf">5.61442590e-02</span><span class="p">,</span>
<span class="mf">6.78542550e-02</span><span class="p">,</span> <span class="mf">8.16388310e-02</span><span class="p">,</span> <span class="mf">9.77756110e-02</span><span class="p">,</span>
<span class="mf">1.16531753e-01</span><span class="p">,</span> <span class="mf">1.37038283e-01</span><span class="p">,</span> <span class="mf">0.00000000e+00</span><span class="p">,</span>
<span class="mf">1.16951050e-02</span><span class="p">,</span> <span class="mf">3.12089850e-02</span><span class="p">,</span> <span class="mf">4.41776550e-02</span><span class="p">,</span>
<span class="mf">5.42877590e-02</span><span class="p">,</span> <span class="mf">6.63321350e-02</span><span class="p">,</span> <span class="mf">8.07655920e-02</span><span class="p">,</span>
<span class="mf">9.70363280e-02</span><span class="p">,</span> <span class="mf">1.15706975e-01</span><span class="p">,</span> <span class="mf">1.36687642e-01</span><span class="p">,</span>
<span class="mf">0.00000000e+00</span><span class="p">,</span> <span class="mf">1.50144640e-02</span><span class="p">,</span> <span class="mf">3.44519970e-02</span><span class="p">,</span>
<span class="mf">4.55907760e-02</span><span class="p">,</span> <span class="mf">5.59556700e-02</span><span class="p">,</span> <span class="mf">6.88450940e-02</span><span class="p">,</span>
<span class="mf">8.41374060e-02</span><span class="p">,</span> <span class="mf">1.01254006e-01</span><span class="p">,</span> <span class="mf">1.20605073e-01</span><span class="p">,</span>
<span class="mf">1.41881288e-01</span><span class="p">,</span> <span class="mf">1.62618058e-01</span><span class="p">])</span>
<span class="n">x</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">array</span><span class="p">([</span><span class="mf">0.00000000e+00</span><span class="p">,</span> <span class="mf">8.82678000e-03</span><span class="p">,</span> <span class="mf">3.25615100e-02</span><span class="p">,</span>
<span class="mf">5.66106800e-02</span><span class="p">,</span> <span class="mf">7.95549800e-02</span><span class="p">,</span> <span class="mf">1.00936330e-01</span><span class="p">,</span>
<span class="mf">1.20351520e-01</span><span class="p">,</span> <span class="mf">1.37442010e-01</span><span class="p">,</span> <span class="mf">1.51858250e-01</span><span class="p">,</span>
<span class="mf">1.64433570e-01</span><span class="p">,</span> <span class="mf">0.00000000e+00</span><span class="p">,</span> <span class="o">-</span><span class="mf">2.12600000e-05</span><span class="p">,</span>
<span class="mf">7.03872000e-03</span><span class="p">,</span> <span class="mf">1.85494500e-02</span><span class="p">,</span> <span class="mf">3.00926700e-02</span><span class="p">,</span>
<span class="mf">4.17617000e-02</span><span class="p">,</span> <span class="mf">5.37279600e-02</span><span class="p">,</span> <span class="mf">6.54941000e-02</span><span class="p">,</span>
<span class="mf">7.68092100e-02</span><span class="p">,</span> <span class="mf">8.76596300e-02</span><span class="p">,</span> <span class="mf">9.80525800e-02</span><span class="p">,</span>
<span class="mf">1.07961810e-01</span><span class="p">,</span> <span class="mf">1.17305210e-01</span><span class="p">,</span> <span class="mf">1.26063930e-01</span><span class="p">,</span>
<span class="mf">1.34180360e-01</span><span class="p">,</span> <span class="mf">1.41725010e-01</span><span class="p">,</span> <span class="mf">1.48629710e-01</span><span class="p">,</span>
<span class="mf">1.55374770e-01</span><span class="p">,</span> <span class="mf">0.00000000e+00</span><span class="p">,</span> <span class="mf">1.65610200e-02</span><span class="p">,</span>
<span class="mf">3.91016100e-02</span><span class="p">,</span> <span class="mf">6.18679400e-02</span><span class="p">,</span> <span class="mf">8.30997400e-02</span><span class="p">,</span>
<span class="mf">1.02132890e-01</span><span class="p">,</span> <span class="mf">1.19011260e-01</span><span class="p">,</span> <span class="mf">1.34620080e-01</span><span class="p">,</span>
<span class="mf">1.49429370e-01</span><span class="p">,</span> <span class="mf">1.63539960e-01</span><span class="p">,</span> <span class="o">-</span><span class="mf">0.00000000e+00</span><span class="p">,</span>
<span class="mf">1.01980300e-02</span><span class="p">,</span> <span class="mf">3.28642800e-02</span><span class="p">,</span> <span class="mf">5.59461900e-02</span><span class="p">,</span>
<span class="mf">7.81388400e-02</span><span class="p">,</span> <span class="mf">9.84458400e-02</span><span class="p">,</span> <span class="mf">1.16270210e-01</span><span class="p">,</span>
<span class="mf">1.31279040e-01</span><span class="p">,</span> <span class="mf">1.45437090e-01</span><span class="p">,</span> <span class="mf">1.59627540e-01</span><span class="p">,</span>
<span class="mf">0.00000000e+00</span><span class="p">,</span> <span class="mf">1.63404300e-02</span><span class="p">,</span> <span class="mf">4.00086000e-02</span><span class="p">,</span>
<span class="mf">6.34390200e-02</span><span class="p">,</span> <span class="mf">8.51085900e-02</span><span class="p">,</span> <span class="mf">1.04787860e-01</span><span class="p">,</span>
<span class="mf">1.22120350e-01</span><span class="p">,</span> <span class="mf">1.36931660e-01</span><span class="p">,</span> <span class="mf">1.50958760e-01</span><span class="p">,</span>
<span class="mf">1.65299640e-01</span><span class="p">,</span> <span class="mf">1.79942720e-01</span><span class="p">])</span>
</pre></div>
</div>
<ol class="arabic simple">
<li><p><a class="reference external" href="#fit-with-known-breakpoint-locations">fit with known breakpoint
locations</a></p></li>
<li><p><a class="reference external" href="#fit-for-specified-number-of-line-segments">fit for specified number of line
segments</a></p></li>
<li><p><a class="reference external" href="#fitfast-for-specified-number-of-line-segments">fitfast for specified number of line
segments</a></p></li>
<li><p><a class="reference external" href="#force-a-fit-through-data-points">force a fit through data
points</a></p></li>
<li><p><a class="reference external" href="#use-custom-optimization-routine">use custom optimization
routine</a></p></li>
<li><p><a class="reference external" href="#pass-differential-evolution-keywords">pass differential evolution
keywords</a></p></li>
<li><p><a class="reference external" href="#find-the-best-number-of-line-segments">find the best number of line
segments</a></p></li>
<li><p><a class="reference external" href="#model-persistence">model persistence</a></p></li>
<li><p><a class="reference external" href="#bad-fits-when-you-have-more-unknowns-than-data">bad fits when you have more unknowns than
data</a></p></li>
<li><p><a class="reference external" href="#fit-with-a-breakpoint-guess">fit with a breakpoint guess</a></p></li>
<li><p><a class="reference external" href="#get-the-linear-regression-matrix">get the linear regression
matrix</a></p></li>
<li><p><a class="reference external" href="#use-of-tensorflow">use of TensorFlow</a></p></li>
<li><p><a class="reference external" href="#fit-constants-or-polynomials">fit constants or polynomials</a></p></li>
<li><p><a class="reference external" href="#specify-breakpoint-bounds">specify breakpoint bounds</a></p></li>
<li><p><a class="reference external" href="#non-linear-standard-errors-and-p-values">non-linear standard errors and
p-values</a></p></li>
<li><p><a class="reference external" href="#obtain-the-equations-of-fitted-pwlf">obtain the equations of fitted
pwlf</a></p></li>
<li><p><a class="reference external" href="#weighted-least-squares-fit">weighted least squares fit</a></p></li>
<li><p><a class="reference external" href="#reproducibleresults">reproducible results</a></p></li>
</ol>
<section id="fit-with-known-breakpoint-locations">
<h2>fit with known breakpoint locations<a class="headerlink" href="#fit-with-known-breakpoint-locations" title="Link to this heading">¶</a></h2>
<p>You can perform a least squares fit if you know the breakpoint
locations.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="c1"># your desired line segment end locations</span>
<span class="n">x0</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">array</span><span class="p">([</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="mf">0.039</span><span class="p">,</span> <span class="mf">0.10</span><span class="p">,</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">)])</span>
<span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># fit the data with the specified break points</span>
<span class="c1"># (ie the x locations of where the line segments</span>
<span class="c1"># will terminate)</span>
<span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit_with_breaks</span><span class="p">(</span><span class="n">x0</span><span class="p">)</span>
<span class="c1"># predict for the determined points</span>
<span class="n">xHat</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="n">num</span><span class="o">=</span><span class="mi">10000</span><span class="p">)</span>
<span class="n">yHat</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">predict</span><span class="p">(</span><span class="n">xHat</span><span class="p">)</span>
<span class="c1"># plot the results</span>
<span class="n">plt</span><span class="o">.</span><span class="n">figure</span><span class="p">()</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">,</span> <span class="s1">'o'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">xHat</span><span class="p">,</span> <span class="n">yHat</span><span class="p">,</span> <span class="s1">'-'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">show</span><span class="p">()</span>
