 [](https://codecov.io/gh/dfki-ric/pytransform3d)
[](https://doi.org/10.21105/joss.01159)
[](https://doi.org/10.5281/zenodo.2553450)
# pytransform3d
A Python library for transformations in three dimensions.
pytransform3d offers...
* operations like concatenation and inversion for most common representations
of rotation (orientation) and translation (position)
* conversions between those representations
* clear documentation of transformation conventions
* tight coupling with matplotlib to quickly visualize (or animate)
transformations
* the TransformManager which manages complex chains of transformations
(with export to graph visualization as PNG, additionally requires pydot)
* the TransformEditor which allows to modify transformations graphically
(additionally requires PyQt4/5)
* the UrdfTransformManager which is able to load transformations from
[URDF](https://wiki.ros.org/urdf) files (additionally requires lxml)
* a matplotlib-like interface to Open3D's visualizer to display and animate
geometries and transformations (additionally requires Open3D)
pytransform3d is used in various domains, for example:
* specifying motions of a robot
* learning robot movements from human demonstration
* sensor fusion for human pose estimation
* collision detection for robots
The API documentation can be found
[here](https://dfki-ric.github.io/pytransform3d/).
I gave a talk at EuroSciPy 2023 about pytransform3d. Slides are available
[here](https://github.com/AlexanderFabisch/pytransform3d_euroscipy2023/).
If you need similar features in JAX (on GPU, vectorized, differentiable),
have a look at the experimental library
[jaxtransform3d](https://github.com/AlexanderFabisch/jaxtransform3d/).
## Installation
Use pip to install the package from PyPI:
```bash
pip install 'pytransform3d[all]'
```
or conda:
```bash
conda install -c conda-forge pytransform3d
```
Take a look at the
[installation instructions](https://dfki-ric.github.io/pytransform3d/install.html)
in the documentation for more details.
## Gallery
The following plots and visualizations have been generated with pytransform3d.
The code for most examples can be found in
[the documentation](https://dfki-ric.github.io/pytransform3d/_auto_examples/index.html).
Left: [Nao robot](https://www.softbankrobotics.com/emea/en/nao) with URDF
from [Bullet3](https://github.com/bulletphysics/bullet3).
Right: [Kuka iiwa](https://www.kuka.com/en-de/products/robot-systems/industrial-robots/lbr-iiwa).
The animation is based on pytransform3d's visualization interface to
[Open3D](http://www.open3d.org/).
[](https://codecov.io/gh/dfki-ric/pytransform3d)
[](https://doi.org/10.21105/joss.01159)
[](https://doi.org/10.5281/zenodo.2553450)
# pytransform3d
A Python library for transformations in three dimensions.
pytransform3d offers...
* operations like concatenation and inversion for most common representations
of rotation (orientation) and translation (position)
* conversions between those representations
* clear documentation of transformation conventions
* tight coupling with matplotlib to quickly visualize (or animate)
transformations
* the TransformManager which manages complex chains of transformations
(with export to graph visualization as PNG, additionally requires pydot)
* the TransformEditor which allows to modify transformations graphically
(additionally requires PyQt4/5)
* the UrdfTransformManager which is able to load transformations from
[URDF](https://wiki.ros.org/urdf) files (additionally requires lxml)
* a matplotlib-like interface to Open3D's visualizer to display and animate
geometries and transformations (additionally requires Open3D)
pytransform3d is used in various domains, for example:
* specifying motions of a robot
* learning robot movements from human demonstration
* sensor fusion for human pose estimation
* collision detection for robots
The API documentation can be found
[here](https://dfki-ric.github.io/pytransform3d/).
I gave a talk at EuroSciPy 2023 about pytransform3d. Slides are available
[here](https://github.com/AlexanderFabisch/pytransform3d_euroscipy2023/).
If you need similar features in JAX (on GPU, vectorized, differentiable),
have a look at the experimental library
[jaxtransform3d](https://github.com/AlexanderFabisch/jaxtransform3d/).
## Installation
Use pip to install the package from PyPI:
```bash
pip install 'pytransform3d[all]'
```
or conda:
```bash
conda install -c conda-forge pytransform3d
```
Take a look at the
[installation instructions](https://dfki-ric.github.io/pytransform3d/install.html)
in the documentation for more details.
## Gallery
The following plots and visualizations have been generated with pytransform3d.
The code for most examples can be found in
[the documentation](https://dfki-ric.github.io/pytransform3d/_auto_examples/index.html).
Left: [Nao robot](https://www.softbankrobotics.com/emea/en/nao) with URDF
from [Bullet3](https://github.com/bulletphysics/bullet3).
Right: [Kuka iiwa](https://www.kuka.com/en-de/products/robot-systems/industrial-robots/lbr-iiwa).
The animation is based on pytransform3d's visualization interface to
[Open3D](http://www.open3d.org/).
