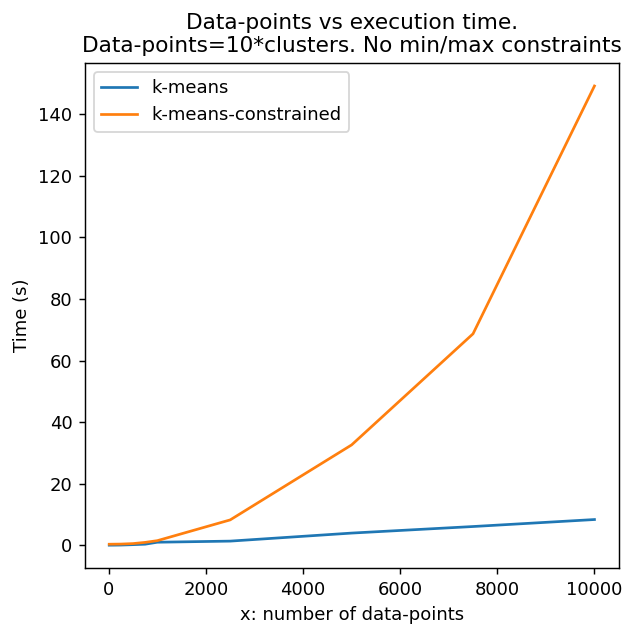

"""k-means-constrained"""
# Authors: Josh Levy-Kramer <josh@levykramer.co.uk>
# Gael Varoquaux <gael.varoquaux@normalesup.org>
# Thomas Rueckstiess <ruecksti@in.tum.de>
# James Bergstra <james.bergstra@umontreal.ca>
# Jan Schlueter <scikit-learn@jan-schlueter.de>
# Nelle Varoquaux
# Peter Prettenhofer <peter.prettenhofer@gmail.com>
# Olivier Grisel <olivier.grisel@ensta.org>
# Mathieu Blondel <mathieu@mblondel.org>
# Robert Layton <robertlayton@gmail.com>
# License: BSD 3 clause
import warnings
import numpy as np
import scipy.sparse as sp
from .sklearn_import.metrics.pairwise import euclidean_distances
from .sklearn_import.utils.extmath import row_norms, squared_norm, cartesian
from .sklearn_import.utils.validation import check_array, check_random_state, as_float_array, check_is_fitted
from joblib import Parallel
from joblib import delayed
# Internal scikit learn methods imported into this project
from k_means_constrained.sklearn_import.cluster._k_means import _centers_dense, _centers_sparse
from k_means_constrained.sklearn_import.cluster.k_means_ import _validate_center_shape, _tolerance, KMeans, \
_init_centroids
from k_means_constrained.mincostflow_vectorized import SimpleMinCostFlowVectorized
def k_means_constrained(X, n_clusters, size_min=None, size_max=None, init='k-means++',
n_init=10, max_iter=300, verbose=False,
tol=1e-4, random_state=None, copy_x=True, n_jobs=1,
return_n_iter=False):
"""K-Means clustering with minimum and maximum cluster size constraints.
Read more in the :ref:`User Guide <k_means>`.
Parameters
----------
X : array-like, shape (n_samples, n_features)
The observations to cluster.
size_min : int, optional, default: None
Constrain the label assignment so that each cluster has a minimum
size of size_min. If None, no constrains will be applied
size_max : int, optional, default: None
Constrain the label assignment so that each cluster has a maximum
size of size_max. If None, no constrains will be applied
n_clusters : int
The number of clusters to form as well as the number of
centroids to generate.
init : {'k-means++', 'random', or ndarray, or a callable}, optional
Method for initialization, default to 'k-means++':
'k-means++' : selects initial cluster centers for k-mean
clustering in a smart way to speed up convergence. See section
Notes in k_init for more details.
'random': generate k centroids from a Gaussian with mean and
variance estimated from the data.
If an ndarray is passed, it should be of shape (n_clusters, n_features)
and gives the initial centers.
If a callable is passed, it should take arguments X, k and
and a random state and return an initialization.
n_init : int, optional, default: 10
Number of time the k-means algorithm will be run with different
centroid seeds. The final results will be the best output of
n_init consecutive runs in terms of inertia.
max_iter : int, optional, default 300
Maximum number of iterations of the k-means algorithm to run.
verbose : boolean, optional
Verbosity mode.
tol : float, optional
Relative tolerance with regards to Frobenius norm of the difference
in the cluster centers of two consecutive iterations to declare
convergence.
random_state : int, RandomState instance or None, optional, default: None
If int, random_state is the seed used by the random number generator;
If RandomState instance, random_state is the random number generator;
If None, the random number generator is the RandomState instance used
by `np.random`.
copy_x : boolean, optional
When pre-computing distances it is more numerically accurate to center
the data first. If copy_x is True, then the original data is not
modified. If False, the original data is modified, and put back before
the function returns, but small numerical differences may be introduced
by subtracting and then adding the data mean.
n_jobs : int
The number of jobs to use for the computation. This works by computing
each of the n_init runs in parallel.
If -1 all CPUs are used. If 1 is given, no parallel computing code is
used at all, which is useful for debugging. For n_jobs below -1,
(n_cpus + 1 + n_jobs) are used. Thus for n_jobs = -2, all CPUs but one
are used.
return_n_iter : bool, optional
Whether or not to return the number of iterations.
Returns
-------
centroid : float ndarray with shape (k, n_features)
Centroids found at the last iteration of k-means.
label : integer ndarray with shape (n_samples,)
label[i] is the code or index of the centroid the
i'th observation is closest to.
inertia : float
The final value of the inertia criterion (sum of squared distances to
the closest centroid for all observations in the training set).
best_n_iter : int
Number of iterations corresponding to the best results.
Returned only if `return_n_iter` is set to True.
"""
if sp.issparse(X):
raise NotImplementedError("Not implemented for sparse X")
if n_init <= 0:
raise ValueError("Invalid number of initializations."
" n_init=%d must be bigger than zero." % n_init)
random_state = check_random_state(random_state)
if max_iter <= 0:
raise ValueError('Number of iterations should be a positive number,'
' got %d instead' % max_iter)
X = as_float_array(X, copy=copy_x)
tol = _tolerance(X, tol)
# Validate init array
if hasattr(init, '__array__'):
init = check_array(init, dtype=X.dtype.type, copy=True)
_validate_center_shape(X, n_clusters, init)
if n_init != 1:
warnings.warn(
'Explicit initial center position passed: '
'performing only one init in k-means instead of n_init=%d'
% n_init, RuntimeWarning, stacklevel=2)
n_init = 1
# subtract of mean of x for more accurate distance computations
if not sp.issparse(X):
X_mean = X.mean(axis=0)
# The copy was already done above
X -= X_mean
if hasattr(init, '__array__'):
init -= X_mean
# precompute squared norms of data points
x_squared_norms = row_norms(X, squared=True)
best_labels, best_inertia, best_centers = None, None, None
if n_jobs == 1:
# For a single thread, less memory is needed if we just store one set
# of the best results (as opposed to one set per run per thread).
for it in range(n_init):
# run a k-means once
labels, inertia, centers, n_iter_ = kmeans_constrained_single(
X, n_clusters,
size_min=size_min, size_max=size_max,
max_iter=max_iter, init=init, verbose=verbose, tol=tol,
x_squared_norms=x_squared_norms, random_state=random_state)
# determine if these results are the best so far
if best_inertia is None or inertia < best_inertia:
best_labels = labels.copy()
best_centers = centers.copy()
best_inertia = inertia
best_n_iter = n_iter_
else:
# parallelisation of k-means runs
seeds = random_state.randint(np.iinfo(np.int32).max, size=n_init)
results = Parallel(n_jobs=n_jobs, verbose=0)(
delayed(kmeans_constrained_single)(X, n_clusters,
size_min=size_min, size_max=size_max,
max_iter=max_iter, init=init,
verbose=verbose, tol=tol,
x_squared_norms=x_squared_norms,
# Change seed to ensure variety
random_state=seed)
for seed in seeds)
# Get results with the lowest inertia
labels, inertia, centers, n_iters = zip(*results)
best = np.argmin(inertia)
best_labels = labels[best]
best_inertia = inertia[best]
best_centers = centers[best]
best_n_iter = n_iters[best]
if not sp.issparse(X):
if not copy_x:
X += X_mean
best_centers += X_mean
if return_n_iter:
return best_centers, best_labels, best_inertia, best_n_iter
else:
return best_centers, best_labels, best_inertia
def kmeans_constrained_single(X, n_clusters, size_min=None, size_max=None,
max_iter=300, init='k-means++',
verbose=False, x_squared_norms=None,
random_state=None, tol=1e-4):
"""A single run of k-means constrained, assumes preparation completed prior.
Parameters
----------
X : array-like of floats, shape (n_samples, n_features)
The observations to cluster.
size_min : int, optional, default: None
Constrain the label assignment so that each cluster has a minimum
size of size_min. If None, no constrains will be applied
size_max : int, optional, default: None
Constrain the label assignment so that each cluster has a maximum
size of size_max. If None, no constrains will be applied
n_clusters : int
The number of clusters to form as well as the number of
centroids to generate.
max_iter : int, optional, default 300
Maximum number of iterations of the k-means algorithm to run.
init : {'k-means++', 'random', or ndarray, or a callable}, optional
Method for initialization, default to 'k-means++':
'k-means++' : selects initial cluster centers for k-mean
clustering in a smart way to speed up convergence. See section
Notes in k_init for more details.