</pre></div>
</div>
<figure class="align-default" id="id1">
<img alt="fit with known breakpoint locations" src="https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/fit_breaks.png" />
<figcaption>
<p><span class="caption-text">fit with known breakpoint locations</span><a class="headerlink" href="#id1" title="Link to this image">¶</a></p>
</figcaption>
</figure>
</section>
<section id="fit-for-specified-number-of-line-segments">
<h2>fit for specified number of line segments<a class="headerlink" href="#fit-for-specified-number-of-line-segments" title="Link to this heading">¶</a></h2>
<p>Use a global optimization to find the breakpoint locations that minimize
the sum of squares error. This uses <a class="reference external" href="https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.differential_evolution.html">Differential
Evolution</a>
from scipy.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># fit the data for four line segments</span>
<span class="n">res</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit</span><span class="p">(</span><span class="mi">4</span><span class="p">)</span>
<span class="c1"># predict for the determined points</span>
<span class="n">xHat</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="n">num</span><span class="o">=</span><span class="mi">10000</span><span class="p">)</span>
<span class="n">yHat</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">predict</span><span class="p">(</span><span class="n">xHat</span><span class="p">)</span>
<span class="c1"># plot the results</span>
<span class="n">plt</span><span class="o">.</span><span class="n">figure</span><span class="p">()</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">,</span> <span class="s1">'o'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">xHat</span><span class="p">,</span> <span class="n">yHat</span><span class="p">,</span> <span class="s1">'-'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">show</span><span class="p">()</span>
</pre></div>
</div>
<figure class="align-default" id="id2">
<img alt="fit for specified number of line segments" src="https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/numberoflines.png" />
<figcaption>
<p><span class="caption-text">fit for specified number of line segments</span><a class="headerlink" href="#id2" title="Link to this image">¶</a></p>
</figcaption>
</figure>
</section>
<section id="fitfast-for-specified-number-of-line-segments">
<h2>fitfast for specified number of line segments<a class="headerlink" href="#fitfast-for-specified-number-of-line-segments" title="Link to this heading">¶</a></h2>
<p>This performs a fit for a specified number of line segments with a
multi-start gradient based optimization. This should be faster than
<a class="reference external" href="https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.differential_evolution.html">Differential
Evolution</a>
for a small number of starting points.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># fit the data for four line segments</span>
<span class="c1"># this performs 3 multi-start optimizations</span>
<span class="n">res</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">fitfast</span><span class="p">(</span><span class="mi">4</span><span class="p">,</span> <span class="n">pop</span><span class="o">=</span><span class="mi">3</span><span class="p">)</span>
<span class="c1"># predict for the determined points</span>
<span class="n">xHat</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="n">num</span><span class="o">=</span><span class="mi">10000</span><span class="p">)</span>
<span class="n">yHat</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">predict</span><span class="p">(</span><span class="n">xHat</span><span class="p">)</span>
<span class="c1"># plot the results</span>
<span class="n">plt</span><span class="o">.</span><span class="n">figure</span><span class="p">()</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">,</span> <span class="s1">'o'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">xHat</span><span class="p">,</span> <span class="n">yHat</span><span class="p">,</span> <span class="s1">'-'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">show</span><span class="p">()</span>
</pre></div>
</div>
<figure class="align-default" id="id3">
<img alt="fitfast for specified number of line segments" src="https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/fitfast.png" />
<figcaption>
<p><span class="caption-text">fitfast for specified number of line segments</span><a class="headerlink" href="#id3" title="Link to this image">¶</a></p>
</figcaption>
</figure>
</section>
<section id="force-a-fit-through-data-points">
<h2>force a fit through data points<a class="headerlink" href="#force-a-fit-through-data-points" title="Link to this heading">¶</a></h2>
<p>Sometimes it’s necessary to force the piecewise continuous model through
a particular data point, or a set of data points. The following example
finds the best 4 line segments that go through two data points.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">myPWLF</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># fit the function with four line segments</span>
<span class="c1"># force the function to go through the data points</span>
<span class="c1"># (0.0, 0.0) and (0.19, 0.16)</span>
<span class="c1"># where the data points are of the form (x, y)</span>
<span class="n">x_c</span> <span class="o">=</span> <span class="p">[</span><span class="mf">0.0</span><span class="p">,</span> <span class="mf">0.19</span><span class="p">]</span>
<span class="n">y_c</span> <span class="o">=</span> <span class="p">[</span><span class="mf">0.0</span><span class="p">,</span> <span class="mf">0.2</span><span class="p">]</span>
<span class="n">res</span> <span class="o">=</span> <span class="n">myPWLF</span><span class="o">.</span><span class="n">fit</span><span class="p">(</span><span class="mi">4</span><span class="p">,</span> <span class="n">x_c</span><span class="p">,</span> <span class="n">y_c</span><span class="p">)</span>
<span class="c1"># predict for the determined points</span>
<span class="n">xHat</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="mf">0.19</span><span class="p">,</span> <span class="n">num</span><span class="o">=</span><span class="mi">10000</span><span class="p">)</span>
<span class="n">yHat</span> <span class="o">=</span> <span class="n">myPWLF</span><span class="o">.</span><span class="n">predict</span><span class="p">(</span><span class="n">xHat</span><span class="p">)</span>
<span class="c1"># plot the results</span>
<span class="n">plt</span><span class="o">.</span><span class="n">figure</span><span class="p">()</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">,</span> <span class="s1">'o'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">xHat</span><span class="p">,</span> <span class="n">yHat</span><span class="p">,</span> <span class="s1">'-'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">show</span><span class="p">()</span>
</pre></div>
</div>
<figure class="align-default" id="id4">
<img alt="force a fit through data points" src="https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/force.png" />
<figcaption>
<p><span class="caption-text">force a fit through data points</span><a class="headerlink" href="#id4" title="Link to this image">¶</a></p>
</figcaption>
</figure>
</section>
<section id="use-custom-optimization-routine">
<h2>use custom optimization routine<a class="headerlink" href="#use-custom-optimization-routine" title="Link to this heading">¶</a></h2>
<p>You can use your favorite optimization routine to find the breakpoint
locations. The following example uses scipy’s
<a class="reference external" href="https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html">minimize</a>
function.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="kn">from</span> <span class="nn">scipy.optimize</span> <span class="kn">import</span> <span class="n">minimize</span>
<span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># initialize custom optimization</span>
<span class="n">number_of_line_segments</span> <span class="o">=</span> <span class="mi">3</span>
<span class="n">my_pwlf</span><span class="o">.</span><span class="n">use_custom_opt</span><span class="p">(</span><span class="n">number_of_line_segments</span><span class="p">)</span>
<span class="c1"># i have number_of_line_segments - 1 number of variables</span>
<span class="c1"># let's guess the correct location of the two unknown variables</span>
<span class="c1"># (the program defaults to have end segments at x0= min(x)</span>
<span class="c1"># and xn=max(x)</span>
<span class="n">xGuess</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">zeros</span><span class="p">(</span><span class="n">number_of_line_segments</span> <span class="o">-</span> <span class="mi">1</span><span class="p">)</span>
<span class="n">xGuess</span><span class="p">[</span><span class="mi">0</span><span class="p">]</span> <span class="o">=</span> <span class="mf">0.02</span>
<span class="n">xGuess</span><span class="p">[</span><span class="mi">1</span><span class="p">]</span> <span class="o">=</span> <span class="mf">0.10</span>
<span class="n">res</span> <span class="o">=</span> <span class="n">minimize</span><span class="p">(</span><span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit_with_breaks_opt</span><span class="p">,</span> <span class="n">xGuess</span><span class="p">)</span>