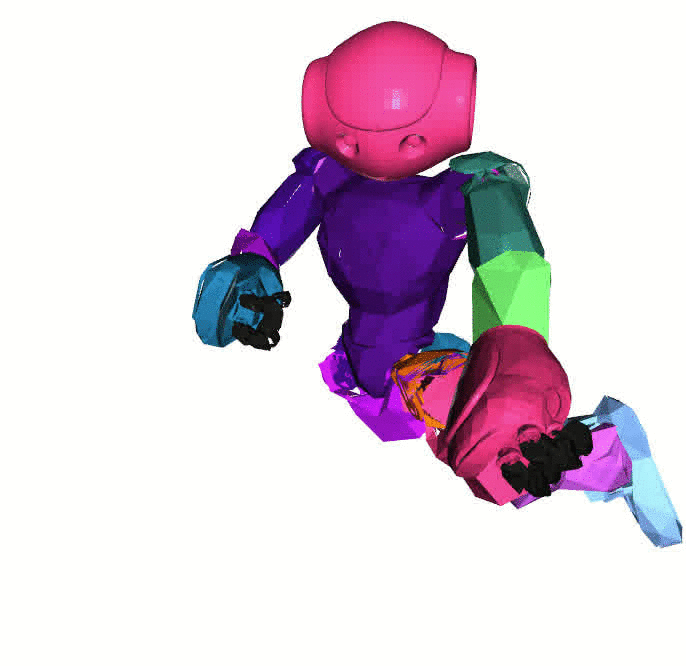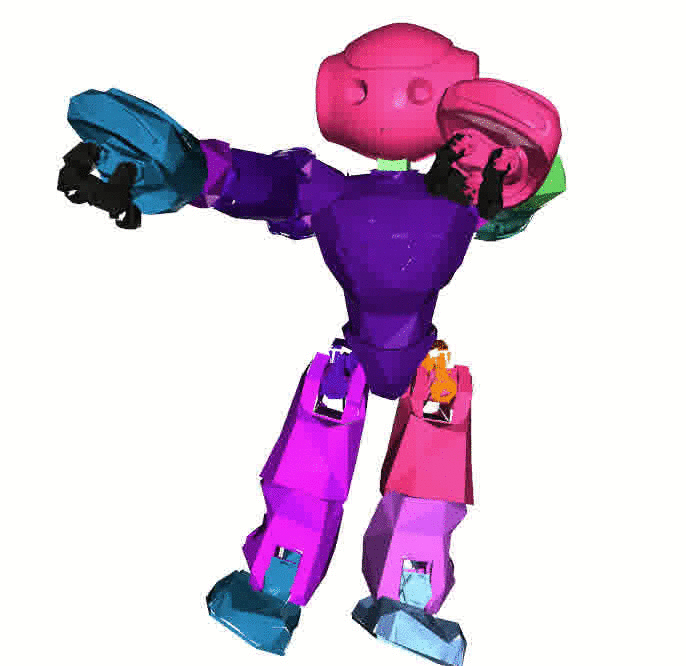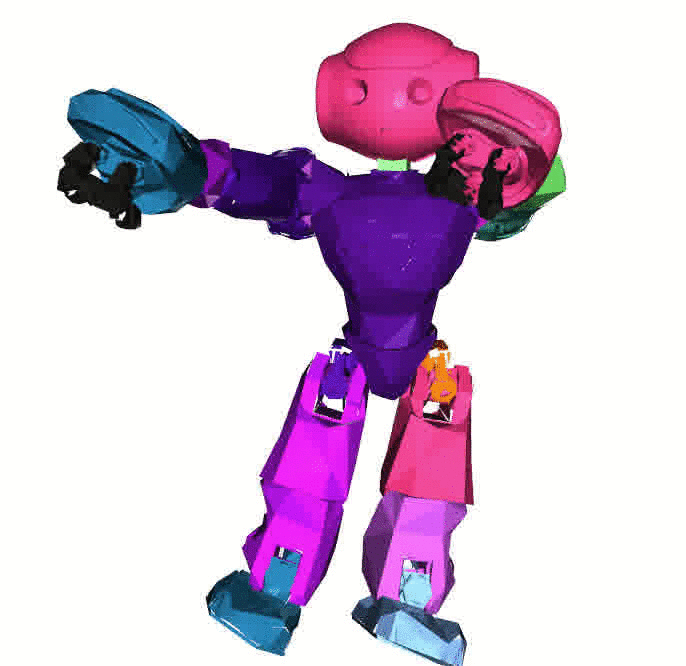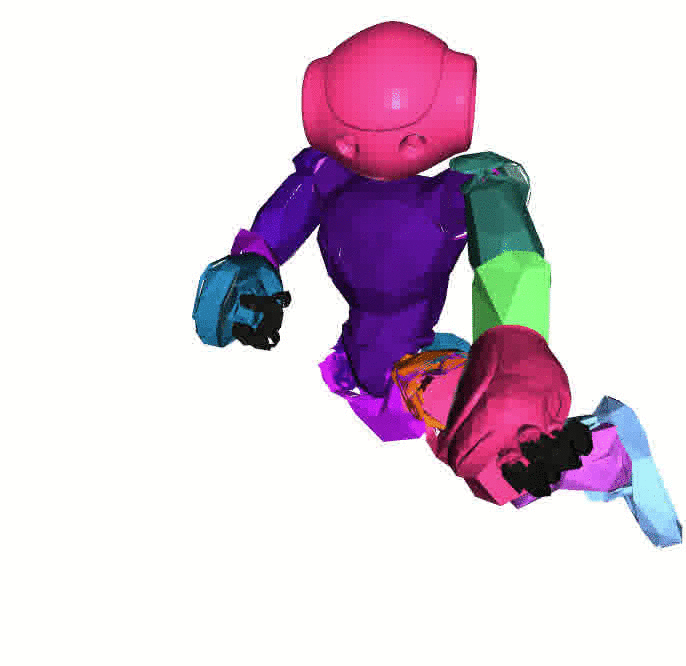
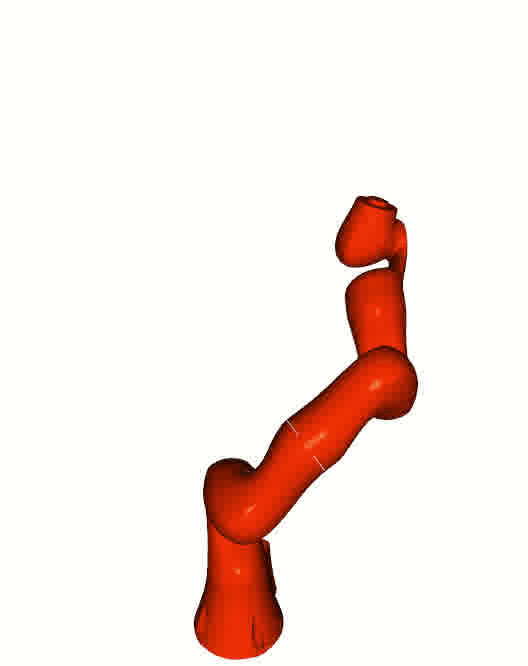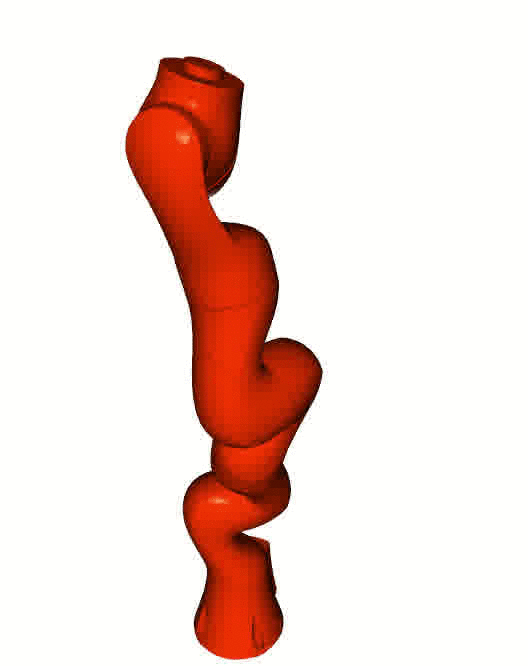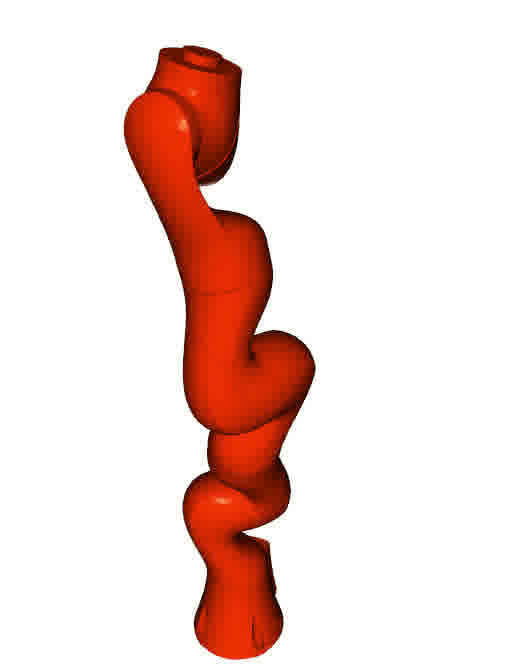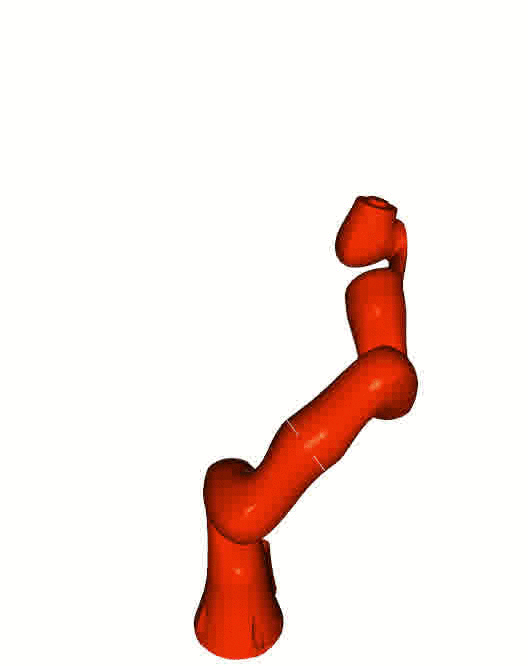