'random': generate k centroids from a Gaussian with mean and
variance estimated from the data.
If an ndarray is passed, it should be of shape (k, p) and gives
the initial centers.
If a callable is passed, it should take arguments X, k and
and a random state and return an initialization.
tol : float, optional
Relative tolerance with regards to Frobenius norm of the difference
in the cluster centers of two consecutive iterations to declare
convergence.
verbose : boolean, optional
Verbosity mode
x_squared_norms : array
Precomputed x_squared_norms.
random_state : int, RandomState instance or None, optional, default: None
If int, random_state is the seed used by the random number generator;
If RandomState instance, random_state is the random number generator;
If None, the random number generator is the RandomState instance used
by `np.random`.
Returns
-------
centroid : float ndarray with shape (k, n_features)
Centroids found at the last iteration of k-means.
label : integer ndarray with shape (n_samples,)
label[i] is the code or index of the centroid the
i'th observation is closest to.
inertia : float
The final value of the inertia criterion (sum of squared distances to
the closest centroid for all observations in the training set).
n_iter : int
Number of iterations run.
"""
if sp.issparse(X):
raise NotImplementedError("Not implemented for sparse X")
random_state = check_random_state(random_state)
n_samples = X.shape[0]
best_labels, best_inertia, best_centers = None, None, None
# init
centers = _init_centroids(X, n_clusters, init, random_state=random_state, x_squared_norms=x_squared_norms)
if verbose:
print("Initialization complete")
# Allocate memory to store the distances for each sample to its
# closer center for reallocation in case of ties
distances = np.zeros(shape=(n_samples,), dtype=X.dtype)
# Determine min and max sizes if non given
if size_min is None:
size_min = 0
if size_max is None:
size_max = n_samples # Number of data points
# Check size min and max
if not ((size_min >= 0) and (size_min <= n_samples)
and (size_max >= 0) and (size_max <= n_samples)):
raise ValueError("size_min and size_max must be a positive number smaller "
"than the number of data points or `None`")
if size_max < size_min:
raise ValueError("size_max must be larger than size_min")
if size_min * n_clusters > n_samples:
raise ValueError("The product of size_min and n_clusters cannot exceed the number of samples (X)")
if size_max * n_clusters < n_samples:
raise ValueError("The product of size_max and n_clusters must be larger than or equal the number of samples (X)")
# iterations
for i in range(max_iter):
centers_old = centers.copy()
# labels assignment is also called the E-step of EM
labels, inertia = \
_labels_constrained(X, centers, size_min, size_max, distances=distances)
# computation of the means is also called the M-step of EM
if sp.issparse(X):
centers = _centers_sparse(X, labels, n_clusters, distances)
else:
centers = _centers_dense(X, labels, n_clusters, distances)
if verbose:
print("Iteration %2d, inertia %.3f" % (i, inertia))
if best_inertia is None or inertia < best_inertia:
best_labels = labels.copy()
best_centers = centers.copy()
best_inertia = inertia
center_shift_total = squared_norm(centers_old - centers)
if center_shift_total <= tol:
if verbose:
print("Converged at iteration %d: "
"center shift %e within tolerance %e"
% (i, center_shift_total, tol))
break
if center_shift_total > 0:
# rerun E-step in case of non-convergence so that predicted labels
# match cluster centers
best_labels, best_inertia = \
_labels_constrained(X, centers, size_min, size_max, distances=distances)
return best_labels, best_inertia, best_centers, i + 1
def _labels_constrained(X, centers, size_min, size_max, distances):
"""Compute labels using the min and max cluster size constraint
This will overwrite the 'distances' array in-place.
Parameters
----------
X : numpy array, shape (n_sample, n_features)
Input data.
size_min : int
Minimum size for each cluster
size_max : int
Maximum size for each cluster
centers : numpy array, shape (n_clusters, n_features)
Cluster centers which data is assigned to.
distances : numpy array, shape (n_samples,)
Pre-allocated array in which distances are stored.
Returns
-------
labels : numpy array, dtype=np.int, shape (n_samples,)
Indices of clusters that samples are assigned to.
inertia : float
Sum of squared distances of samples to their closest cluster center.
"""
C = centers
# Distances to each centre C. (the `distances` parameter is the distance to the closest centre)
# K-mean original uses squared distances but this equivalent for constrained k-means
D = euclidean_distances(X, C, squared=False)
edges, costs, capacities, supplies, n_C, n_X = minimum_cost_flow_problem_graph(X, C, D, size_min, size_max)
labels = solve_min_cost_flow_graph(edges, costs, capacities, supplies, n_C, n_X)
# cython k-means M step code assumes int32 inputs
labels = labels.astype(np.int32)
# Change distances in-place
distances[:] = D[np.arange(D.shape[0]), labels] ** 2 # Square for M step of EM
inertia = distances.sum()
return labels, inertia
def minimum_cost_flow_problem_graph(X, C, D, size_min, size_max):
# Setup minimum cost flow formulation graph
# Vertices indexes:
# X-nodes: [0, n(x)-1], C-nodes: [n(X), n(X)+n(C)-1], C-dummy nodes:[n(X)+n(C), n(X)+2*n(C)-1],
# Artificial node: [n(X)+2*n(C), n(X)+2*n(C)+1-1]
# Create indices of nodes
n_X = X.shape[0]
n_C = C.shape[0]
X_ix = np.arange(n_X)
C_dummy_ix = np.arange(X_ix[-1] + 1, X_ix[-1] + 1 + n_C)
C_ix = np.arange(C_dummy_ix[-1] + 1, C_dummy_ix[-1] + 1 + n_C)
art_ix = C_ix[-1] + 1
# Edges
edges_X_C_dummy = cartesian([X_ix, C_dummy_ix]) # All X's connect to all C dummy nodes (C')
edges_C_dummy_C = np.stack([C_dummy_ix, C_ix], axis=1) # Each C' connects to a corresponding C (centroid)
edges_C_art = np.stack([C_ix, art_ix * np.ones(n_C)], axis=1) # All C connect to artificial node
edges = np.concatenate([edges_X_C_dummy, edges_C_dummy_C, edges_C_art])
# Costs
costs_X_C_dummy = D.reshape(D.size)
costs = np.concatenate([costs_X_C_dummy, np.zeros(edges.shape[0] - len(costs_X_C_dummy))])
# Capacities - can set for max-k
capacities_C_dummy_C = size_max * np.ones(n_C)
cap_non = n_X # The total supply and therefore wont restrict flow
capacities = np.concatenate([
np.ones(edges_X_C_dummy.shape[0]),
capacities_C_dummy_C,
cap_non * np.ones(n_C)
])
# Sources and sinks
supplies_X = np.ones(n_X)
supplies_C = -1 * size_min * np.ones(n_C) # Demand node
supplies_art = -1 * (n_X - n_C * size_min) # Demand node
supplies = np.concatenate([
supplies_X,
np.zeros(n_C), # C_dummies
supplies_C,
[supplies_art]
])
# All arrays must be of int dtype for `SimpleMinCostFlow`
edges = edges.astype('int32')
costs = np.around(costs * 1000, 0).astype('int32') # Times by 1000 to give extra precision
capacities = capacities.astype('int32')
supplies = supplies.astype('int32')
return edges, costs, capacities, supplies, n_C, n_X
def solve_min_cost_flow_graph(edges, costs, capacities, supplies, n_C, n_X):
# Instantiate a SimpleMinCostFlow solver.
min_cost_flow = SimpleMinCostFlowVectorized()
if (edges.dtype != 'int32') or (costs.dtype != 'int32') \
or (capacities.dtype != 'int32') or (supplies.dtype != 'int32'):
raise ValueError("`edges`, `costs`, `capacities`, `supplies` must all be int dtype")
N_edges = edges.shape[0]
N_nodes = len(supplies)
# Add each edge with associated capacities and cost
min_cost_flow.AddArcWithCapacityAndUnitCostVectorized(edges[:, 0], edges[:, 1], capacities, costs)
# Add node supplies
min_cost_flow.SetNodeSupplyVectorized(np.arange(N_nodes, dtype='int32'), supplies)
# Find the minimum cost flow between node 0 and node 4.
if min_cost_flow.Solve() != min_cost_flow.OPTIMAL:
raise Exception('There was an issue with the min cost flow input.')
# Assignment
labels_M = min_cost_flow.FlowVectorized(np.arange(n_X * n_C, dtype='int32')).reshape(n_X, n_C)
labels = labels_M.argmax(axis=1)
return labels
[docs]class KMeansConstrained(KMeans):
"""K-Means clustering with minimum and maximum cluster size constraints
Parameters
----------
n_clusters : int, optional, default: 8
The number of clusters to form as well as the number of
centroids to generate.
size_min : int, optional, default: None
Constrain the label assignment so that each cluster has a minimum
size of size_min. If None, no constrains will be applied
size_max : int, optional, default: None
Constrain the label assignment so that each cluster has a maximum
size of size_max. If None, no constrains will be applied
init : {'k-means++', 'random' or an ndarray}
Method for initialization, defaults to 'k-means++':
'k-means++' : selects initial cluster centers for k-mean
clustering in a smart way to speed up convergence. See section
Notes in k_init for more details.
'random': choose k observations (rows) at random from data for
the initial centroids.