<span class="c1"># set up the break point locations</span>
<span class="n">x0</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">zeros</span><span class="p">(</span><span class="n">number_of_line_segments</span> <span class="o">+</span> <span class="mi">1</span><span class="p">)</span>
<span class="n">x0</span><span class="p">[</span><span class="mi">0</span><span class="p">]</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">min</span><span class="p">(</span><span class="n">x</span><span class="p">)</span>
<span class="n">x0</span><span class="p">[</span><span class="o">-</span><span class="mi">1</span><span class="p">]</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">max</span><span class="p">(</span><span class="n">x</span><span class="p">)</span>
<span class="n">x0</span><span class="p">[</span><span class="mi">1</span><span class="p">:</span><span class="o">-</span><span class="mi">1</span><span class="p">]</span> <span class="o">=</span> <span class="n">res</span><span class="o">.</span><span class="n">x</span>
<span class="c1"># calculate the parameters based on the optimal break point locations</span>
<span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit_with_breaks</span><span class="p">(</span><span class="n">x0</span><span class="p">)</span>
<span class="c1"># predict for the determined points</span>
<span class="n">xHat</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="n">num</span><span class="o">=</span><span class="mi">10000</span><span class="p">)</span>
<span class="n">yHat</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">predict</span><span class="p">(</span><span class="n">xHat</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">figure</span><span class="p">()</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">,</span> <span class="s1">'o'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">xHat</span><span class="p">,</span> <span class="n">yHat</span><span class="p">,</span> <span class="s1">'-'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">show</span><span class="p">()</span>
</pre></div>
</div>
</section>
<section id="pass-differential-evolution-keywords">
<h2>pass differential evolution keywords<a class="headerlink" href="#pass-differential-evolution-keywords" title="Link to this heading">¶</a></h2>
<p>You can pass keyword arguments from the <code class="docutils literal notranslate"><span class="pre">fit</span></code> function into scipy’s
<a class="reference external" href="https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.differential_evolution.html">Differential
Evolution</a>.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># fit the data for four line segments</span>
<span class="c1"># this sets DE to have an absolute tolerance of 0.1</span>
<span class="n">res</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit</span><span class="p">(</span><span class="mi">4</span><span class="p">,</span> <span class="n">atol</span><span class="o">=</span><span class="mf">0.1</span><span class="p">)</span>
<span class="c1"># predict for the determined points</span>
<span class="n">xHat</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="n">num</span><span class="o">=</span><span class="mi">10000</span><span class="p">)</span>
<span class="n">yHat</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">predict</span><span class="p">(</span><span class="n">xHat</span><span class="p">)</span>
<span class="c1"># plot the results</span>
<span class="n">plt</span><span class="o">.</span><span class="n">figure</span><span class="p">()</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">,</span> <span class="s1">'o'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">xHat</span><span class="p">,</span> <span class="n">yHat</span><span class="p">,</span> <span class="s1">'-'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">show</span><span class="p">()</span>
</pre></div>
</div>
</section>
<section id="find-the-best-number-of-line-segments">
<h2>find the best number of line segments<a class="headerlink" href="#find-the-best-number-of-line-segments" title="Link to this heading">¶</a></h2>
<p>This example uses EGO (bayesian optimization) and a penalty function to
find the best number of line segments. This will require careful use of
the penalty parameter <code class="docutils literal notranslate"><span class="pre">l</span></code>. Use this template to automatically find the
best number of line segments.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="kn">from</span> <span class="nn">GPyOpt.methods</span> <span class="kn">import</span> <span class="n">BayesianOptimization</span>
<span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># define your objective function</span>
<span class="k">def</span> <span class="nf">my_obj</span><span class="p">(</span><span class="n">x</span><span class="p">):</span>
<span class="c1"># define some penalty parameter l</span>
<span class="c1"># you'll have to arbitrarily pick this</span>
<span class="c1"># it depends upon the noise in your data,</span>
<span class="c1"># and the value of your sum of square of residuals</span>
<span class="n">l</span> <span class="o">=</span> <span class="n">y</span><span class="o">.</span><span class="n">mean</span><span class="p">()</span><span class="o">*</span><span class="mf">0.001</span>
<span class="n">f</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">zeros</span><span class="p">(</span><span class="n">x</span><span class="o">.</span><span class="n">shape</span><span class="p">[</span><span class="mi">0</span><span class="p">])</span>
<span class="k">for</span> <span class="n">i</span><span class="p">,</span> <span class="n">j</span> <span class="ow">in</span> <span class="nb">enumerate</span><span class="p">(</span><span class="n">x</span><span class="p">):</span>
<span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit</span><span class="p">(</span><span class="n">j</span><span class="p">[</span><span class="mi">0</span><span class="p">])</span>
<span class="n">f</span><span class="p">[</span><span class="n">i</span><span class="p">]</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">ssr</span> <span class="o">+</span> <span class="p">(</span><span class="n">l</span><span class="o">*</span><span class="n">j</span><span class="p">[</span><span class="mi">0</span><span class="p">])</span>
<span class="k">return</span> <span class="n">f</span>
<span class="c1"># define the lower and upper bound for the number of line segments</span>
<span class="n">bounds</span> <span class="o">=</span> <span class="p">[{</span><span class="s1">'name'</span><span class="p">:</span> <span class="s1">'var_1'</span><span class="p">,</span> <span class="s1">'type'</span><span class="p">:</span> <span class="s1">'discrete'</span><span class="p">,</span>
<span class="s1">'domain'</span><span class="p">:</span> <span class="n">np</span><span class="o">.</span><span class="n">arange</span><span class="p">(</span><span class="mi">2</span><span class="p">,</span> <span class="mi">40</span><span class="p">)}]</span>
<span class="n">np</span><span class="o">.</span><span class="n">random</span><span class="o">.</span><span class="n">seed</span><span class="p">(</span><span class="mi">12121</span><span class="p">)</span>
<span class="n">myBopt</span> <span class="o">=</span> <span class="n">BayesianOptimization</span><span class="p">(</span><span class="n">my_obj</span><span class="p">,</span> <span class="n">domain</span><span class="o">=</span><span class="n">bounds</span><span class="p">,</span> <span class="n">model_type</span><span class="o">=</span><span class="s1">'GP'</span><span class="p">,</span>
<span class="n">initial_design_numdata</span><span class="o">=</span><span class="mi">10</span><span class="p">,</span>
<span class="n">initial_design_type</span><span class="o">=</span><span class="s1">'latin'</span><span class="p">,</span>
<span class="n">exact_feval</span><span class="o">=</span><span class="kc">True</span><span class="p">,</span> <span class="n">verbosity</span><span class="o">=</span><span class="kc">True</span><span class="p">,</span>
<span class="n">verbosity_model</span><span class="o">=</span><span class="kc">False</span><span class="p">)</span>
<span class="n">max_iter</span> <span class="o">=</span> <span class="mi">30</span>
<span class="c1"># perform the bayesian optimization to find the optimum number</span>
<span class="c1"># of line segments</span>
<span class="n">myBopt</span><span class="o">.</span><span class="n">run_optimization</span><span class="p">(</span><span class="n">max_iter</span><span class="o">=</span><span class="n">max_iter</span><span class="p">,</span> <span class="n">verbosity</span><span class="o">=</span><span class="kc">True</span><span class="p">)</span>
<span class="nb">print</span><span class="p">(</span><span class="s1">'</span><span class="se">\n</span><span class="s1"> </span><span class="se">\n</span><span class="s1"> Opt found </span><span class="se">\n</span><span class="s1">'</span><span class="p">)</span>
<span class="nb">print</span><span class="p">(</span><span class="s1">'Optimum number of line segments:'</span><span class="p">,</span> <span class="n">myBopt</span><span class="o">.</span><span class="n">x_opt</span><span class="p">)</span>
<span class="nb">print</span><span class="p">(</span><span class="s1">'Function value:'</span><span class="p">,</span> <span class="n">myBopt</span><span class="o">.</span><span class="n">fx_opt</span><span class="p">)</span>