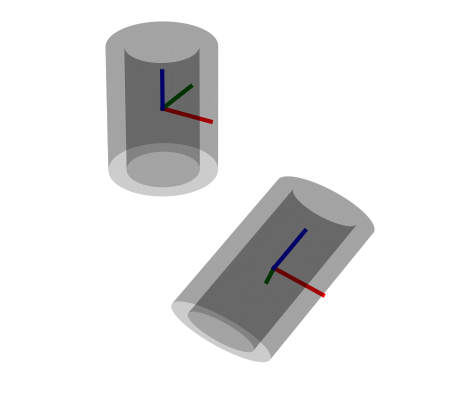
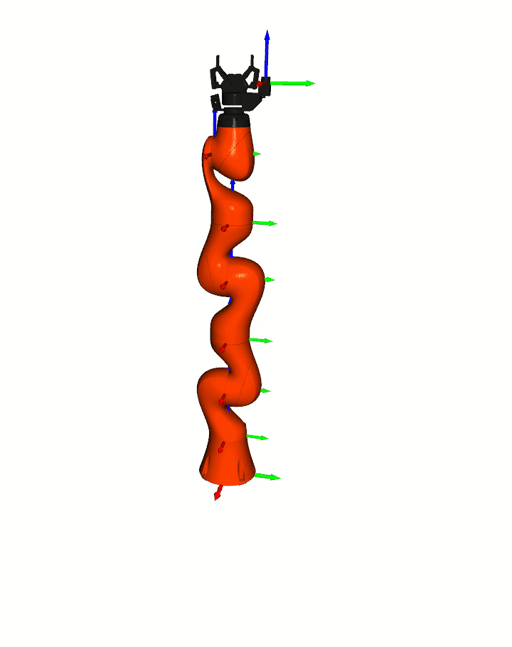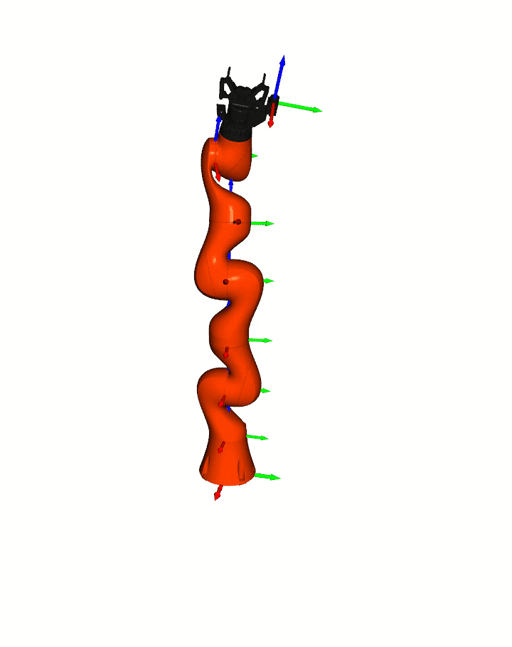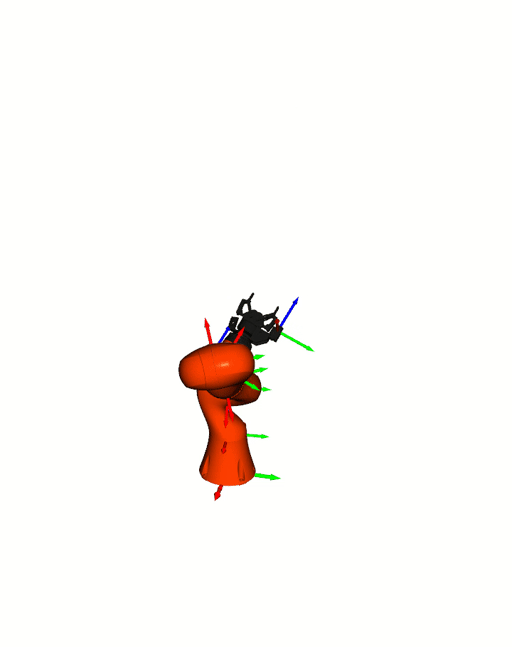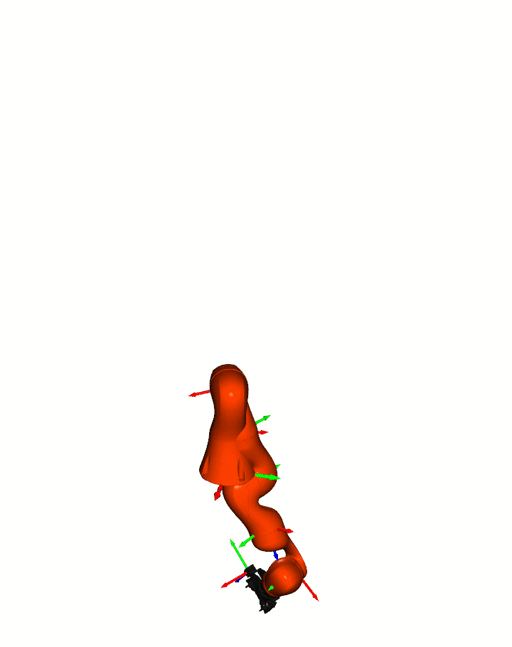 Visualizations based on [Open3D](http://www.open3d.org/).