If an ndarray is passed, it should be of shape (n_clusters, n_features)
and gives the initial centers.
n_init : int, default: 10
Number of times the k-means algorithm will be run with different
centroid seeds. The final results will be the best output of
n_init consecutive runs in terms of inertia.
max_iter : int, default: 300
Maximum number of iterations of the k-means algorithm for a
single run.
tol : float, default: 1e-4
Relative tolerance with regards to Frobenius norm of the difference
in the cluster centers of two consecutive iterations to declare
convergence.
verbose : int, default 0
Verbosity mode.
random_state : int, RandomState instance or None, optional, default: None
If int, random_state is the seed used by the random number generator;
If RandomState instance, random_state is the random number generator;
If None, the random number generator is the RandomState instance used
by `np.random`.
copy_x : boolean, default True
When pre-computing distances it is more numerically accurate to center
the data first. If copy_x is True, then the original data is not
modified. If False, the original data is modified, and put back before
the function returns, but small numerical differences may be introduced
by subtracting and then adding the data mean.
n_jobs : int
The number of jobs to use for the computation. This works by computing
each of the n_init runs in parallel.
If -1 all CPUs are used. If 1 is given, no parallel computing code is
used at all, which is useful for debugging. For n_jobs below -1,
(n_cpus + 1 + n_jobs) are used. Thus for n_jobs = -2, all CPUs but one
are used.
Attributes
----------
cluster_centers_ : array, [n_clusters, n_features]
Coordinates of cluster centers
labels_ :
Labels of each point
inertia_ : float
Sum of squared distances of samples to their closest cluster center.
Examples
--------
>>> from k_means_constrained import KMeansConstrained
>>> import numpy as np
>>> X = np.array([[1, 2], [1, 4], [1, 0],
... [4, 2], [4, 4], [4, 0]])
>>> clf = KMeansConstrained(
... n_clusters=2,
... size_min=2,
... size_max=5,
... random_state=0
... )
>>> clf.fit_predict(X)
array([0, 0, 0, 1, 1, 1], dtype=int32)
>>> clf.cluster_centers_
array([[ 1., 2.],
[ 4., 2.]])
>>> clf.labels_
array([0, 0, 0, 1, 1, 1], dtype=int32)
Notes
------
K-means problem constrained with a minimum and/or maximum size for each cluster.
The constrained assignment is formulated as a Minimum Cost Flow (MCF) linear network optimisation
problem. This is then solved using a cost-scaling push-relabel algorithm. The implementation used is
Google's Operations Research tools's `SimpleMinCostFlow`.
Ref:
1. Bradley, P. S., K. P. Bennett, and Ayhan Demiriz. "Constrained k-means clustering."
Microsoft Research, Redmond (2000): 1-8.
2. Google's SimpleMinCostFlow implementation:
https://github.com/google/or-tools/blob/master/ortools/graph/min_cost_flow.h
"""
def __init__(self, n_clusters=8, size_min=None, size_max=None, init='k-means++', n_init=10, max_iter=300, tol=1e-4,
verbose=False, random_state=None, copy_x=True, n_jobs=1):
self.size_min = size_min
self.size_max = size_max
super().__init__(n_clusters=n_clusters, init=init, n_init=n_init, max_iter=max_iter, tol=tol,
verbose=verbose, random_state=random_state, copy_x=copy_x, n_jobs=n_jobs)
[docs] def fit(self, X, y=None):
"""Compute k-means clustering with given constants.
Parameters
----------
X : array-like, shape=(n_samples, n_features)
Training instances to cluster.
y : Ignored
"""
if sp.issparse(X):
raise NotImplementedError("Not implemented for sparse X")
random_state = check_random_state(self.random_state)
X = self._check_fit_data(X)
self.cluster_centers_, self.labels_, self.inertia_, self.n_iter_ = \
k_means_constrained(
X, n_clusters=self.n_clusters,
size_min=self.size_min, size_max=self.size_max,
init=self.init,
n_init=self.n_init, max_iter=self.max_iter, verbose=self.verbose,
tol=self.tol, random_state=random_state, copy_x=self.copy_x,
n_jobs=self.n_jobs,
return_n_iter=True)
return self
[docs] def predict(self, X, size_min='init', size_max='init'):
"""
Predict the closest cluster each sample in X belongs to given the provided constraints.
The constraints can be temporally overridden when determining which cluster each datapoint is assigned to.
Only computes the assignment step. It does not re-fit the cluster positions.
Parameters
----------
X : array-like, shape = [n_samples, n_features]
New data to predict.
size_min : int, optional, default: size_min provided with initialisation
Constrain the label assignment so that each cluster has a minimum
size of size_min. If None, no constrains will be applied.
If 'init' the value provided during initialisation of the
class will be used.
size_max : int, optional, default: size_max provided with initialisation
Constrain the label assignment so that each cluster has a maximum
size of size_max. If None, no constrains will be applied.
If 'init' the value provided during initialisation of the
class will be used.
Returns
-------
labels : array, shape [n_samples,]
Index of the cluster each sample belongs to.
"""
if sp.issparse(X):
raise NotImplementedError("Not implemented for sparse X")
if size_min == 'init':
size_min = self.size_min
if size_max == 'init':
size_max = self.size_max
n_clusters = self.n_clusters
n_samples = X.shape[0]
check_is_fitted(self, 'cluster_centers_')
X = self._check_test_data(X)
# Allocate memory to store the distances for each sample to its
# closer center for reallocation in case of ties
distances = np.zeros(shape=(n_samples,), dtype=X.dtype)
# Determine min and max sizes if non given
if size_min is None:
size_min = 0
if size_max is None:
size_max = n_samples # Number of data points
# Check size min and max
if not ((size_min >= 0) and (size_min <= n_samples)
and (size_max >= 0) and (size_max <= n_samples)):
raise ValueError("size_min and size_max must be a positive number smaller "
"than the number of data points or `None`")
if size_max < size_min:
raise ValueError("size_max must be larger than size_min")
if size_min * n_clusters > n_samples:
raise ValueError("The product of size_min and n_clusters cannot exceed the number of samples (X)")
labels, inertia = \
_labels_constrained(X, self.cluster_centers_, size_min, size_max, distances=distances)
return labels
[docs] def fit_predict(self, X, y=None):
"""Compute cluster centers and predict cluster index for each sample.
Equivalent to calling fit(X) followed by predict(X) but also more efficient.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to transform.
Returns
-------
labels : array, shape [n_samples,]
Index of the cluster each sample belongs to.
"""
return self.fit(X).labels_
"""K-means clustering"""
# Authors: Gael Varoquaux <gael.varoquaux@normalesup.org>
# Thomas Rueckstiess <ruecksti@in.tum.de>
# James Bergstra <james.bergstra@umontreal.ca>
# Jan Schlueter <scikit-learn@jan-schlueter.de>
# Nelle Varoquaux
# Peter Prettenhofer <peter.prettenhofer@gmail.com>
# Olivier Grisel <olivier.grisel@ensta.org>
# Mathieu Blondel <mathieu@mblondel.org>
# Robert Layton <robertlayton@gmail.com>
# License: BSD 3 clause
import warnings
import numpy as np
import scipy.sparse as sp
from sklearn.base import BaseEstimator, ClusterMixin, TransformerMixin
from sklearn.externals.six import string_types
from sklearn.metrics.pairwise import euclidean_distances
from sklearn.metrics.pairwise import pairwise_distances_argmin_min
from sklearn.utils import check_array
from sklearn.utils import check_random_state
from sklearn.utils.extmath import row_norms, stable_cumsum
from sklearn.utils.sparsefuncs import mean_variance_axis
from sklearn.utils.validation import FLOAT_DTYPES
from sklearn.utils.validation import check_is_fitted
from . import _k_means
###############################################################################
# Initialization heuristic
def _k_init(X, n_clusters, x_squared_norms, random_state, n_local_trials=None):
"""Init n_clusters seeds according to k-means++
Parameters
-----------
X : array or sparse matrix, shape (n_samples, n_features)
The data to pick seeds for. To avoid memory copy, the input data
should be double precision (dtype=np.float64).
n_clusters : integer
The number of seeds to choose
x_squared_norms : array, shape (n_samples,)
Squared Euclidean norm of each data point.
random_state : numpy.RandomState
The generator used to initialize the centers.
n_local_trials : integer, optional
The number of seeding trials for each center (except the first),
of which the one reducing inertia the most is greedily chosen.
Set to None to make the number of trials depend logarithmically
on the number of seeds (2+log(k)); this is the default.
Notes
-----
Selects initial cluster centers for k-mean clustering in a smart way
to speed up convergence. see: Arthur, D. and Vassilvitskii, S.
"k-means++: the advantages of careful seeding". ACM-SIAM symposium
on Discrete algorithms. 2007
Version ported from http://www.stanford.edu/~darthur/kMeansppTest.zip,
which is the implementation used in the aforementioned paper.