<span class="n">myBopt</span><span class="o">.</span><span class="n">plot_acquisition</span><span class="p">()</span>
<span class="n">myBopt</span><span class="o">.</span><span class="n">plot_convergence</span><span class="p">()</span>
<span class="c1"># perform the fit for the optimum</span>
<span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit</span><span class="p">(</span><span class="n">myBopt</span><span class="o">.</span><span class="n">x_opt</span><span class="p">)</span>
<span class="c1"># predict for the determined points</span>
<span class="n">xHat</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="n">num</span><span class="o">=</span><span class="mi">10000</span><span class="p">)</span>
<span class="n">yHat</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">predict</span><span class="p">(</span><span class="n">xHat</span><span class="p">)</span>
<span class="c1"># plot the results</span>
<span class="n">plt</span><span class="o">.</span><span class="n">figure</span><span class="p">()</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">,</span> <span class="s1">'o'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">xHat</span><span class="p">,</span> <span class="n">yHat</span><span class="p">,</span> <span class="s1">'-'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">show</span><span class="p">()</span>
</pre></div>
</div>
</section>
<section id="model-persistence">
<h2>model persistence<a class="headerlink" href="#model-persistence" title="Link to this heading">¶</a></h2>
<p>You can save fitted models with pickle. Alternatively see
<a class="reference external" href="https://joblib.readthedocs.io/en/latest/">joblib</a>.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="c1"># if you use Python 2.x you should import cPickle</span>
<span class="c1"># import cPickle as pickle</span>
<span class="c1"># if you use Python 3.x you can just use pickle</span>
<span class="kn">import</span> <span class="nn">pickle</span>
<span class="c1"># your desired line segment end locations</span>
<span class="n">x0</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">array</span><span class="p">([</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="mf">0.039</span><span class="p">,</span> <span class="mf">0.10</span><span class="p">,</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">)])</span>
<span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># fit the data with the specified break points</span>
<span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit_with_breaks</span><span class="p">(</span><span class="n">x0</span><span class="p">)</span>
<span class="c1"># save the fitted model</span>
<span class="k">with</span> <span class="nb">open</span><span class="p">(</span><span class="s1">'my_fit.pkl'</span><span class="p">,</span> <span class="s1">'wb'</span><span class="p">)</span> <span class="k">as</span> <span class="n">f</span><span class="p">:</span>
<span class="n">pickle</span><span class="o">.</span><span class="n">dump</span><span class="p">(</span><span class="n">my_pwlf</span><span class="p">,</span> <span class="n">f</span><span class="p">,</span> <span class="n">pickle</span><span class="o">.</span><span class="n">HIGHEST_PROTOCOL</span><span class="p">)</span>
<span class="c1"># load the fitted model</span>
<span class="k">with</span> <span class="nb">open</span><span class="p">(</span><span class="s1">'my_fit.pkl'</span><span class="p">,</span> <span class="s1">'rb'</span><span class="p">)</span> <span class="k">as</span> <span class="n">f</span><span class="p">:</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pickle</span><span class="o">.</span><span class="n">load</span><span class="p">(</span><span class="n">f</span><span class="p">)</span>
</pre></div>
</div>
</section>
<section id="bad-fits-when-you-have-more-unknowns-than-data">
<h2>bad fits when you have more unknowns than data<a class="headerlink" href="#bad-fits-when-you-have-more-unknowns-than-data" title="Link to this heading">¶</a></h2>
<p>You can get very bad fits with pwlf when you have more unknowns than
data points. The following example will fit 99 line segments to the 59
data points. While this will result in an error of zero, the model will
have very weird predictions within the data. You should not fit more
unknowns than you have data with pwlf!</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="n">break_locations</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="n">num</span><span class="o">=</span><span class="mi">100</span><span class="p">)</span>
<span class="c1"># initialize piecewise linear fit with your x and y data</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit_with_breaks</span><span class="p">(</span><span class="n">break_locations</span><span class="p">)</span>
<span class="c1"># predict for the determined points</span>
<span class="n">xHat</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="nb">min</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="nb">max</span><span class="p">(</span><span class="n">x</span><span class="p">),</span> <span class="n">num</span><span class="o">=</span><span class="mi">10000</span><span class="p">)</span>
<span class="n">yHat</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">predict</span><span class="p">(</span><span class="n">xHat</span><span class="p">)</span>
<span class="c1"># plot the results</span>
<span class="n">plt</span><span class="o">.</span><span class="n">figure</span><span class="p">()</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">,</span> <span class="s1">'o'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">plot</span><span class="p">(</span><span class="n">xHat</span><span class="p">,</span> <span class="n">yHat</span><span class="p">,</span> <span class="s1">'-'</span><span class="p">)</span>
<span class="n">plt</span><span class="o">.</span><span class="n">show</span><span class="p">()</span>
</pre></div>
</div>
<figure class="align-default" id="id5">
<img alt="bad fits when you have more unknowns than data" src="https://raw.githubusercontent.com/cjekel/piecewise_linear_fit_py/master/examples/figs/badfit.png" />
<figcaption>
<p><span class="caption-text">bad fits when you have more unknowns than data</span><a class="headerlink" href="#id5" title="Link to this image">¶</a></p>
</figcaption>
</figure>
</section>
<section id="fit-with-a-breakpoint-guess">
<h2>fit with a breakpoint guess<a class="headerlink" href="#fit-with-a-breakpoint-guess" title="Link to this heading">¶</a></h2>
<p>In this example we see two distinct linear regions, and we believe a
breakpoint occurs at 6.0. We’ll use the fit_guess() function to find the
best breakpoint location starting with this guess. These fits should be
much faster than the <code class="docutils literal notranslate"><span class="pre">fit</span></code> or <code class="docutils literal notranslate"><span class="pre">fitfast</span></code> function when you have a
reasonable idea where the breakpoints occur.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="kn">import</span> <span class="nn">numpy</span> <span class="k">as</span> <span class="nn">np</span>
<span class="kn">import</span> <span class="nn">pwlf</span>
<span class="n">x</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">array</span><span class="p">([</span><span class="mf">4.</span><span class="p">,</span> <span class="mf">5.</span><span class="p">,</span> <span class="mf">6.</span><span class="p">,</span> <span class="mf">7.</span><span class="p">,</span> <span class="mf">8.</span><span class="p">])</span>
<span class="n">y</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">array</span><span class="p">([</span><span class="mf">11.</span><span class="p">,</span> <span class="mf">13.</span><span class="p">,</span> <span class="mf">16.</span><span class="p">,</span> <span class="mf">28.92</span><span class="p">,</span> <span class="mf">42.81</span><span class="p">])</span>
<span class="n">my_pwlf</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="n">breaks</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit_guess</span><span class="p">([</span><span class="mf">6.0</span><span class="p">])</span>
</pre></div>
</div>
<p>Note specifying one breakpoint will result in two line segments. If we
wanted three line segments, we’ll have to specify two breakpoints.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="n">breaks</span> <span class="o">=</span> <span class="n">my_pwlf</span><span class="o">.</span><span class="n">fit_guess</span><span class="p">([</span><span class="mf">5.5</span><span class="p">,</span> <span class="mf">6.0</span><span class="p">])</span>
</pre></div>
</div>
</section>
<section id="get-the-linear-regression-matrix">
<h2>get the linear regression matrix<a class="headerlink" href="#get-the-linear-regression-matrix" title="Link to this heading">¶</a></h2>
<p>In some cases it may be desirable to work with the linear regression
matrix directly. The following example grabs the linear regression
matrix <code class="docutils literal notranslate"><span class="pre">A</span></code> for a specific set of breakpoints. In this case we assume
that the breakpoints occur at each of the data points. Please see the
<a class="reference external" href="https://github.com/cjekel/piecewise_linear_fit_py/tree/master/paper">paper</a>