Visualizations based on [Open3D](http://www.open3d.org/).
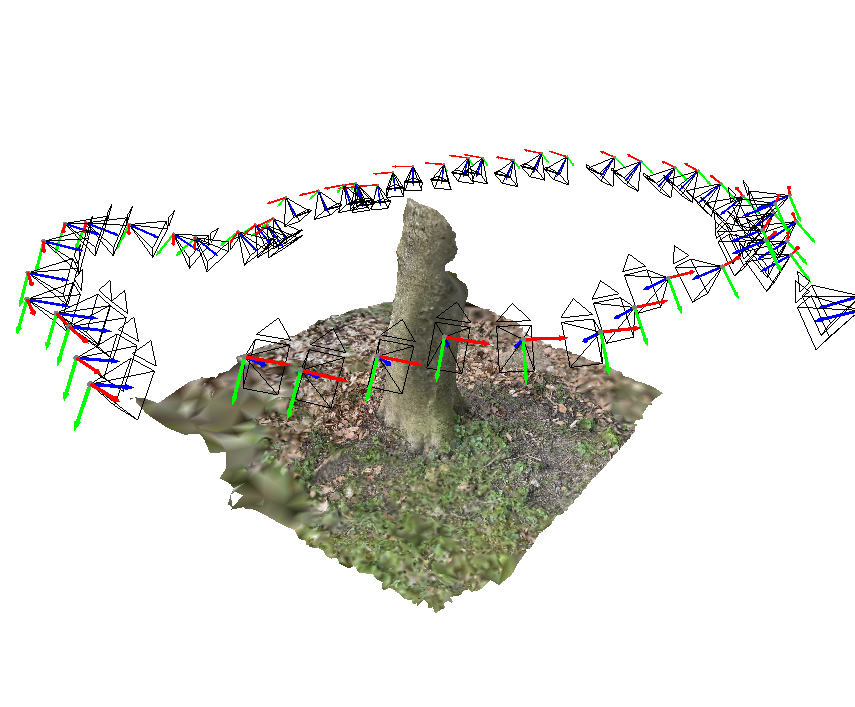
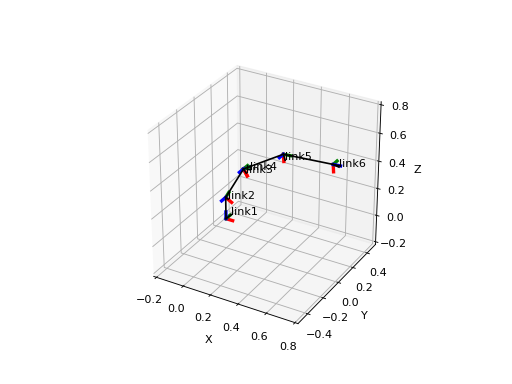
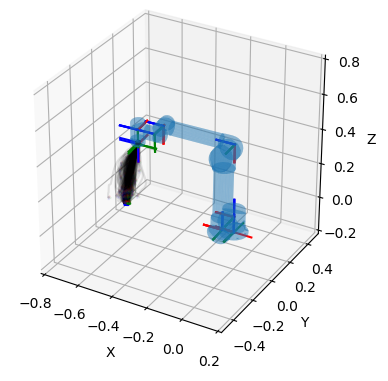
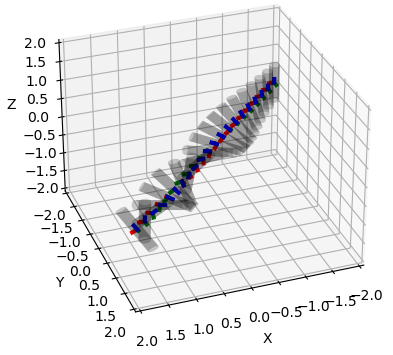
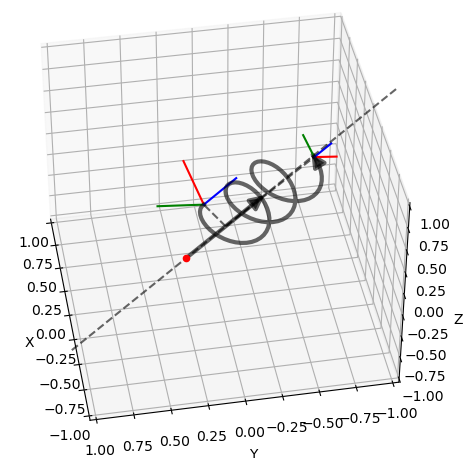
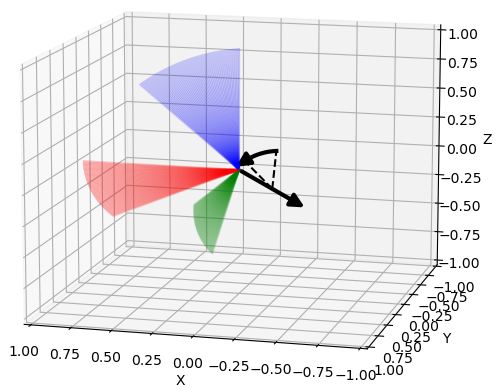