"""
n_samples, n_features = X.shape
centers = np.empty((n_clusters, n_features), dtype=X.dtype)
assert x_squared_norms is not None, 'x_squared_norms None in _k_init'
# Set the number of local seeding trials if none is given
if n_local_trials is None:
# This is what Arthur/Vassilvitskii tried, but did not report
# specific results for other than mentioning in the conclusion
# that it helped.
n_local_trials = 2 + int(np.log(n_clusters))
# Pick first center randomly
center_id = random_state.randint(n_samples)
if sp.issparse(X):
centers[0] = X[center_id].toarray()
else:
centers[0] = X[center_id]
# Initialize list of closest distances and calculate current potential
closest_dist_sq = euclidean_distances(
centers[0, np.newaxis], X, Y_norm_squared=x_squared_norms,
squared=True)
current_pot = closest_dist_sq.sum()
# Pick the remaining n_clusters-1 points
for c in range(1, n_clusters):
# Choose center candidates by sampling with probability proportional
# to the squared distance to the closest existing center
rand_vals = random_state.random_sample(n_local_trials) * current_pot
candidate_ids = np.searchsorted(stable_cumsum(closest_dist_sq),
rand_vals)
# Compute distances to center candidates
distance_to_candidates = euclidean_distances(
X[candidate_ids], X, Y_norm_squared=x_squared_norms, squared=True)
# Decide which candidate is the best
best_candidate = None
best_pot = None
best_dist_sq = None
for trial in range(n_local_trials):
# Compute potential when including center candidate
new_dist_sq = np.minimum(closest_dist_sq,
distance_to_candidates[trial])
new_pot = new_dist_sq.sum()
# Store result if it is the best local trial so far
if (best_candidate is None) or (new_pot < best_pot):
best_candidate = candidate_ids[trial]
best_pot = new_pot
best_dist_sq = new_dist_sq
# Permanently add best center candidate found in local tries
if sp.issparse(X):
centers[c] = X[best_candidate].toarray()
else:
centers[c] = X[best_candidate]
current_pot = best_pot
closest_dist_sq = best_dist_sq
return centers
###############################################################################
# K-means batch estimation by EM (expectation maximization)
def _validate_center_shape(X, n_centers, centers):
"""Check if centers is compatible with X and n_centers"""
if len(centers) != n_centers:
raise ValueError('The shape of the initial centers (%s) '
'does not match the number of clusters %i'
% (centers.shape, n_centers))
if centers.shape[1] != X.shape[1]:
raise ValueError(
"The number of features of the initial centers %s "
"does not match the number of features of the data %s."
% (centers.shape[1], X.shape[1]))
def _tolerance(X, tol):
"""Return a tolerance which is independent of the dataset"""
if sp.issparse(X):
variances = mean_variance_axis(X, axis=0)[1]
else:
variances = np.var(X, axis=0)
return np.mean(variances) * tol
def _labels_inertia_precompute_dense(X, x_squared_norms, centers, distances):
"""Compute labels and inertia using a full distance matrix.
This will overwrite the 'distances' array in-place.
Parameters
----------
X : numpy array, shape (n_sample, n_features)
Input data.
x_squared_norms : numpy array, shape (n_samples,)
Precomputed squared norms of X.
centers : numpy array, shape (n_clusters, n_features)
Cluster centers which data is assigned to.
distances : numpy array, shape (n_samples,)
Pre-allocated array in which distances are stored.
Returns
-------
labels : numpy array, dtype=np.int, shape (n_samples,)
Indices of clusters that samples are assigned to.
inertia : float
Sum of distances of samples to their closest cluster center.
"""
n_samples = X.shape[0]
# Breakup nearest neighbor distance computation into batches to prevent
# memory blowup in the case of a large number of samples and clusters.
# TODO: Once PR #7383 is merged use check_inputs=False in metric_kwargs.
labels, mindist = pairwise_distances_argmin_min(
X=X, Y=centers, metric='euclidean', metric_kwargs={'squared': True})
# cython k-means code assumes int32 inputs
labels = labels.astype(np.int32)
if n_samples == distances.shape[0]:
# distances will be changed in-place
distances[:] = mindist
inertia = mindist.sum()
return labels, inertia
def _labels_inertia(X, x_squared_norms, centers,
precompute_distances=True, distances=None):
"""E step of the K-means EM algorithm.
Compute the labels and the inertia of the given samples and centers.
This will compute the distances in-place.
Parameters
----------
X : float64 array-like or CSR sparse matrix, shape (n_samples, n_features)
The input samples to assign to the labels.
x_squared_norms : array, shape (n_samples,)
Precomputed squared euclidean norm of each data point, to speed up
computations.
centers : float array, shape (k, n_features)
The cluster centers.
precompute_distances : boolean, default: True
Precompute distances (faster but takes more memory).
distances : float array, shape (n_samples,)
Pre-allocated array to be filled in with each sample's distance
to the closest center.
Returns
-------
labels : int array of shape(n)
The resulting assignment
inertia : float
Sum of distances of samples to their closest cluster center.
"""
n_samples = X.shape[0]
# set the default value of centers to -1 to be able to detect any anomaly
# easily
labels = -np.ones(n_samples, np.int32)
if distances is None:
distances = np.zeros(shape=(0,), dtype=X.dtype)
# distances will be changed in-place
if sp.issparse(X):
inertia = _k_means._assign_labels_csr(
X, x_squared_norms, centers, labels, distances=distances)
else:
if precompute_distances:
return _labels_inertia_precompute_dense(X, x_squared_norms,
centers, distances)
inertia = _k_means._assign_labels_array(
X, x_squared_norms, centers, labels, distances=distances)
return labels, inertia
def _init_centroids(X, k, init, random_state=None, x_squared_norms=None,
init_size=None):
"""Compute the initial centroids
Parameters
----------
X : array, shape (n_samples, n_features)
k : int
number of centroids
init : {'k-means++', 'random' or ndarray or callable} optional
Method for initialization
random_state : int, RandomState instance or None, optional, default: None
If int, random_state is the seed used by the random number generator;
If RandomState instance, random_state is the random number generator;
If None, the random number generator is the RandomState instance used
by `np.random`.
x_squared_norms : array, shape (n_samples,), optional
Squared euclidean norm of each data point. Pass it if you have it at
hands already to avoid it being recomputed here. Default: None
init_size : int, optional
Number of samples to randomly sample for speeding up the
initialization (sometimes at the expense of accuracy): the
only algorithm is initialized by running a batch KMeans on a
random subset of the data. This needs to be larger than k.
Returns
-------
centers : array, shape(k, n_features)
"""
random_state = check_random_state(random_state)
n_samples = X.shape[0]
if x_squared_norms is None:
x_squared_norms = row_norms(X, squared=True)
if init_size is not None and init_size < n_samples:
if init_size < k:
warnings.warn(
"init_size=%d should be larger than k=%d. "
"Setting it to 3*k" % (init_size, k),
RuntimeWarning, stacklevel=2)
init_size = 3 * k
init_indices = random_state.randint(0, n_samples, init_size)
X = X[init_indices]
x_squared_norms = x_squared_norms[init_indices]
n_samples = X.shape[0]
elif n_samples < k:
raise ValueError(
"n_samples=%d should be larger than k=%d" % (n_samples, k))
if isinstance(init, string_types) and init == 'k-means++':
centers = _k_init(X, k, random_state=random_state,
x_squared_norms=x_squared_norms)
elif isinstance(init, string_types) and init == 'random':
seeds = random_state.permutation(n_samples)[:k]
centers = X[seeds]
elif hasattr(init, '__array__'):
# ensure that the centers have the same dtype as X
# this is a requirement of fused types of cython
centers = np.array(init, dtype=X.dtype)
elif callable(init):
centers = init(X, k, random_state=random_state)
centers = np.asarray(centers, dtype=X.dtype)
else:
raise ValueError("the init parameter for the k-means should "
"be 'k-means++' or 'random' or an ndarray, "
"'%s' (type '%s') was passed." % (init, type(init)))
if sp.issparse(centers):
centers = centers.toarray()
_validate_center_shape(X, k, centers)
return centers
class KMeans(BaseEstimator, ClusterMixin, TransformerMixin):
"""K-Means clustering
Read more in the :ref:`User Guide <k_means>`.
Parameters
----------
n_clusters : int, optional, default: 8
The number of clusters to form as well as the number of
centroids to generate.
init : {'k-means++', 'random' or an ndarray}
Method for initialization, defaults to 'k-means++':
'k-means++' : selects initial cluster centers for k-mean
clustering in a smart way to speed up convergence. See section
Notes in k_init for more details.
'random': choose k observations (rows) at random from data for
the initial centroids.
If an ndarray is passed, it should be of shape (n_clusters, n_features)
and gives the initial centers.
n_init : int, default: 10
Number of time the k-means algorithm will be run with different
centroid seeds. The final results will be the best output of
n_init consecutive runs in terms of inertia.
max_iter : int, default: 300
Maximum number of iterations of the k-means algorithm for a
single run.
tol : float, default: 1e-4
Relative tolerance with regards to inertia to declare convergence
precompute_distances : {'auto', True, False}
Precompute distances (faster but takes more memory).
'auto' : do not precompute distances if n_samples * n_clusters > 12
million. This corresponds to about 100MB overhead per job using
double precision.
True : always precompute distances
False : never precompute distances
verbose : int, default 0
Verbosity mode.
random_state : int, RandomState instance or None, optional, default: None
If int, random_state is the seed used by the random number generator;
If RandomState instance, random_state is the random number generator;
If None, the random number generator is the RandomState instance used
by `np.random`.
copy_x : boolean, default True
When pre-computing distances it is more numerically accurate to center
the data first. If copy_x is True, then the original data is not
modified. If False, the original data is modified, and put back before
the function returns, but small numerical differences may be introduced
by subtracting and then adding the data mean.
n_jobs : int
The number of jobs to use for the computation. This works by computing
each of the n_init runs in parallel.
If -1 all CPUs are used. If 1 is given, no parallel computing code is
used at all, which is useful for debugging. For n_jobs below -1,
(n_cpus + 1 + n_jobs) are used. Thus for n_jobs = -2, all CPUs but one
are used.
algorithm : "auto", "full" or "elkan", default="auto"
K-means algorithm to use. The classical EM-style algorithm is "full".