for details about the regression matrix <code class="docutils literal notranslate"><span class="pre">A</span></code>.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="kn">import</span> <span class="nn">numpy</span> <span class="k">as</span> <span class="nn">np</span>
<span class="kn">import</span> <span class="nn">pwlf</span>
<span class="c1"># select random seed for reproducibility</span>
<span class="n">np</span><span class="o">.</span><span class="n">random</span><span class="o">.</span><span class="n">seed</span><span class="p">(</span><span class="mi">123</span><span class="p">)</span>
<span class="c1"># generate sin wave data</span>
<span class="n">x</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">linspace</span><span class="p">(</span><span class="mi">0</span><span class="p">,</span> <span class="mi">10</span><span class="p">,</span> <span class="n">num</span><span class="o">=</span><span class="mi">100</span><span class="p">)</span>
<span class="n">y</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">sin</span><span class="p">(</span><span class="n">x</span> <span class="o">*</span> <span class="n">np</span><span class="o">.</span><span class="n">pi</span> <span class="o">/</span> <span class="mi">2</span><span class="p">)</span>
<span class="n">ytrue</span> <span class="o">=</span> <span class="n">y</span><span class="o">.</span><span class="n">copy</span><span class="p">()</span>
<span class="c1"># add noise to the data</span>
<span class="n">y</span> <span class="o">=</span> <span class="n">np</span><span class="o">.</span><span class="n">random</span><span class="o">.</span><span class="n">normal</span><span class="p">(</span><span class="mi">0</span><span class="p">,</span> <span class="mf">0.05</span><span class="p">,</span> <span class="mi">100</span><span class="p">)</span> <span class="o">+</span> <span class="n">ytrue</span>
<span class="n">my_pwlf_en</span> <span class="o">=</span> <span class="n">pwlf</span><span class="o">.</span><span class="n">PiecewiseLinFit</span><span class="p">(</span><span class="n">x</span><span class="p">,</span> <span class="n">y</span><span class="p">)</span>
<span class="c1"># copy the x data to use as break points</span>
<span class="n">breaks</span> <span class="o">=</span> <span class="n">my_pwlf_en</span><span class="o">.</span><span class="n">x_data</span><span class="o">.</span><span class="n">copy</span><span class="p">()</span>
<span class="c1"># create the linear regression matrix A</span>
<span class="n">A</span> <span class="o">=</span> <span class="n">my_pwlf_en</span><span class="o">.</span><span class="n">assemble_regression_matrix</span><span class="p">(</span><span class="n">breaks</span><span class="p">,</span> <span class="n">my_pwlf_en</span><span class="o">.</span><span class="n">x_data</span><span class="p">)</span>
</pre></div>
</div>
<p>We can perform fits that are more complicated than a least squares fit
when we have the regression matrix. The following uses the Elastic Net
regularizer to perform an interesting fit with the regression matrix.</p>
<div class="highlight-python notranslate"><div class="highlight"><pre><span></span><span class="kn">from</span> <span class="nn">sklearn.linear_model</span> <span class="kn">import</span> <span class="n">ElasticNetCV</span>
<span class="c1"># set up the elastic net</span>
<span class="n">en_model</span> <span class="o">=</span> <span class="n">ElasticNetCV</span><span class="p">(</span><span class="n">cv</span><span class="o">=</span><span class="mi">5</span><span class="p">,</span>
<span class="n">l1_ratio</span><span class="o">=</span><span class="p">[</span><span class="mf">.1</span><span class="p">,</span> <span class="mf">.5</span><span class="p">,</span> <span class="mf">.7</span><span class="p">,</span> <span class="mf">.9</span><span class="p">,</span>
<span class="mf">.95</span><span class="p">,</span> <span class="mf">.99</span><span class="p">,</span> <span class="mi">1</span><span class="p">],</span>
<span class="n">fit_intercept</span><span class="o">=</span><span class="kc">False</span><span class="p">,</span>
<span class="n">max_iter</span><span class="o">=</span><span class="mi">1000000</span><span class="p">,</span> <span class="n">n_jobs</span><span class="o">=-</span><span class="mi">1</span><span class="p">)</span>
<span class="c1"># fit the model using the elastic net</span>
<span class="n">en_model</span><span class="o">.</span><span class="n">fit</span><span class="p">(</span><span class="n">A</span><span class="p">,</span> <span class="n">my_pwlf_en</span><span class="o">.</span><span class="n">y_data</span><span class="p">)</span>
<span class="c1"># predict from the elastic net parameters</span>
<span class="n">xhat</span> <span class="o">=</span> <sp
gitextract_642gn0fy/
├── .github/
│ └── workflows/
│ ├── ci.yml
│ └── cron.yml
├── .gitignore
├── CHANGELOG.md
├── LICENSE
├── MANIFEST.in
├── README.md
├── README.rst
├── convert_README_to_RST.sh
├── docs/
│ ├── .buildinfo
│ ├── .nojekyll
│ ├── _sources/
│ │ ├── about.rst.txt
│ │ ├── examples.rst.txt
│ │ ├── how_it_works.rst.txt
│ │ ├── index.rst.txt
│ │ ├── installation.rst.txt
│ │ ├── license.rst.txt
│ │ ├── modules.rst.txt
│ │ ├── pwlf.rst.txt
│ │ ├── requirements.rst.txt
│ │ └── stubs/
│ │ └── pwlf.PiecewiseLinFit.rst.txt
│ ├── _static/
│ │ ├── alabaster.css
│ │ ├── basic.css
│ │ ├── custom.css
│ │ ├── doctools.js
│ │ ├── documentation_options.js
│ │ ├── language_data.js
│ │ ├── pygments.css
│ │ ├── searchtools.js
│ │ └── sphinx_highlight.js
│ ├── about.html
│ ├── examples.html
│ ├── genindex.html
│ ├── how_it_works.html
│ ├── index.html
│ ├── installation.html
│ ├── license.html
│ ├── modules.html
│ ├── objects.inv
│ ├── pwlf.html
│ ├── requirements.html
│ ├── search.html
│ ├── searchindex.js
│ └── stubs/
│ └── pwlf.PiecewiseLinFit.html
├── doctrees/
│ ├── about.doctree
│ ├── environment.pickle
│ ├── examples.doctree
│ ├── how_it_works.doctree
│ ├── index.doctree
│ ├── installation.doctree
│ ├── license.doctree
│ ├── modules.doctree
│ ├── pwlf.doctree
│ ├── requirements.doctree
│ └── stubs/
│ └── pwlf.PiecewiseLinFit.doctree
├── examples/
│ ├── EGO_integer_only.ipynb
│ ├── README.md
│ ├── README.rst
│ ├── ex_data/
│ │ └── saved_parameters.npy
│ ├── experiment_with_batch_process.py.ipynb
│ ├── fitForSpecifiedNumberOfLineSegments.py
│ ├── fitForSpecifiedNumberOfLineSegments_passDiffEvoKeywords.py
│ ├── fitForSpecifiedNumberOfLineSegments_standard_deviation.py
│ ├── fitWithKnownLineSegmentLocations.py
│ ├── fit_begin_and_end.py
│ ├── min_length_demo.ipynb
│ ├── mixed_degree.py
│ ├── mixed_degree_forcing_slope.py
│ ├── model_persistence.py
│ ├── model_persistence_prediction.py
│ ├── prediction_variance.py
│ ├── prediction_variance_degree2.py
│ ├── robust_regression.py
│ ├── run_opt_to_find_best_number_of_line_segments.py
│ ├── sineWave.py
│ ├── sineWave_custom_opt_bounds.py
│ ├── sineWave_degrees.py
│ ├── sineWave_time_compare.py
│ ├── slope_constraint_demo.ipynb
│ ├── stack_overflow_example.py
│ ├── standard_errrors_and_p-values.py
│ ├── standard_errrors_and_p-values_non-linear.py
│ ├── test0.py
│ ├── test_for_model_significance.py
│ ├── test_if_breaks_exact.py
│ ├── tf/
│ │ ├── fit_begin_and_end.py
│ │ ├── sine_benchmark_fixed_20_break_points.py
│ │ ├── sine_benchmark_six_segments.py
│ │ └── test_fit.py
│ ├── understanding_higher_degrees/
│ │ └── polynomials_in_pwlf.ipynb
│ ├── useCustomOptimizationRoutine.py
│ ├── weighted_least_squares_ex.py
│ └── weighted_least_squares_ex_stats.py
├── pwlf/
│ ├── __init__.py
│ ├── pwlf.py
│ └── version.py
├── setup.py
├── sphinx_build_script.sh
├── sphinxdocs/
│ ├── Makefile
│ ├── make.bat
│ └── source/
│ ├── about.rst
│ ├── conf.py
│ ├── examples.rst
│ ├── how_it_works.rst
│ ├── index.rst
│ ├── installation.rst
│ ├── license.rst
│ ├── modules.rst
│ ├── pwlf.rst
│ └── requirements.rst
└── tests/
├── __init__.py
└── tests.py
SYMBOL INDEX (98 symbols across 9 files)
FILE: docs/_static/doctools.js
constant BLACKLISTED_KEY_CONTROL_ELEMENTS (line 6) | const BLACKLISTED_KEY_CONTROL_ELEMENTS = new Set([
FILE: docs/_static/documentation_options.js
constant DOCUMENTATION_OPTIONS (line 1) | const DOCUMENTATION_OPTIONS = {
FILE: docs/_static/searchtools.js
class SearchResultKind (line 44) | class SearchResultKind {
method index (line 45) | static get index() { return "index"; }
method object (line 46) | static get object() { return "object"; }
method text (line 47) | static get text() { return "text"; }
method title (line 48) | static get title() { return "title"; }
FILE: docs/_static/sphinx_highlight.js
constant SPHINX_HIGHLIGHT_ENABLED (line 4) | const SPHINX_HIGHLIGHT_ENABLED = true
FILE: examples/mixed_degree_forcing_slope.py
function my_fun (line 15) | def my_fun(beta):
FILE: examples/robust_regression.py
function my_fun (line 20) | def my_fun(breaks_and_beta):
FILE: examples/run_opt_to_find_best_number_of_line_segments.py
function my_obj (line 57) | def my_obj(x):
FILE: pwlf/pwlf.py
class PiecewiseLinFit (line 34) | class PiecewiseLinFit(object):
method __init__ (line 36) | def __init__(
method _switch_to_np_array (line 265) | def _switch_to_np_array(input_):
method _get_breaks (line 283) | def _get_breaks(self, input_):
method _fit_one_segment (line 302) | def _fit_one_segment(self):
method _fit_one_segment_force_points (line 308) | def _fit_one_segment_force_points(self, x_c, y_c):
method _check_mixed_degree_list (line 315) | def _check_mixed_degree_list(self, n_segments):
method assemble_regression_matrix (line 327) | def assemble_regression_matrix(self, breaks, x):
method fit_with_breaks (line 426) | def fit_with_breaks(self, breaks):
method fit_with_breaks_force_points (line 495) | def fit_with_breaks_force_points(self, breaks, x_c, y_c):