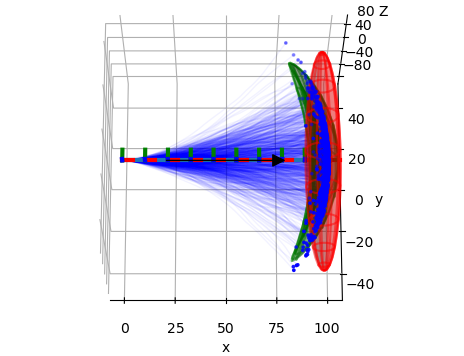
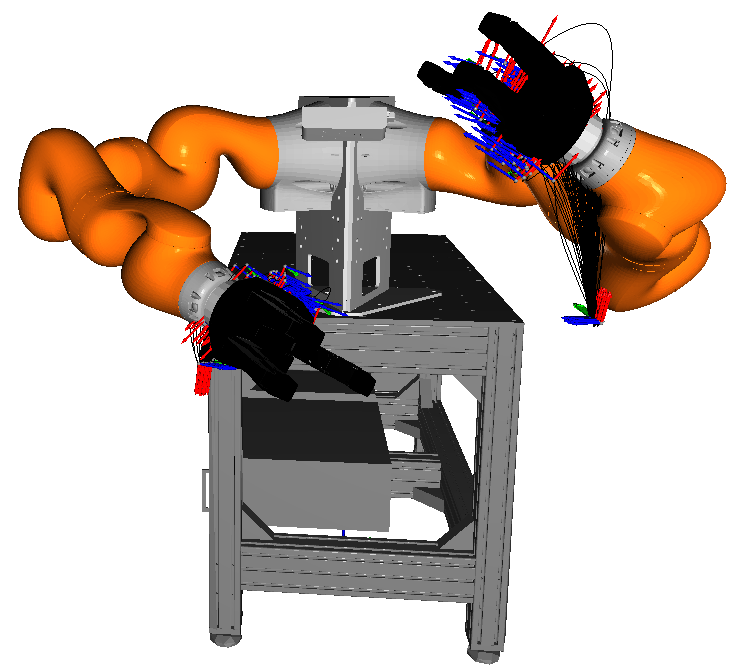 Various plots based on Matplotlib.
Various plots based on Matplotlib.
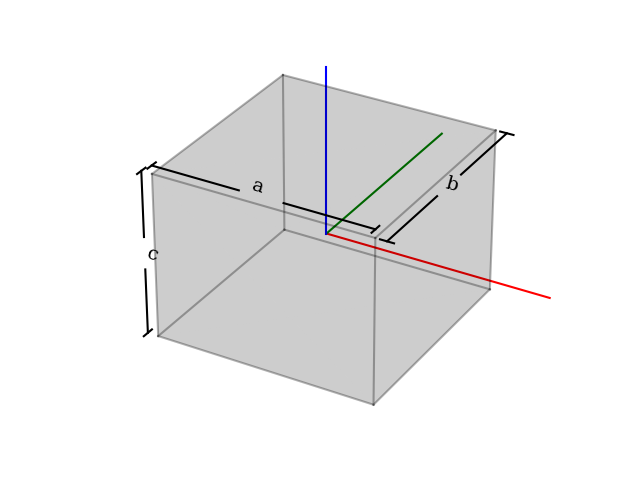
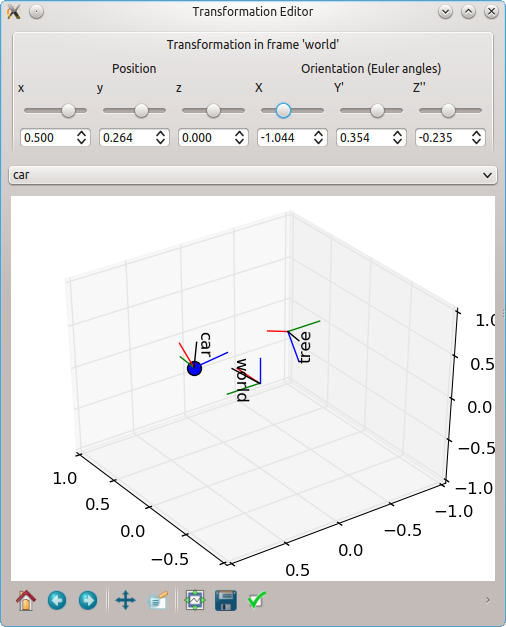
 Transformation editor based on Qt.