The "elkan" variation is more efficient by using the triangle
inequality, but currently doesn't support sparse data. "auto" chooses
"elkan" for dense data and "full" for sparse data.
Attributes
----------
cluster_centers_ : array, [n_clusters, n_features]
Coordinates of cluster centers
labels_ :
Labels of each point
inertia_ : float
Sum of distances of samples to their closest cluster center.
Examples
--------
>>> from sklearn.cluster import KMeans
>>> import numpy as np
>>> X = np.array([[1, 2], [1, 4], [1, 0],
... [4, 2], [4, 4], [4, 0]])
>>> kmeans = KMeans(n_clusters=2, random_state=0).fit(X)
>>> kmeans.labels_
array([0, 0, 0, 1, 1, 1], dtype=int32)
>>> kmeans.predict([[0, 0], [4, 4]])
array([0, 1], dtype=int32)
>>> kmeans.cluster_centers_
array([[ 1., 2.],
[ 4., 2.]])
See also
--------
MiniBatchKMeans
Alternative online implementation that does incremental updates
of the centers positions using mini-batches.
For large scale learning (say n_samples > 10k) MiniBatchKMeans is
probably much faster than the default batch implementation.
Notes
------
The k-means problem is solved using Lloyd's algorithm.
The average complexity is given by O(k n T), were n is the number of
samples and T is the number of iteration.
The worst case complexity is given by O(n^(k+2/p)) with
n = n_samples, p = n_features. (D. Arthur and S. Vassilvitskii,
'How slow is the k-means method?' SoCG2006)
In practice, the k-means algorithm is very fast (one of the fastest
clustering algorithms available), but it falls in local minima. That's why
it can be useful to restart it several times.
"""
def __init__(self, n_clusters=8, init='k-means++', n_init=10,
max_iter=300, tol=1e-4, precompute_distances='auto',
verbose=0, random_state=None, copy_x=True,
n_jobs=1, algorithm='auto'):
self.n_clusters = n_clusters
self.init = init
self.max_iter = max_iter
self.tol = tol
self.precompute_distances = precompute_distances
self.n_init = n_init
self.verbose = verbose
self.random_state = random_state
self.copy_x = copy_x
self.n_jobs = n_jobs
self.algorithm = algorithm
def _check_fit_data(self, X):
"""Verify that the number of samples given is larger than k"""
X = check_array(X, accept_sparse='csr', dtype=[np.float64, np.float32])
if X.shape[0] < self.n_clusters:
raise ValueError("n_samples=%d should be >= n_clusters=%d" % (
X.shape[0], self.n_clusters))
return X
def _check_test_data(self, X):
X = check_array(X, accept_sparse='csr', dtype=FLOAT_DTYPES)
n_samples, n_features = X.shape
expected_n_features = self.cluster_centers_.shape[1]
if not n_features == expected_n_features:
raise ValueError("Incorrect number of features. "
"Got %d features, expected %d" % (
n_features, expected_n_features))
return X
def fit(self, X, y=None):
"""Compute k-means clustering.
Parameters
----------
X : array-like or sparse matrix, shape=(n_samples, n_features)
Training instances to cluster.
"""
# Added to remove scikit-learn internal dependenceies
raise NotImplemented
def fit_predict(self, X, y=None):
"""Compute cluster centers and predict cluster index for each sample.
Convenience method; equivalent to calling fit(X) followed by
predict(X).
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to transform.
Returns
-------
labels : array, shape [n_samples,]
Index of the cluster each sample belongs to.
"""
return self.fit(X).labels_
def fit_transform(self, X, y=None):
"""Compute clustering and transform X to cluster-distance space.
Equivalent to fit(X).transform(X), but more efficiently implemented.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to transform.
Returns
-------
X_new : array, shape [n_samples, k]
X transformed in the new space.
"""
# Currently, this just skips a copy of the data if it is not in
# np.array or CSR format already.
# XXX This skips _check_test_data, which may change the dtype;
# we should refactor the input validation.
X = self._check_fit_data(X)
return self.fit(X)._transform(X)
def transform(self, X):
"""Transform X to a cluster-distance space.
In the new space, each dimension is the distance to the cluster
centers. Note that even if X is sparse, the array returned by
`transform` will typically be dense.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to transform.
Returns
-------
X_new : array, shape [n_samples, k]
X transformed in the new space.
"""
check_is_fitted(self, 'cluster_centers_')
X = self._check_test_data(X)
return self._transform(X)
def _transform(self, X):
"""guts of transform method; no input validation"""
return euclidean_distances(X, self.cluster_centers_)
def predict(self, X):
"""Predict the closest cluster each sample in X belongs to.
In the vector quantization literature, `cluster_centers_` is called
the code book and each value returned by `predict` is the index of
the closest code in the code book.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to predict.
Returns
-------
labels : array, shape [n_samples,]
Index of the cluster each sample belongs to.
"""
check_is_fitted(self, 'cluster_centers_')
X = self._check_test_data(X)
x_squared_norms = row_norms(X, squared=True)
return _labels_inertia(X, x_squared_norms, self.cluster_centers_)[0]
def score(self, X, y=None):
"""Opposite of the value of X on the K-means objective.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data.
Returns
-------
score : float
Opposite of the value of X on the K-means objective.
"""
check_is_fitted(self, 'cluster_centers_')
X = self._check_test_data(X)
x_squared_norms = row_norms(X, squared=True)
return -_labels_inertia(X, x_squared_norms, self.cluster_centers_)[1]
import warnings
from collections import defaultdict
import numpy as np
import six
from k_means_constrained.sklearn_import import __version__
from k_means_constrained.sklearn_import.funcsigs import signature
class BaseEstimator(object):
"""Base class for all estimators in scikit-learn
Notes
-----
All estimators should specify all the parameters that can be set
at the class level in their ``__init__`` as explicit keyword
arguments (no ``*args`` or ``**kwargs``).
"""
@classmethod
def _get_param_names(cls):
"""Get parameter names for the estimator"""
# fetch the constructor or the original constructor before
# deprecation wrapping if any
init = getattr(cls.__init__, 'deprecated_original', cls.__init__)
if init is object.__init__:
# No explicit constructor to introspect
return []
# introspect the constructor arguments to find the model parameters
# to represent
init_signature = signature(init)
# Consider the constructor parameters excluding 'self'
parameters = [p for p in init_signature.parameters.values()
if p.name != 'self' and p.kind != p.VAR_KEYWORD]
for p in parameters:
if p.kind == p.VAR_POSITIONAL:
raise RuntimeError("scikit-learn estimators should always "
"specify their parameters in the signature"
" of their __init__ (no varargs)."
" %s with constructor %s doesn't "
" follow this convention."
% (cls, init_signature))
# Extract and sort argument names excluding 'self'
return sorted([p.name for p in parameters])
def get_params(self, deep=True):
"""Get parameters for this estimator.
Parameters
----------
deep : boolean, optional
If True, will return the parameters for this estimator and
contained subobjects that are estimators.
Returns
-------
params : mapping of string to any
Parameter names mapped to their values.
"""
out = dict()
for key in self._get_param_names():
# We need deprecation warnings to always be on in order to
# catch deprecated param values.
# This is set in utils/__init__.py but it gets overwritten
# when running under python3 somehow.
warnings.simplefilter("always", DeprecationWarning)
try:
with warnings.catch_warnings(record=True) as w:
value = getattr(self, key, None)
if len(w) and w[0].category == DeprecationWarning:
# if the parameter is deprecated, don't show it
continue
finally:
warnings.filters.pop(0)
# XXX: should we rather test if instance of estimator?
if deep and hasattr(value, 'get_params'):
deep_items = value.get_params().items()
out.update((key + '__' + k, val) for k, val in deep_items)
out[key] = value
return out
def set_params(self, **params):
"""Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects
(such as pipelines). The latter have parameters of the form
``<component>__<parameter>`` so that it's possible to update each
component of a nested object.
Returns
-------
self
"""
if not params:
# Simple optimization to gain speed (inspect is slow)
return self
valid_params = self.get_params(deep=True)
nested_params = defaultdict(dict) # grouped by prefix
for key, value in params.items():
key, delim, sub_key = key.partition('__')
if key not in valid_params:
raise ValueError('Invalid parameter %s for estimator %s. '
'Check the list of available parameters '
'with `estimator.get_params().keys()`.' %
(key, self))
if delim:
nested_params[key][sub_key] = value
else:
setattr(self, key, value)
for key, sub_params in nested_params.items():
valid_params[key].set_params(**sub_params)
return self
def __repr__(self):
class_name = self.__class__.__name__
return '%s(%s)' % (class_name, _pprint(self.get_params(deep=False),
offset=len(class_name),),)
def __getstate__(self):
try:
state = super(BaseEstimator, self).__getstate__()
except AttributeError:
state = self.__dict__.copy()
if type(self).__module__.startswith('sklearn.'):
return dict(state.items(), _sklearn_version=__version__)
else:
return state
def __setstate__(self, state):
if type(self).__module__.startswith('sklearn.'):
pickle_version = state.pop("_sklearn_version", "pre-0.18")
if pickle_version != __version__:
warnings.warn(
"Trying to unpickle estimator {0} from version {1} when "
"using version {2}. This might lead to breaking code or "
"invalid results. Use at your own risk.".format(
self.__class__.__name__, pickle_version, __version__),
UserWarning)
try:
super(BaseEstimator, self).__setstate__(state)
except AttributeError:
self.__dict__.update(state)
class ClusterMixin(object):
"""Mixin class for all cluster estimators in scikit-learn."""