method predict (line 571) | def predict(self, x, beta=None, breaks=None):
method fit_with_breaks_opt (line 629) | def fit_with_breaks_opt(self, var):
method fit_force_points_opt (line 684) | def fit_force_points_opt(self, var):
method fit (line 733) | def fit(self, n_segments, x_c=None, y_c=None, bounds=None, **kwargs):
method fitfast (line 896) | def fitfast(self, n_segments, pop=2, bounds=None, **kwargs):
method fit_guess (line 1049) | def fit_guess(self, guess_breakpoints, bounds=None, **kwargs):
method use_custom_opt (line 1156) | def use_custom_opt(self, n_segments, x_c=None, y_c=None):
method calc_slopes (line 1220) | def calc_slopes(self):
method standard_errors (line 1258) | def standard_errors(self, method="linear", step_size=1e-4):
method prediction_variance (line 1391) | def prediction_variance(self, x):
method r_squared (line 1479) | def r_squared(self):
method p_values (line 1526) | def p_values(self, method="linear", step_size=1e-4):
method lstsq (line 1627) | def lstsq(self, A, calc_slopes=True):
method conlstsq (line 1679) | def conlstsq(self, A, calc_slopes=True):
FILE: tests/tests.py
class TestEverything (line 6) | class TestEverything(unittest.TestCase):
method test_break_point_spot_on (line 13) | def test_break_point_spot_on(self):
method test_break_point_spot_on_list (line 20) | def test_break_point_spot_on_list(self):
method test_degree_not_supported (line 27) | def test_degree_not_supported(self):
method test_x_c_not_supplied (line 35) | def test_x_c_not_supplied(self):
method test_y_c_not_supplied (line 44) | def test_y_c_not_supplied(self):
method test_lapack_driver (line 53) | def test_lapack_driver(self):
method test_assembly (line 61) | def test_assembly(self):
method test_assembly_list (line 72) | def test_assembly_list(self):
method test_break_point_spot_on_r2 (line 83) | def test_break_point_spot_on_r2(self):
method test_break_point_diff_x0_0 (line 91) | def test_break_point_diff_x0_0(self):
method test_break_point_diff_x0_1 (line 100) | def test_break_point_diff_x0_1(self):
method test_break_point_diff_x0_2 (line 108) | def test_break_point_diff_x0_2(self):
method test_break_point_diff_x0_3 (line 116) | def test_break_point_diff_x0_3(self):
method test_break_point_diff_x0_4 (line 124) | def test_break_point_diff_x0_4(self):
method test_diff_evo (line 132) | def test_diff_evo(self):
method test_custom_bounds1 (line 137) | def test_custom_bounds1(self):
method test_custom_bounds2 (line 148) | def test_custom_bounds2(self):
method test_predict (line 159) | def test_predict(self):
method test_custom_opt (line 168) | def test_custom_opt(self):
method test_custom_opt_with_con (line 176) | def test_custom_opt_with_con(self):
method test_single_force_break_point1 (line 184) | def test_single_force_break_point1(self):
method test_single_force_break_point2 (line 192) | def test_single_force_break_point2(self):
method test_single_force_break_point_degree (line 200) | def test_single_force_break_point_degree(self):
method test_single_force_break_point_degree_zero (line 208) | def test_single_force_break_point_degree_zero(self):
method test_opt_fit_with_force_points (line 216) | def test_opt_fit_with_force_points(self):
method test_opt_fit_single_segment (line 228) | def test_opt_fit_single_segment(self):
method test_opt_fit_with_force_points_single_segment (line 238) | def test_opt_fit_with_force_points_single_segment(self):
method test_se (line 250) | def test_se(self):
method test_p (line 257) | def test_p(self):
method test_nonlinear_p_and_se (line 264) | def test_nonlinear_p_and_se(self):
method test_pv (line 280) | def test_pv(self):
method test_predict_with_custom_param (line 288) | def test_predict_with_custom_param(self):
method test_fit_guess (line 295) | def test_fit_guess(self):
method test_fit_guess_kwrds (line 302) | def test_fit_guess_kwrds(self):
method test_multi_start_fitfast (line 313) | def test_multi_start_fitfast(self):
method test_n_parameters_correct (line 320) | def test_n_parameters_correct(self):
method test_force_fits_through_points_other_degrees (line 337) | def test_force_fits_through_points_other_degrees(self):
method test_fitfast (line 356) | def test_fitfast(self):
method test_fit (line 376) | def test_fit(self):
method test_se_no_fit (line 391) | def test_se_no_fit(self):
method test_se_no_method (line 399) | def test_se_no_method(self):
method test_pv_list (line 409) | def test_pv_list(self):
method test_pv_no_fit (line 415) | def test_pv_no_fit(self):
method test_r2_no_fit (line 423) | def test_r2_no_fit(self):
method test_pvalue_no_fit (line 431) | def test_pvalue_no_fit(self):
method test_pvalues_wrong_method (line 439) | def test_pvalues_wrong_method(self):
method test_all_stats (line 449) | def test_all_stats(self):
method test_weighted_same_as_ols (line 479) | def test_weighted_same_as_ols(self):
method test_heteroscedastic_data (line 494) | def test_heteroscedastic_data(self):
method test_not_supported_fit (line 504) | def test_not_supported_fit(self):
method test_not_supported_fit_with_breaks_force_points (line 517) | def test_not_supported_fit_with_breaks_force_points(self):
method test_custom_opt_not_supported (line 530) | def test_custom_opt_not_supported(self):
method test_random_seed_fit (line 539) | def test_random_seed_fit(self):
method test_random_seed_fitfast (line 551) | def test_random_seed_fitfast(self):
method test_one_segment_fits (line 564) | def test_one_segment_fits(self):
method test_float32 (line 573) | def test_float32(self):
method test_lfloat128 (line 581) | def test_lfloat128(self):
method test_mixed_degree1 (line 592) | def test_mixed_degree1(self):
method test_mixed_degree2 (line 604) | def test_mixed_degree2(self):
method test_mixed_degree3 (line 616) | def test_mixed_degree3(self):
method test_mixed_degree_wrong_list (line 628) | def test_mixed_degree_wrong_list(self):
method test_mixed_degree_force_point (line 649) | def test_mixed_degree_force_point(self):
Condensed preview — 112 files, each showing path, character count, and a content snippet. Download the .json file or copy for the full structured content (1,093K chars).
[
{
"path": ".github/workflows/ci.yml",
"chars": 1097,
"preview": "name: pwlf ci\n\non:\n push:\n\njobs:\n build:\n\n runs-on: ubuntu-22.04\n strategy:\n matrix:\n python-version"
},
{
"path": ".github/workflows/cron.yml",
"chars": 791,
"preview": "name: pwlf cron\n\non:\n schedule:\n # Also run every 24 hours\n - cron: '* */24 * * *'\n\njobs:\n build:\n\n runs-on:"
},
{
"path": ".gitignore",
"chars": 333,
"preview": "*.pyc\n.coverage\nMANIFEST\n/build/\nsetup.cfg\n# Setuptools distribution folder.\n/dist/\n\n# Python egg metadata, regenerated "
},
{
"path": "CHANGELOG.md",
"chars": 15169,
"preview": "# Changelog\nAll notable changes to this project will be documented in this file.\n\nThe format is based on [Keep a Changel"
},
{
"path": "LICENSE",
"chars": 1075,
"preview": "MIT License\n\nCopyright (c) 2017-2022 Charles Jekel\n\nPermission is hereby granted, free of charge, to any person obtainin"
},
{
"path": "MANIFEST.in",
"chars": 35,
"preview": "include LICENSE\ninclude README.rst\n"
},
{
"path": "README.md",
"chars": 5181,
"preview": "# About\nA library for fitting continuous piecewise linear functions to data. Just specify the number of line segments yo"
},
{
"path": "README.rst",
"chars": 5693,
"preview": "About\n=====\n\nA library for fitting continuous piecewise linear functions to data.\nJust specify the number of line segmen"
},
{
"path": "convert_README_to_RST.sh",
"chars": 82,
"preview": "#!/usr/bin/env bash\npandoc --from=markdown --to=rst --output=README.rst README.md\n"
},
{
"path": "docs/.buildinfo",
"chars": 230,
"preview": "# Sphinx build info version 1\n# This file hashes the configuration used when building these files. When it is not found,"
},
{
"path": "docs/.nojekyll",
"chars": 0,
"preview": ""
},
{
"path": "docs/_sources/about.rst.txt",
"chars": 873,
"preview": "About\n============\n\nA library for fitting continuous piecewise linear functions to data. Just specify the number of line"
},
{
"path": "docs/_sources/examples.rst.txt",
"chars": 29927,
"preview": "Examples\n========\n\nAll of these examples will use the following data and imports.\n\n.. code:: python\n\n import numpy as "
},
{
"path": "docs/_sources/how_it_works.rst.txt",
"chars": 1232,
"preview": "How it works\n============\n\nThis\n`paper <https://github.com/cjekel/piecewise_linear_fit_py/raw/master/paper/pwlf_Jekel_Ve"
},
{
"path": "docs/_sources/index.rst.txt",
"chars": 298,
"preview": "pwlf: piecewise linear fitting\n================================\n\nFit piecewise linear functions to data!\n\n.. toctree::\n "
},
{
"path": "docs/_sources/installation.rst.txt",
"chars": 459,
"preview": "Installation\n============\n\nPython Package Index (PyPI)\n---------------------------\n\nYou can now install with pip.\n\n::\n\n "
},
{
"path": "docs/_sources/license.rst.txt",
"chars": 1092,
"preview": "License\n=======\n\nMIT License\n\nCopyright (c) 2017-2020 Charles Jekel\n\nPermission is hereby granted, free of charge, to an"
},
{
"path": "docs/_sources/modules.rst.txt",
"chars": 49,
"preview": "pwlf\n====\n\n.. toctree::\n :maxdepth: 4\n\n pwlf\n"
},
{
"path": "docs/_sources/pwlf.rst.txt",
"chars": 195,
"preview": "pwlf package contents\n============\n\n.. autosummary::\n :toctree: stubs\n\n pwlf.PiecewiseLinFit\n\n.. autoclass:: pwl"
},
{
"path": "docs/_sources/requirements.rst.txt",
"chars": 196,
"preview": "Requirements\n============\n\n`NumPy <https://pypi.org/project/numpy/>`__ (>= 1.14.0)\n\n`SciPy <https://pypi.org/project/sci"