Transformation editor based on Qt.
 ## Example
This is just one simple example. You can find more examples in the subfolder
`examples/`.
```python
import numpy as np
import matplotlib.pyplot as plt
from pytransform3d import rotations as pr
from pytransform3d import transformations as pt
from pytransform3d.transform_manager import TransformManager
rng = np.random.default_rng(0)
ee2robot = pt.transform_from_pq(
np.hstack((np.array([0.4, -0.3, 0.5]),
pr.random_quaternion(rng))))
cam2robot = pt.transform_from_pq(
np.hstack((np.array([0.0, 0.0, 0.8]), pr.q_id)))
object2cam = pt.transform_from(
pr.active_matrix_from_intrinsic_euler_xyz(np.array([0.0, 0.0, -0.5])),
np.array([0.5, 0.1, 0.1]))
tm = TransformManager()
tm.add_transform("end-effector", "robot", ee2robot)
tm.add_transform("camera", "robot", cam2robot)
tm.add_transform("object", "camera", object2cam)
ee2object = tm.get_transform("end-effector", "object")
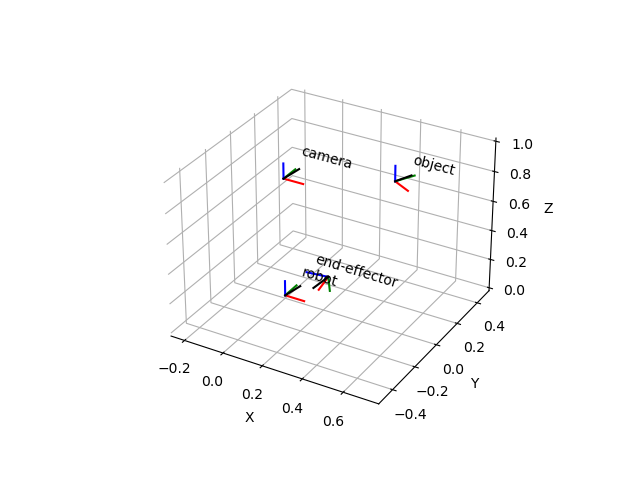
ax = tm.plot_frames_in("robot", s=0.1)
ax.set_xlim((-0.25, 0.75))
ax.set_ylim((-0.5, 0.5))
ax.set_zlim((0.0, 1.0))
plt.show()
```
## Documentation
The API documentation can be found
[here](https://dfki-ric.github.io/pytransform3d/).
The documentation can be found in the directory `doc`.
To build the documentation, run e.g. (on linux):
```bash
cd doc
make html
```
The HTML documentation is now located at `doc/build/html/index.html`.
Execute the following command in the main folder of the repository
to install the dependencies:
```bash
pip install -e '.[doc]'
```
## Tests
You can use pytest to run the tests of this project in the root directory:
```bash
pytest
```
A coverage report will be located at `htmlcov/index.html`.
Note that you have to install `pytest` to run the tests and `pytest-cov` to
obtain the code coverage report.
## Contributing
If you wish to report bugs, please use the
[issue tracker](https://github.com/dfki-ric/pytransform3d/issues) at
Github. If you would like to contribute to pytransform3d, just open an issue
or a [pull request](https://github.com/dfki-ric/pytransform3d/pulls).
The target branch for pull requests is the develop branch.
The development branch will be merged to main for new releases.
If you have questions about the software, you should ask them in the
[discussion section](https://github.com/dfki-ric/pytransform3d/discussions).
The recommended workflow to add a new feature, add documentation, or fix a bug
is the following:
* Push your changes to a branch (e.g. `feature/x`, `doc/y`, or `fix/z`) of your
fork of the pytransform3d repository.
* Open a pull request to the latest development branch. There is usually an
open merge request from the latest development branch to the main branch.
* When the latest development branch is merged to the main branch, a new
release will be made.
Note that there is a
[checklist](https://github.com/dfki-ric/pytransform3d/wiki#checklist-for-new-features)
for new features.
It is forbidden to directly push to the main branch. Each new version
has its own development branch from which a pull request will be opened to the
main branch. Only the maintainer of the software is allowed to merge a
development branch to the main branch.
## License
The library is distributed under the
[3-Clause BSD license](https://github.com/dfki-ric/pytransform3d/blob/main/LICENSE).
## Citation
If you use pytransform3d for a scientific publication, I would appreciate
citation of the following paper:
Fabisch, A. (2019). pytransform3d: 3D Transformations for Python.