_estimator_type = "clusterer"
def fit_predict(self, X, y=None):
"""Performs clustering on X and returns cluster labels.
Parameters
----------
X : ndarray, shape (n_samples, n_features)
Input data.
Returns
-------
y : ndarray, shape (n_samples,)
cluster labels
"""
# non-optimized default implementation; override when a better
# method is possible for a given clustering algorithm
self.fit(X)
return self.labels_
class TransformerMixin(object):
"""Mixin class for all transformers in scikit-learn."""
def fit_transform(self, X, y=None, **fit_params):
"""Fit to data, then transform it.
Fits transformer to X and y with optional parameters fit_params
and returns a transformed version of X.
Parameters
----------
X : numpy array of shape [n_samples, n_features]
Training set.
y : numpy array of shape [n_samples]
Target values.
Returns
-------
X_new : numpy array of shape [n_samples, n_features_new]
Transformed array.
"""
# non-optimized default implementation; override when a better
# method is possible for a given clustering algorithm
if y is None:
# fit method of arity 1 (unsupervised transformation)
return self.fit(X, **fit_params).transform(X)
else:
# fit method of arity 2 (supervised transformation)
return self.fit(X, y, **fit_params).transform(X)
def _pprint(params, offset=0, printer=repr):
"""Pretty print the dictionary 'params'
Parameters
----------
params : dict
The dictionary to pretty print
offset : int
The offset in characters to add at the begin of each line.
printer : callable
The function to convert entries to strings, typically
the builtin str or repr
"""
# Do a multi-line justified repr:
options = np.get_printoptions()
np.set_printoptions(precision=5, threshold=64, edgeitems=2)
params_list = list()
this_line_length = offset
line_sep = ',\n' + (1 + offset // 2) * ' '
for i, (k, v) in enumerate(sorted(six.iteritems(params))):
if type(v) is float:
# use str for representing floating point numbers
# this way we get consistent representation across
# architectures and versions.
this_repr = '%s=%s' % (k, str(v))
else:
# use repr of the rest
this_repr = '%s=%s' % (k, printer(v))
if len(this_repr) > 500:
this_repr = this_repr[:300] + '...' + this_repr[-100:]
if i > 0:
if (this_line_length + len(this_repr) >= 75 or '\n' in this_repr):
params_list.append(line_sep)
this_line_length = len(line_sep)
else:
params_list.append(', ')
this_line_length += 2
params_list.append(this_repr)
this_line_length += len(this_repr)
np.set_printoptions(**options)
lines = ''.join(params_list)
# Strip trailing space to avoid nightmare in doctests
lines = '\n'.join(l.rstrip(' ') for l in lines.split('\n'))
return lines
"""K-means clustering"""
# Authors: Gael Varoquaux <gael.varoquaux@normalesup.org>
# Thomas Rueckstiess <ruecksti@in.tum.de>
# James Bergstra <james.bergstra@umontreal.ca>
# Jan Schlueter <scikit-learn@jan-schlueter.de>
# Nelle Varoquaux
# Peter Prettenhofer <peter.prettenhofer@gmail.com>
# Olivier Grisel <olivier.grisel@ensta.org>
# Mathieu Blondel <mathieu@mblondel.org>
# Robert Layton <robertlayton@gmail.com>
# License: BSD 3 clause
import warnings
import numpy as np
import scipy.sparse as sp
from k_means_constrained.sklearn_import.base import BaseEstimator, ClusterMixin, TransformerMixin
from six import string_types
from k_means_constrained.sklearn_import.metrics.pairwise import euclidean_distances, pairwise_distances_argmin_min
from k_means_constrained.sklearn_import.utils.validation import check_array, check_random_state, FLOAT_DTYPES, \
check_is_fitted
from k_means_constrained.sklearn_import.utils.extmath import row_norms, stable_cumsum
from k_means_constrained.sklearn_import.utils.sparsefuncs import mean_variance_axis
from k_means_constrained.sklearn_import.cluster import _k_means
###############################################################################
# Initialization heuristic
def _k_init(X, n_clusters, x_squared_norms, random_state, n_local_trials=None):
"""Init n_clusters seeds according to k-means++
Parameters
-----------
X : array or sparse matrix, shape (n_samples, n_features)
The data to pick seeds for. To avoid memory copy, the input data
should be double precision (dtype=np.float64).
n_clusters : integer
The number of seeds to choose
x_squared_norms : array, shape (n_samples,)
Squared Euclidean norm of each data point.
random_state : numpy.RandomState
The generator used to initialize the centers.
n_local_trials : integer, optional
The number of seeding trials for each center (except the first),
of which the one reducing inertia the most is greedily chosen.
Set to None to make the number of trials depend logarithmically
on the number of seeds (2+log(k)); this is the default.
Notes
-----
Selects initial cluster centers for k-mean clustering in a smart way
to speed up convergence. see: Arthur, D. and Vassilvitskii, S.
"k-means++: the advantages of careful seeding". ACM-SIAM symposium
on Discrete algorithms. 2007
Version ported from http://www.stanford.edu/~darthur/kMeansppTest.zip,
which is the implementation used in the aforementioned paper.
"""
n_samples, n_features = X.shape
centers = np.empty((n_clusters, n_features), dtype=X.dtype)
assert x_squared_norms is not None, 'x_squared_norms None in _k_init'
# Set the number of local seeding trials if none is given
if n_local_trials is None:
# This is what Arthur/Vassilvitskii tried, but did not report
# specific results for other than mentioning in the conclusion
# that it helped.
n_local_trials = 2 + int(np.log(n_clusters))
# Pick first center randomly
center_id = random_state.randint(n_samples)
if sp.issparse(X):
centers[0] = X[center_id].toarray()
else:
centers[0] = X[center_id]
# Initialize list of closest distances and calculate current potential
closest_dist_sq = euclidean_distances(
centers[0, np.newaxis], X, Y_norm_squared=x_squared_norms,
squared=True)
current_pot = closest_dist_sq.sum()
# Pick the remaining n_clusters-1 points
for c in range(1, n_clusters):
# Choose center candidates by sampling with probability proportional
# to the squared distance to the closest existing center
rand_vals = random_state.random_sample(n_local_trials) * current_pot
candidate_ids = np.searchsorted(stable_cumsum(closest_dist_sq),
rand_vals)
# Compute distances to center candidates
distance_to_candidates = euclidean_distances(
X[candidate_ids], X, Y_norm_squared=x_squared_norms, squared=True)
# Decide which candidate is the best
best_candidate = None
best_pot = None
best_dist_sq = None
for trial in range(n_local_trials):
# Compute potential when including center candidate
new_dist_sq = np.minimum(closest_dist_sq,
distance_to_candidates[trial])
new_pot = new_dist_sq.sum()
# Store result if it is the best local trial so far
if (best_candidate is None) or (new_pot < best_pot):
best_candidate = candidate_ids[trial]
best_pot = new_pot
best_dist_sq = new_dist_sq
# Permanently add best center candidate found in local tries
if sp.issparse(X):
centers[c] = X[best_candidate].toarray()
else:
centers[c] = X[best_candidate]
current_pot = best_pot
closest_dist_sq = best_dist_sq
return centers
###############################################################################
# K-means batch estimation by EM (expectation maximization)
def _validate_center_shape(X, n_centers, centers):
"""Check if centers is compatible with X and n_centers"""
if len(centers) != n_centers:
raise ValueError('The shape of the initial centers (%s) '
'does not match the number of clusters %i'
% (centers.shape, n_centers))
if centers.shape[1] != X.shape[1]:
raise ValueError(
"The number of features of the initial centers %s "
"does not match the number of features of the data %s."
% (centers.shape[1], X.shape[1]))
def _tolerance(X, tol):
"""Return a tolerance which is independent of the dataset"""
if sp.issparse(X):
variances = mean_variance_axis(X, axis=0)[1]
else:
variances = np.var(X, axis=0)
return np.mean(variances) * tol
def _labels_inertia_precompute_dense(X, x_squared_norms, centers, distances):
"""Compute labels and inertia using a full distance matrix.
This will overwrite the 'distances' array in-place.
Parameters
----------
X : numpy array, shape (n_sample, n_features)
Input data.
x_squared_norms : numpy array, shape (n_samples,)
Precomputed squared norms of X.
centers : numpy array, shape (n_clusters, n_features)
Cluster centers which data is assigned to.
distances : numpy array, shape (n_samples,)
Pre-allocated array in which distances are stored.
Returns
-------
labels : numpy array, dtype=np.int, shape (n_samples,)
Indices of clusters that samples are assigned to.
inertia : float
Sum of distances of samples to their closest cluster center.
"""
n_samples = X.shape[0]
# Breakup nearest neighbor distance computation into batches to prevent
# memory blowup in the case of a large number of samples and clusters.
# TODO: Once PR #7383 is merged use check_inputs=False in metric_kwargs.
labels, mindist = pairwise_distances_argmin_min(
X=X, Y=centers, metric='euclidean', metric_kwargs={'squared': True})
# cython k-means code assumes int32 inputs
labels = labels.astype(np.int32)
if n_samples == distances.shape[0]:
# distances will be changed in-place
distances[:] = mindist
inertia = mindist.sum()
return labels, inertia
def _labels_inertia(X, x_squared_norms, centers,
precompute_distances=True, distances=None):
"""E step of the K-means EM algorithm.
Compute the labels and the inertia of the given samples and centers.
This will compute the distances in-place.