},
{
"path": "docs/_sources/stubs/pwlf.PiecewiseLinFit.rst.txt",
"chars": 869,
"preview": "pwlf.PiecewiseLinFit\n====================\n\n.. currentmodule:: pwlf\n\n.. autoclass:: PiecewiseLinFit\n\n \n .. autometho"
},
{
"path": "docs/_static/alabaster.css",
"chars": 10776,
"preview": "/* -- page layout ----------------------------------------------------------- */\n\nbody {\n font-family: Georgia, serif"
},
{
"path": "docs/_static/basic.css",
"chars": 14817,
"preview": "/*\n * Sphinx stylesheet -- basic theme.\n */\n\n/* -- main layout ---------------------------------------------------------"
},
{
"path": "docs/_static/custom.css",
"chars": 42,
"preview": "/* This file intentionally left blank. */\n"
},
{
"path": "docs/_static/doctools.js",
"chars": 4322,
"preview": "/*\n * Base JavaScript utilities for all Sphinx HTML documentation.\n */\n\"use strict\";\n\nconst BLACKLISTED_KEY_CONTROL_ELEM"
},
{
"path": "docs/_static/documentation_options.js",
"chars": 328,
"preview": "const DOCUMENTATION_OPTIONS = {\n VERSION: '2.5.2',\n LANGUAGE: 'en',\n COLLAPSE_INDEX: false,\n BUILDER: 'html'"
},
{
"path": "docs/_static/language_data.js",
"chars": 4598,
"preview": "/*\n * This script contains the language-specific data used by searchtools.js,\n * namely the list of stopwords, stemmer, "
},
{
"path": "docs/_static/pygments.css",
"chars": 5359,
"preview": "pre { line-height: 125%; }\ntd.linenos .normal { color: inherit; background-color: transparent; padding-left: 5px; paddin"
},
{
"path": "docs/_static/searchtools.js",
"chars": 21463,
"preview": "/*\n * Sphinx JavaScript utilities for the full-text search.\n */\n\"use strict\";\n\n/**\n * Simple result scoring code.\n */\nif"
},
{
"path": "docs/_static/sphinx_highlight.js",
"chars": 5123,
"preview": "/* Highlighting utilities for Sphinx HTML documentation. */\n\"use strict\";\n\nconst SPHINX_HIGHLIGHT_ENABLED = true\n\n/**\n *"
},
{
"path": "docs/about.html",
"chars": 6051,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/examples.html",
"chars": 114464,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/genindex.html",
"chars": 8065,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/how_it_works.html",
"chars": 5673,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/index.html",
"chars": 8696,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/installation.html",
"chars": 6604,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/license.html",
"chars": 5219,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/modules.html",
"chars": 8637,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/pwlf.html",
"chars": 79178,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/requirements.html",
"chars": 4544,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/search.html",
"chars": 4034,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"./\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport\""
},
{
"path": "docs/searchindex.js",
"chars": 22548,
"preview": "Search.setIndex({\"alltitles\": {\"About\": [[0, null]], \"Conda\": [[4, \"conda\"]], \"Examples\": [[1, null]], \"From source\": [["
},
{
"path": "docs/stubs/pwlf.PiecewiseLinFit.html",
"chars": 32535,
"preview": "<!DOCTYPE html>\n\n<html lang=\"en\" data-content_root=\"../\">\n <head>\n <meta charset=\"utf-8\" />\n <meta name=\"viewport"
},
{
"path": "examples/EGO_integer_only.ipynb",
"chars": 101680,
"preview": "{\n \"cells\": [\n {\n \"cell_type\": \"code\",\n \"execution_count\": 1,\n \"metadata\": {},\n \"outputs\": [],\n \"source\": [\n "
},
{
"path": "examples/README.md",
"chars": 27051,
"preview": "# Examples\nAll of these examples will use the following data and imports.\n```python\nimport numpy as np\nimport matplotlib"
},
{
"path": "examples/README.rst",
"chars": 29927,
"preview": "Examples\n========\n\nAll of these examples will use the following data and imports.\n\n.. code:: python\n\n import numpy as "
},
{
"path": "examples/experiment_with_batch_process.py.ipynb",
"chars": 49457,
"preview": "{\n \"cells\": [\n {\n \"cell_type\": \"code\",\n \"execution_count\": 1,\n \"metadata\": {},\n \"outputs\": [],\n \"source\": [\n "
},
{
"path": "examples/fitForSpecifiedNumberOfLineSegments.py",
"chars": 4075,
"preview": "# fit for a specified number of line segments\n# you specify the number of line segments you want, the library does the r"
},
{
"path": "examples/fitForSpecifiedNumberOfLineSegments_passDiffEvoKeywords.py",
"chars": 3685,
"preview": "# fit for a specified number of line segments\n# you specify the number of line segments you want, the library does the r"
},
{
"path": "examples/fitForSpecifiedNumberOfLineSegments_standard_deviation.py",
"chars": 3584,
"preview": "# fit for a specified number of line segments\n# you specify the number of line segments you want, the library does the r"
},
{
"path": "examples/fitWithKnownLineSegmentLocations.py",
"chars": 3127,
"preview": "# fit and predict with known line segment x locations\n\n# import our libraries\nimport numpy as np\nimport matplotlib.pyplo"
},
{
"path": "examples/fit_begin_and_end.py",
"chars": 3207,
"preview": "# fit and predict between a known begging and known ending\n\n# import our libraries\nimport numpy as np\nimport matplotlib."
},
{
"path": "examples/min_length_demo.ipynb",
"chars": 24298,
"preview": "{\n \"cells\": [\n {\n \"cell_type\": \"code\",\n \"execution_count\": 1,\n \"metadata\": {},\n \"outputs\": [],\n \"source\": [\n "
},
{
"path": "examples/mixed_degree.py",
"chars": 414,
"preview": "import numpy as np\nimport matplotlib.pyplot as plt\nimport pwlf\n\nx = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]\ny = [1, 2, 3, 4, 4.2"
},
{
"path": "examples/mixed_degree_forcing_slope.py",
"chars": 1296,
"preview": "import numpy as np\nfrom scipy.optimize import differential_evolution\nimport matplotlib.pyplot as plt\nimport pwlf\n\nx = [1"
},
{
"path": "examples/model_persistence.py",
"chars": 3182,
"preview": "# import our libraries\nimport numpy as np\nimport pwlf\n\n# if you use Python 2.x you should import cPickle\n# import cPickl"
},
{
"path": "examples/model_persistence_prediction.py",
"chars": 3094,
"preview": "# import our libraries\nimport numpy as np\nimport pwlf\n\n# your data\ny = np.array([0.00000000e+00, 9.69801700e-03, 2.94350"
},
{
"path": "examples/prediction_variance.py",
"chars": 3409,
"preview": "# fit for a specified number of line segments\n# you specify the number of line segments you want, the library does the r"
},
{
"path": "examples/prediction_variance_degree2.py",
"chars": 3419,
"preview": "# fit for a specified number of line segments\n# you specify the number of line segments you want, the library does the r"
},
{
"path": "examples/robust_regression.py",
"chars": 2980,
"preview": "# import our libraries\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport pwlf\nfrom scipy.optimize import least_s"
},
{
"path": "examples/run_opt_to_find_best_number_of_line_segments.py",
"chars": 4221,
"preview": "# Running an optimization to find the best number of line segments\n\n# import our libraries\nimport numpy as np\nimport mat"
},
{
"path": "examples/sineWave.py",
"chars": 1029,
"preview": "# fit for a specified number of line segments\n# you specify the number of line segments you want, the library does the r"
},
{
"path": "examples/sineWave_custom_opt_bounds.py",
"chars": 1012,
"preview": "# fit for a specified number of line segments\n# you specify the number of line segments you want, the library does the r"
},
{
"path": "examples/sineWave_degrees.py",
"chars": 1136,
"preview": "# import our libraries\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport pwlf\n\n# generate sin wave data\nx = np.l"
},
{
"path": "examples/sineWave_time_compare.py",
"chars": 1593,
"preview": "# fit for a specified number of line segments\n# you specify the number of line segments you want, the library does the r"
},
{
"path": "examples/slope_constraint_demo.ipynb",
"chars": 22441,
"preview": "{\n \"cells\": [\n {\n \"cell_type\": \"code\",\n \"execution_count\": 1,\n \"metadata\": {},\n \"outputs\": [\n {\n \"name\":"
},
{
"path": "examples/stack_overflow_example.py",
"chars": 876,
"preview": "import numpy as np\nimport matplotlib.pyplot as plt\nimport pwlf\n\n# https://stackoverflow.com/questions/29382903/how-to-ap"
},
{
"path": "examples/standard_errrors_and_p-values.py",
"chars": 3655,
"preview": "# example of how to calculate standard errors and p-values\nfrom __future__ import print_function\nimport numpy as np\nimpo"
},
{
"path": "examples/standard_errrors_and_p-values_non-linear.py",
"chars": 1515,
"preview": "from __future__ import print_function\nimport numpy as np\nimport pwlf\n# import matplotlib.pyplot as plt\n\n# generate a tru"
},
{
"path": "examples/test0.py",
"chars": 3477,
"preview": "import numpy as np\nimport matplotlib.pyplot as plt\nimport pwlf\nfrom scipy.optimize import minimize\n\ny = np.array([0.0000"
},
{
"path": "examples/test_for_model_significance.py",
"chars": 4239,
"preview": "# example of how to calculate standard errors and p-values\nfrom __future__ import print_function\nimport numpy as np\nimpo"
},
{
"path": "examples/test_if_breaks_exact.py",
"chars": 993,
"preview": "import numpy as np\nimport pwlf\n\nx = np.array((0.0, 1.0, 1.5, 2.0))\ny = np.array((0.0, 1.0, 1.1, 1.5))\n\nmy_fit1 = pwlf.Pi"
},
{
"path": "examples/tf/fit_begin_and_end.py",
"chars": 3285,
"preview": "# fit and predict between a known begging and known ending\n\n# import our libraries\nimport numpy as np\nimport matplotlib."