Journal of Open Source Software, 4(33), 1159,
[](https://doi.org/10.21105/joss.01159)
Bibtex entry:
```bibtex
@article{Fabisch2019,
doi = {10.21105/joss.01159},
url = {https://doi.org/10.21105/joss.01159},
year = {2019},
publisher = {The Open Journal},
volume = {4},
number = {33},
pages = {1159},
author = {Alexander Fabisch},
title = {pytransform3d: 3D Transformations for Python},
journal = {Journal of Open Source Software}
}
```
================================================
FILE: doc/Makefile
================================================
# Makefile for Sphinx documentation
#
# You can set these variables from the command line.
SPHINXOPTS =
SPHINXBUILD = sphinx-build
PAPER =
BUILDDIR = build
# User-friendly check for sphinx-build
ifeq ($(shell which $(SPHINXBUILD) >/dev/null 2>&1; echo $$?), 1)
$(error The '$(SPHINXBUILD)' command was not found. Make sure you have Sphinx installed, then set the SPHINXBUILD environment variable to point to the full path of the '$(SPHINXBUILD)' executable. Alternatively you can add the directory with the executable to your PATH. If you don't have Sphinx installed, grab it from http://sphinx-doc.org/)
endif
# Internal variables.
PAPEROPT_a4 = -D latex_paper_size=a4
PAPEROPT_letter = -D latex_paper_size=letter
ALLSPHINXOPTS = -d $(BUILDDIR)/doctrees $(PAPEROPT_$(PAPER)) $(SPHINXOPTS) source
# the i18n builder cannot share the environment and doctrees with the others
I18NSPHINXOPTS = $(PAPEROPT_$(PAPER)) $(SPHINXOPTS) source
.PHONY: help clean html dirhtml singlehtml pickle json htmlhelp qthelp devhelp epub latex latexpdf text man changes linkcheck doctest gettext
help:
@echo "Please use \`make
## Example
This is just one simple example. You can find more examples in the subfolder
`examples/`.
```python
import numpy as np
import matplotlib.pyplot as plt
from pytransform3d import rotations as pr
from pytransform3d import transformations as pt
from pytransform3d.transform_manager import TransformManager
rng = np.random.default_rng(0)
ee2robot = pt.transform_from_pq(
np.hstack((np.array([0.4, -0.3, 0.5]),
pr.random_quaternion(rng))))
cam2robot = pt.transform_from_pq(
np.hstack((np.array([0.0, 0.0, 0.8]), pr.q_id)))
object2cam = pt.transform_from(
pr.active_matrix_from_intrinsic_euler_xyz(np.array([0.0, 0.0, -0.5])),
np.array([0.5, 0.1, 0.1]))
tm = TransformManager()
tm.add_transform("end-effector", "robot", ee2robot)
tm.add_transform("camera", "robot", cam2robot)
tm.add_transform("object", "camera", object2cam)
ee2object = tm.get_transform("end-effector", "object")
ax = tm.plot_frames_in("robot", s=0.1)
ax.set_xlim((-0.25, 0.75))
ax.set_ylim((-0.5, 0.5))
ax.set_zlim((0.0, 1.0))
plt.show()
```

## Documentation
The API documentation can be found
[here](https://dfki-ric.github.io/pytransform3d/).
The documentation can be found in the directory `doc`.
To build the documentation, run e.g. (on linux):
```bash
cd doc
make html
```
The HTML documentation is now located at `doc/build/html/index.html`.
Execute the following command in the main folder of the repository
to install the dependencies:
```bash
pip install -e '.[doc]'
```
## Tests
You can use pytest to run the tests of this project in the root directory:
```bash
pytest
```
A coverage report will be located at `htmlcov/index.html`.
Note that you have to install `pytest` to run the tests and `pytest-cov` to
obtain the code coverage report.
## Contributing
If you wish to report bugs, please use the
[issue tracker](https://github.com/dfki-ric/pytransform3d/issues) at
Github. If you would like to contribute to pytransform3d, just open an issue
or a [pull request](https://github.com/dfki-ric/pytransform3d/pulls).
The target branch for pull requests is the develop branch.
The development branch will be merged to main for new releases.
If you have questions about the software, you should ask them in the
[discussion section](https://github.com/dfki-ric/pytransform3d/discussions).
The recommended workflow to add a new feature, add documentation, or fix a bug
is the following:
* Push your changes to a branch (e.g. `feature/x`, `doc/y`, or `fix/z`) of your
fork of the pytransform3d repository.
* Open a pull request to the latest development branch. There is usually an
open merge request from the latest development branch to the main branch.
* When the latest development branch is merged to the main branch, a new
release will be made.
Note that there is a
[checklist](https://github.com/dfki-ric/pytransform3d/wiki#checklist-for-new-features)
for new features.
It is forbidden to directly push to the main branch. Each new version
has its own development branch from which a pull request will be opened to the
main branch. Only the maintainer of the software is allowed to merge a
development branch to the main branch.
## License
The library is distributed under the
[3-Clause BSD license](https://github.com/dfki-ric/pytransform3d/blob/main/LICENSE).
## Citation
If you use pytransform3d for a scientific publication, I would appreciate
citation of the following paper:
Fabisch, A. (2019). pytransform3d: 3D Transformations for Python.
Journal of Open Source Software, 4(33), 1159,
[](https://doi.org/10.21105/joss.01159)
Bibtex entry:
```bibtex
@article{Fabisch2019,
doi = {10.21105/joss.01159},
url = {https://doi.org/10.21105/joss.01159},
year = {2019},
publisher = {The Open Journal},
volume = {4},
number = {33},
pages = {1159},
author = {Alexander Fabisch},
title = {pytransform3d: 3D Transformations for Python},
journal = {Journal of Open Source Software}
}
```
================================================
FILE: doc/Makefile
================================================
# Makefile for Sphinx documentation
#
# You can set these variables from the command line.
SPHINXOPTS =
SPHINXBUILD = sphinx-build
PAPER =
BUILDDIR = build
# User-friendly check for sphinx-build
ifeq ($(shell which $(SPHINXBUILD) >/dev/null 2>&1; echo $$?), 1)
$(error The '$(SPHINXBUILD)' command was not found. Make sure you have Sphinx installed, then set the SPHINXBUILD environment variable to point to the full path of the '$(SPHINXBUILD)' executable. Alternatively you can add the directory with the executable to your PATH. If you don't have Sphinx installed, grab it from http://sphinx-doc.org/)