Parameters
----------
X : float64 array-like or CSR sparse matrix, shape (n_samples, n_features)
The input samples to assign to the labels.
x_squared_norms : array, shape (n_samples,)
Precomputed squared euclidean norm of each data point, to speed up
computations.
centers : float array, shape (k, n_features)
The cluster centers.
precompute_distances : boolean, default: True
Precompute distances (faster but takes more memory).
distances : float array, shape (n_samples,)
Pre-allocated array to be filled in with each sample's distance
to the closest center.
Returns
-------
labels : int array of shape(n)
The resulting assignment
inertia : float
Sum of distances of samples to their closest cluster center.
"""
n_samples = X.shape[0]
# set the default value of centers to -1 to be able to detect any anomaly
# easily
labels = -np.ones(n_samples, np.int32)
if distances is None:
distances = np.zeros(shape=(0,), dtype=X.dtype)
# distances will be changed in-place
if sp.issparse(X):
inertia = _k_means._assign_labels_csr(
X, x_squared_norms, centers, labels, distances=distances)
else:
if precompute_distances:
return _labels_inertia_precompute_dense(X, x_squared_norms,
centers, distances)
inertia = _k_means._assign_labels_array(
X, x_squared_norms, centers, labels, distances=distances)
return labels, inertia
def _init_centroids(X, k, init, random_state=None, x_squared_norms=None,
init_size=None):
"""Compute the initial centroids
Parameters
----------
X : array, shape (n_samples, n_features)
k : int
number of centroids
init : {'k-means++', 'random' or ndarray or callable} optional
Method for initialization
random_state : int, RandomState instance or None, optional, default: None
If int, random_state is the seed used by the random number generator;
If RandomState instance, random_state is the random number generator;
If None, the random number generator is the RandomState instance used
by `np.random`.
x_squared_norms : array, shape (n_samples,), optional
Squared euclidean norm of each data point. Pass it if you have it at
hands already to avoid it being recomputed here. Default: None
init_size : int, optional
Number of samples to randomly sample for speeding up the
initialization (sometimes at the expense of accuracy): the
only algorithm is initialized by running a batch KMeans on a
random subset of the data. This needs to be larger than k.
Returns
-------
centers : array, shape(k, n_features)
"""
random_state = check_random_state(random_state)
n_samples = X.shape[0]
if x_squared_norms is None:
x_squared_norms = row_norms(X, squared=True)
if init_size is not None and init_size < n_samples:
if init_size < k:
warnings.warn(
"init_size=%d should be larger than k=%d. "
"Setting it to 3*k" % (init_size, k),
RuntimeWarning, stacklevel=2)
init_size = 3 * k
init_indices = random_state.randint(0, n_samples, init_size)
X = X[init_indices]
x_squared_norms = x_squared_norms[init_indices]
n_samples = X.shape[0]
elif n_samples < k:
raise ValueError(
"n_samples=%d should be larger than k=%d" % (n_samples, k))
if isinstance(init, string_types) and init == 'k-means++':
centers = _k_init(X, k, random_state=random_state,
x_squared_norms=x_squared_norms)
elif isinstance(init, string_types) and init == 'random':
seeds = random_state.permutation(n_samples)[:k]
centers = X[seeds]
elif hasattr(init, '__array__'):
# ensure that the centers have the same dtype as X
# this is a requirement of fused types of cython
centers = np.array(init, dtype=X.dtype)
elif callable(init):
centers = init(X, k, random_state=random_state)
centers = np.asarray(centers, dtype=X.dtype)
else:
raise ValueError("the init parameter for the k-means should "
"be 'k-means++' or 'random' or an ndarray, "
"'%s' (type '%s') was passed." % (init, type(init)))
if sp.issparse(centers):
centers = centers.toarray()
_validate_center_shape(X, k, centers)
return centers
class KMeans(BaseEstimator, ClusterMixin, TransformerMixin):
"""K-Means clustering
Read more in the :ref:`User Guide <k_means>`.
Parameters
----------
n_clusters : int, optional, default: 8
The number of clusters to form as well as the number of
centroids to generate.
init : {'k-means++', 'random' or an ndarray}
Method for initialization, defaults to 'k-means++':
'k-means++' : selects initial cluster centers for k-mean
clustering in a smart way to speed up convergence. See section
Notes in k_init for more details.
'random': choose k observations (rows) at random from data for
the initial centroids.
If an ndarray is passed, it should be of shape (n_clusters, n_features)
and gives the initial centers.
n_init : int, default: 10
Number of time the k-means algorithm will be run with different
centroid seeds. The final results will be the best output of
n_init consecutive runs in terms of inertia.
max_iter : int, default: 300
Maximum number of iterations of the k-means algorithm for a
single run.
tol : float, default: 1e-4
Relative tolerance with regards to inertia to declare convergence
precompute_distances : {'auto', True, False}
Precompute distances (faster but takes more memory).
'auto' : do not precompute distances if n_samples * n_clusters > 12
million. This corresponds to about 100MB overhead per job using
double precision.
True : always precompute distances
False : never precompute distances
verbose : int, default 0
Verbosity mode.
random_state : int, RandomState instance or None, optional, default: None
If int, random_state is the seed used by the random number generator;
If RandomState instance, random_state is the random number generator;
If None, the random number generator is the RandomState instance used
by `np.random`.
copy_x : boolean, default True
When pre-computing distances it is more numerically accurate to center
the data first. If copy_x is True, then the original data is not
modified. If False, the original data is modified, and put back before
the function returns, but small numerical differences may be introduced
by subtracting and then adding the data mean.
n_jobs : int
The number of jobs to use for the computation. This works by computing
each of the n_init runs in parallel.
If -1 all CPUs are used. If 1 is given, no parallel computing code is
used at all, which is useful for debugging. For n_jobs below -1,
(n_cpus + 1 + n_jobs) are used. Thus for n_jobs = -2, all CPUs but one
are used.
algorithm : "auto", "full" or "elkan", default="auto"
K-means algorithm to use. The classical EM-style algorithm is "full".
The "elkan" variation is more efficient by using the triangle
inequality, but currently doesn't support sparse data. "auto" chooses
"elkan" for dense data and "full" for sparse data.
Attributes
----------
cluster_centers_ : array, [n_clusters, n_features]
Coordinates of cluster centers
labels_ :
Labels of each point
inertia_ : float
Sum of distances of samples to their closest cluster center.
Examples
--------
>>> from sklearn.cluster import KMeans
>>> import numpy as np
>>> X = np.array([[1, 2], [1, 4], [1, 0],
... [4, 2], [4, 4], [4, 0]])
>>> kmeans = KMeans(n_clusters=2, random_state=0).fit(X)
>>> kmeans.labels_
array([0, 0, 0, 1, 1, 1], dtype=int32)
>>> kmeans.predict([[0, 0], [4, 4]])
array([0, 1], dtype=int32)
>>> kmeans.cluster_centers_
array([[ 1., 2.],
[ 4., 2.]])
See also
--------
MiniBatchKMeans
Alternative online implementation that does incremental updates
of the centers positions using mini-batches.
For large scale learning (say n_samples > 10k) MiniBatchKMeans is
probably much faster than the default batch implementation.
Notes
------
The k-means problem is solved using Lloyd's algorithm.
The average complexity is given by O(k n T), were n is the number of
samples and T is the number of iteration.
The worst case complexity is given by O(n^(k+2/p)) with
n = n_samples, p = n_features. (D. Arthur and S. Vassilvitskii,
'How slow is the k-means method?' SoCG2006)
In practice, the k-means algorithm is very fast (one of the fastest
clustering algorithms available), but it falls in local minima. That's why
it can be useful to restart it several times.
"""
def __init__(self, n_clusters=8, init='k-means++', n_init=10,
max_iter=300, tol=1e-4, precompute_distances='auto',
verbose=0, random_state=None, copy_x=True,
n_jobs=1, algorithm='auto'):
self.n_clusters = n_clusters
self.init = init
self.max_iter = max_iter
self.tol = tol
self.precompute_distances = precompute_distances
self.n_init = n_init
self.verbose = verbose
self.random_state = random_state
self.copy_x = copy_x
self.n_jobs = n_jobs
self.algorithm = algorithm
def _check_fit_data(self, X):
"""Verify that the number of samples given is larger than k"""
X = check_array(X, accept_sparse='csr', dtype=[np.float64, np.float32])
if X.shape[0] < self.n_clusters:
raise ValueError("n_samples=%d should be >= n_clusters=%d" % (
X.shape[0], self.n_clusters))
return X
def _check_test_data(self, X):
X = check_array(X, accept_sparse='csr', dtype=FLOAT_DTYPES)
n_samples, n_features = X.shape
expected_n_features = self.cluster_centers_.shape[1]
if not n_features == expected_n_features:
raise ValueError("Incorrect number of features. "
"Got %d features, expected %d" % (
n_features, expected_n_features))
return X
def fit(self, X, y=None):
"""Compute k-means clustering.
Parameters
----------
X : array-like or sparse matrix, shape=(n_samples, n_features)
Training instances to cluster.
"""
# Added to remove scikit-learn internal dependenceies
raise NotImplemented
def fit_predict(self, X, y=None):
"""Compute cluster centers and predict cluster index for each sample.
Convenience method; equivalent to calling fit(X) followed by
predict(X).
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to transform.
Returns
-------
labels : array, shape [n_samples,]
Index of the cluster each sample belongs to.
"""
return self.fit(X).labels_
def fit_transform(self, X, y=None):
"""Compute clustering and transform X to cluster-distance space.