},
{
"path": "examples/tf/sine_benchmark_fixed_20_break_points.py",
"chars": 975,
"preview": "import numpy as np\nimport pwlf\nfrom time import time\nimport os\n\nbreaks = np.linspace(0.0, 10.0, num=21)\n\nn = np.logspace"
},
{
"path": "examples/tf/sine_benchmark_six_segments.py",
"chars": 840,
"preview": "import numpy as np\nimport matplotlib.pyplot as plt\nimport pwlf\nfrom time import time\n\n# set random seed\nnp.random.seed(2"
},
{
"path": "examples/tf/test_fit.py",
"chars": 3135,
"preview": "# fit and predict with known line segment x locations\n\n# import our libraries\nimport numpy as np\nimport matplotlib.pyplo"
},
{
"path": "examples/understanding_higher_degrees/polynomials_in_pwlf.ipynb",
"chars": 150359,
"preview": "{\n \"cells\": [\n {\n \"cell_type\": \"code\",\n \"execution_count\": 1,\n \"metadata\": {},\n \"outputs\": [],\n \"source\": [\n "
},
{
"path": "examples/useCustomOptimizationRoutine.py",
"chars": 3674,
"preview": "# use your own custom optimization routine to find the optimal location\n# of line segment locations\n\n# import our librar"
},
{
"path": "examples/weighted_least_squares_ex.py",
"chars": 1245,
"preview": "from time import time\nimport os\nos.environ['OMP_NUM_THREADS'] = '1'\nimport numpy as np\nimport matplotlib.pyplot as plt\ni"
},
{
"path": "examples/weighted_least_squares_ex_stats.py",
"chars": 1460,
"preview": "from time import time\nimport os\nos.environ['OMP_NUM_THREADS'] = '1'\nimport numpy as np\nimport matplotlib.pyplot as plt\ni"
},
{
"path": "pwlf/__init__.py",
"chars": 93,
"preview": "from .pwlf import PiecewiseLinFit # noqa F401\nfrom .version import __version__ # noqa F401\n"
},
{
"path": "pwlf/pwlf.py",
"chars": 64960,
"preview": "# -- coding: utf-8 --\n# MIT License\n#\n# Copyright (c) 2017-2022 Charles Jekel\n#\n# Permission is hereby granted, free of "
},
{
"path": "pwlf/version.py",
"chars": 22,
"preview": "__version__ = \"2.5.2\"\n"
},
{
"path": "setup.py",
"chars": 782,
"preview": "import io\nfrom distutils.core import setup\n\n# load the version from version.py\nversion = {}\nwith open(\"pwlf/version.py\")"
},
{
"path": "sphinx_build_script.sh",
"chars": 536,
"preview": "\n#!/usr/bin/env bash\npip install .\n# clean docs and doctrees\nrm -r docs/*\nrm -r doctrees/*\n# make the documentation in g"
},
{
"path": "sphinxdocs/Makefile",
"chars": 584,
"preview": "# Minimal makefile for Sphinx documentation\n#\n\n# You can set these variables from the command line.\nSPHINXOPTS =\nSPHI"
},
{
"path": "sphinxdocs/make.bat",
"chars": 756,
"preview": "@ECHO OFF\n\npushd %~dp0\n\nREM Command file for Sphinx documentation\n\nif \"%SPHINXBUILD%\" == \"\" (\n\tset SPHINXBUILD=sphinx-bu"
},
{
"path": "sphinxdocs/source/about.rst",
"chars": 873,
"preview": "About\n============\n\nA library for fitting continuous piecewise linear functions to data. Just specify the number of line"
},
{
"path": "sphinxdocs/source/conf.py",
"chars": 5596,
"preview": "# -*- coding: utf-8 -*-\n#\n# Configuration file for the Sphinx documentation builder.\n#\n# This file does only contain a s"
},
{
"path": "sphinxdocs/source/examples.rst",
"chars": 29927,
"preview": "Examples\n========\n\nAll of these examples will use the following data and imports.\n\n.. code:: python\n\n import numpy as "
},
{
"path": "sphinxdocs/source/how_it_works.rst",
"chars": 1232,
"preview": "How it works\n============\n\nThis\n`paper <https://github.com/cjekel/piecewise_linear_fit_py/raw/master/paper/pwlf_Jekel_Ve"
},
{
"path": "sphinxdocs/source/index.rst",
"chars": 298,
"preview": "pwlf: piecewise linear fitting\n================================\n\nFit piecewise linear functions to data!\n\n.. toctree::\n "
},
{
"path": "sphinxdocs/source/installation.rst",
"chars": 459,
"preview": "Installation\n============\n\nPython Package Index (PyPI)\n---------------------------\n\nYou can now install with pip.\n\n::\n\n "
},
{
"path": "sphinxdocs/source/license.rst",
"chars": 1092,
"preview": "License\n=======\n\nMIT License\n\nCopyright (c) 2017-2020 Charles Jekel\n\nPermission is hereby granted, free of charge, to an"
},
{
"path": "sphinxdocs/source/modules.rst",
"chars": 49,
"preview": "pwlf\n====\n\n.. toctree::\n :maxdepth: 4\n\n pwlf\n"
},
{
"path": "sphinxdocs/source/pwlf.rst",
"chars": 195,
"preview": "pwlf package contents\n============\n\n.. autosummary::\n :toctree: stubs\n\n pwlf.PiecewiseLinFit\n\n.. autoclass:: pwl"
},
{
"path": "sphinxdocs/source/requirements.rst",
"chars": 196,
"preview": "Requirements\n============\n\n`NumPy <https://pypi.org/project/numpy/>`__ (>= 1.14.0)\n\n`SciPy <https://pypi.org/project/sci"
},
{
"path": "tests/__init__.py",
"chars": 0,
"preview": ""
},
{
"path": "tests/tests.py",
"chars": 24700,
"preview": "import numpy as np\nimport unittest\nimport pwlf\n\n\nclass TestEverything(unittest.TestCase):\n\n x_small = np.array((0.0, "
}
]
// ... and 13 more files (download for full content)
About this extraction
This page contains the full source code of the cjekel/piecewise_linear_fit_py GitHub repository, extracted and formatted as plain text for AI agents and large language models (LLMs). The extraction includes 112 files (1.0 MB), approximately 441.2k tokens, and a symbol index with 98 extracted functions, classes, methods, constants, and types. Use this with OpenClaw, Claude, ChatGPT, Cursor, Windsurf, or any other AI tool that accepts text input. You can copy the full output to your clipboard or download it as a .txt file.
Extracted by GitExtract — free GitHub repo to text converter for AI. Built by Nikandr Surkov.