endif
# Internal variables.
PAPEROPT_a4 = -D latex_paper_size=a4
PAPEROPT_letter = -D latex_paper_size=letter
ALLSPHINXOPTS = -d $(BUILDDIR)/doctrees $(PAPEROPT_$(PAPER)) $(SPHINXOPTS) source
# the i18n builder cannot share the environment and doctrees with the others
I18NSPHINXOPTS = $(PAPEROPT_$(PAPER)) $(SPHINXOPTS) source
.PHONY: help clean html dirhtml singlehtml pickle json htmlhelp qthelp devhelp epub latex latexpdf text man changes linkcheck doctest gettext
help:
@echo "Please use \`make | Right-handed | Left-handed |
|---|---|
| .. plot:: :width: 400px import numpy as np import matplotlib.pyplot as plt from pytransform3d.plot_utils import make_3d_axis plt.figure() ax = make_3d_axis(1) plt.setp(ax, xlim=(-0.05, 1.05), ylim=(-0.05, 1.05), zlim=(-0.05, 1.05), xlabel="X", ylabel="Y", zlabel="Z") basis = np.eye(3) for d, c in enumerate(["r", "g", "b"]): ax.plot([0.0, basis[0, d]], [0.0, basis[1, d]], [0.0, basis[2, d]], color=c, lw=5) plt.show() .. raw:: html | .. plot:: :width: 400px import numpy as np import matplotlib.pyplot as plt from pytransform3d.plot_utils import make_3d_axis plt.figure() ax = make_3d_axis(1) plt.setp(ax, xlim=(-0.05, 1.05), ylim=(-0.05, 1.05), zlim=(-1.05, 0.05), xlabel="X", ylabel="Y", zlabel="Z") basis = np.eye(3) basis[:, 2] *= -1.0 for d, c in enumerate(["r", "g", "b"]): ax.plot([0.0, basis[0, d]], [0.0, basis[1, d]], [0.0, basis[2, d]], color=c, lw=5) plt.show() .. raw:: html |
| Active | Passive |
|---|---|
| .. plot:: :width: 400px import numpy as np import matplotlib.pyplot as plt from pytransform3d.transformations import transform, plot_transform from pytransform3d.plot_utils import make_3d_axis, Arrow3D plt.figure() ax = make_3d_axis(1) plt.setp(ax, xlim=(-1.05, 1.05), ylim=(-0.55, 1.55), zlim=(-1.05, 1.05), xlabel="X", ylabel="Y", zlabel="Z") ax.view_init(elev=90, azim=-90) ax.set_xticks(()) ax.set_yticks(()) ax.set_zticks(()) rng = np.random.default_rng(42) PA = np.ones((10, 4)) PA[:, :3] = 0.1 * rng.standard_normal(size=(10, 3)) PA[:, 0] += 0.3 PA[:, :3] += 0.3 x_translation = -0.1 y_translation = 0.2 z_rotation = np.pi / 4.0 A2B = np.array([ [np.cos(z_rotation), -np.sin(z_rotation), 0.0, x_translation], [np.sin(z_rotation), np.cos(z_rotation), 0.0, y_translation], [0.0, 0.0, 1.0, 0.0], [0.0, 0.0, 0.0, 1.0] ]) PB = transform(A2B, PA) plot_transform(ax=ax, A2B=np.eye(4)) ax.scatter(PA[:, 0], PA[:, 1], PA[:, 2], c="orange") plot_transform(ax=ax, A2B=A2B, ls="--", alpha=0.5) ax.scatter(PB[:, 0], PB[:, 1], PB[:, 2], c="cyan") axis_arrow = Arrow3D( [0.7, 0.3], [0.4, 0.9], [0.2, 0.2], mutation_scale=20, lw=3, arrowstyle="-|>", color="k") ax.add_artist(axis_arrow) plt.tight_layout() plt.show() .. raw:: html | .. plot:: :width: 400px import numpy as np import matplotlib.pyplot as plt from mpl_toolkits.mplot3d import proj3d from pytransform3d.transformations import transform, plot_transform from pytransform3d.plot_utils import make_3d_axis, Arrow3D plt.figure() ax = make_3d_axis(1) plt.setp(ax, xlim=(-1.05, 1.05), ylim=(-0.55, 1.55), zlim=(-1.05, 1.05), xlabel="X", ylabel="Y", zlabel="Z") ax.view_init(elev=90, azim=-90) ax.set_xticks(()) ax.set_yticks(()) ax.set_zticks(()) rng = np.random.default_rng(42) PA = np.ones((10, 4)) PA[:, :3] = 0.1 * rng.standard_normal(size=(10, 3)) PA[:, 0] += 0.3 PA[:, :3] += 0.3 x_translation = -0.1 y_translation = 0.2 z_rotation = np.pi / 4.0 A2B = np.array([ [np.cos(z_rotation), -np.sin(z_rotation), 0.0, x_translation], [np.sin(z_rotation), np.cos(z_rotation), 0.0, y_translation], [0.0, 0.0, 1.0, 0.0], [0.0, 0.0, 0.0, 1.0] ]) plot_transform(ax=ax, A2B=np.eye(4), ls="--", alpha=0.5) ax.scatter(PA[:, 0], PA[:, 1], PA[:, 2], c="orange") plot_transform(ax=ax, A2B=A2B) axis_arrow = Arrow3D( [0.0, -0.1], [0.0, 0.2], [0.2, 0.2], mutation_scale=20, lw=3, arrowstyle="-|>", color="k") ax.add_artist(axis_arrow) plt.tight_layout() plt.show() .. raw:: html |