Equivalent to fit(X).transform(X), but more efficiently implemented.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to transform.
Returns
-------
X_new : array, shape [n_samples, k]
X transformed in the new space.
"""
# Currently, this just skips a copy of the data if it is not in
# np.array or CSR format already.
# XXX This skips _check_test_data, which may change the dtype;
# we should refactor the input validation.
X = self._check_fit_data(X)
return self.fit(X)._transform(X)
def transform(self, X):
"""Transform X to a cluster-distance space.
In the new space, each dimension is the distance to the cluster
centers. Note that even if X is sparse, the array returned by
`transform` will typically be dense.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to transform.
Returns
-------
X_new : array, shape [n_samples, k]
X transformed in the new space.
"""
check_is_fitted(self, 'cluster_centers_')
X = self._check_test_data(X)
return self._transform(X)
def _transform(self, X):
"""guts of transform method; no input validation"""
return euclidean_distances(X, self.cluster_centers_)
def predict(self, X):
"""Predict the closest cluster each sample in X belongs to.
In the vector quantization literature, `cluster_centers_` is called
the code book and each value returned by `predict` is the index of
the closest code in the code book.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data to predict.
Returns
-------
labels : array, shape [n_samples,]
Index of the cluster each sample belongs to.
"""
check_is_fitted(self, 'cluster_centers_')
X = self._check_test_data(X)
x_squared_norms = row_norms(X, squared=True)
return _labels_inertia(X, x_squared_norms, self.cluster_centers_)[0]
def score(self, X, y=None):
"""Opposite of the value of X on the K-means objective.
Parameters
----------
X : {array-like, sparse matrix}, shape = [n_samples, n_features]
New data.
Returns
-------
score : float
Opposite of the value of X on the K-means objective.
"""
check_is_fitted(self, 'cluster_centers_')
X = self._check_test_data(X)
x_squared_norms = row_norms(X, squared=True)
return -_labels_inertia(X, x_squared_norms, self.cluster_centers_)[1]
"""Base classes for all estimators."""
# Author: Gael Varoquaux <gael.varoquaux@normalesup.org>
# License: BSD 3 clause
import copy
import warnings
from collections import defaultdict
import platform
import inspect
import re
import numpy as np
from . import __version__
from .utils import _IS_32BIT
_DEFAULT_TAGS = {
'non_deterministic': False,
'requires_positive_data': False,
'X_types': ['2darray'],
'poor_score': False,
'no_validation': False,
'multioutput': False,
"allow_nan": False,
'stateless': False,
'multilabel': False,
'_skip_test': False,
'multioutput_only': False}
def clone(estimator, safe=True):
"""Constructs a new estimator with the same parameters.
Clone does a deep copy of the model in an estimator
without actually copying attached data. It yields a new estimator
with the same parameters that has not been fit on any data.
Parameters
----------
estimator : estimator object, or list, tuple or set of objects
The estimator or group of estimators to be cloned
safe : boolean, optional
If safe is false, clone will fall back to a deep copy on objects
that are not estimators.
"""
estimator_type = type(estimator)
# XXX: not handling dictionaries
if estimator_type in (list, tuple, set, frozenset):
return estimator_type([clone(e, safe=safe) for e in estimator])
elif not hasattr(estimator, 'get_params') or isinstance(estimator, type):
if not safe:
return copy.deepcopy(estimator)
else:
raise TypeError("Cannot clone object '%s' (type %s): "
"it does not seem to be a scikit-learn estimator "
"as it does not implement a 'get_params' methods."
% (repr(estimator), type(estimator)))
klass = estimator.__class__
new_object_params = estimator.get_params(deep=False)
for name, param in new_object_params.items():
new_object_params[name] = clone(param, safe=False)
new_object = klass(**new_object_params)
params_set = new_object.get_params(deep=False)
# quick sanity check of the parameters of the clone
for name in new_object_params:
param1 = new_object_params[name]
param2 = params_set[name]
if param1 is not param2:
raise RuntimeError('Cannot clone object %s, as the constructor '
'either does not set or modifies parameter %s' %
(estimator, name))
return new_object
def _pprint(params, offset=0, printer=repr):
"""Pretty print the dictionary 'params'
Parameters
----------
params : dict
The dictionary to pretty print
offset : int
The offset in characters to add at the begin of each line.
printer : callable
The function to convert entries to strings, typically
the builtin str or repr
"""
# Do a multi-line justified repr:
options = np.get_printoptions()
np.set_printoptions(precision=5, threshold=64, edgeitems=2)
params_list = list()
this_line_length = offset
line_sep = ',\n' + (1 + offset // 2) * ' '
for i, (k, v) in enumerate(sorted(params.items())):
if type(v) is float:
# use str for representing floating point numbers
# this way we get consistent representation across
# architectures and versions.
this_repr = '%s=%s' % (k, str(v))
else:
# use repr of the rest
this_repr = '%s=%s' % (k, printer(v))
if len(this_repr) > 500:
this_repr = this_repr[:300] + '...' + this_repr[-100:]
if i > 0:
if (this_line_length + len(this_repr) >= 75 or '\n' in this_repr):
params_list.append(line_sep)
this_line_length = len(line_sep)
else:
params_list.append(', ')
this_line_length += 2
params_list.append(this_repr)
this_line_length += len(this_repr)
np.set_printoptions(**options)
lines = ''.join(params_list)
# Strip trailing space to avoid nightmare in doctests
lines = '\n'.join(l.rstrip(' ') for l in lines.split('\n'))
return lines
def _update_if_consistent(dict1, dict2):
common_keys = set(dict1.keys()).intersection(dict2.keys())
for key in common_keys:
if dict1[key] != dict2[key]:
raise TypeError("Inconsistent values for tag {}: {} != {}".format(
key, dict1[key], dict2[key]
))
dict1.update(dict2)
return dict1
class BaseEstimator:
"""Base class for all estimators in scikit-learn
Notes
-----
All estimators should specify all the parameters that can be set
at the class level in their ``__init__`` as explicit keyword
arguments (no ``*args`` or ``**kwargs``).
"""
@classmethod
def _get_param_names(cls):
"""Get parameter names for the estimator"""
# fetch the constructor or the original constructor before
# deprecation wrapping if any
init = getattr(cls.__init__, 'deprecated_original', cls.__init__)
if init is object.__init__:
# No explicit constructor to introspect
return []
# introspect the constructor arguments to find the model parameters
# to represent
init_signature = inspect.signature(init)
# Consider the constructor parameters excluding 'self'
parameters = [p for p in init_signature.parameters.values()
if p.name != 'self' and p.kind != p.VAR_KEYWORD]
for p in parameters:
if p.kind == p.VAR_POSITIONAL:
raise RuntimeError("scikit-learn estimators should always "
"specify their parameters in the signature"
" of their __init__ (no varargs)."
" %s with constructor %s doesn't "
" follow this convention."
% (cls, init_signature))
# Extract and sort argument names excluding 'self'
return sorted([p.name for p in parameters])
def get_params(self, deep=True):
"""Get parameters for this estimator.
Parameters
----------
deep : boolean, optional
If True, will return the parameters for this estimator and
contained subobjects that are estimators.
Returns
-------
params : mapping of string to any
Parameter names mapped to their values.
"""
out = dict()
for key in self._get_param_names():
value = getattr(self, key, None)
if deep and hasattr(value, 'get_params'):
deep_items = value.get_params().items()
out.update((key + '__' + k, val) for k, val in deep_items)
out[key] = value
return out
def set_params(self, **params):
"""Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects
(such as pipelines). The latter have parameters of the form
``<component>__<parameter>`` so that it's possible to update each
component of a nested object.
Returns
-------
self
"""
if not params:
# Simple optimization to gain speed (inspect is slow)
return self
valid_params = self.get_params(deep=True)
nested_params = defaultdict(dict) # grouped by prefix
for key, value in params.items():
key, delim, sub_key = key.partition('__')
if key not in valid_params:
raise ValueError('Invalid parameter %s for estimator %s. '
'Check the list of available parameters '
'with `estimator.get_params().keys()`.' %
(key, self))
if delim:
nested_params[key][sub_key] = value
else:
setattr(self, key, value)
valid_params[key] = value
for key, sub_params in nested_params.items():
valid_params[key].set_params(**sub_params)
return self
def __repr__(self, N_CHAR_MAX=700):
# N_CHAR_MAX is the (approximate) maximum number of non-blank
# characters to render. We pass it as an optional parameter to ease
# the tests.
from .utils._pprint import _EstimatorPrettyPrinter
N_MAX_ELEMENTS_TO_SHOW = 30 # number of elements to show in sequences
# use ellipsis for sequences with a lot of elements
pp = _EstimatorPrettyPrinter(
compact=True, indent=1, indent_at_name=True,
n_max_elements_to_show=N_MAX_ELEMENTS_TO_SHOW)
repr_ = pp.pformat(self)
# Use bruteforce ellipsis when there are a lot of non-blank characters
n_nonblank = len(''.join(repr_.split()))
if n_nonblank > N_CHAR_MAX:
lim = N_CHAR_MAX // 2 # apprx number of chars to keep on both ends
regex = r'^(\s*\S){%d}' % lim
# The regex '^(\s*\S){%d}' % n
# matches from the start of the string until the nth non-blank
# character:
# - ^ matches the start of string
# - (pattern){n} matches n repetitions of pattern
# - \s*\S matches a non-blank char following zero or more blanks
left_lim = re.match(regex, repr_).end()
right_lim = re.match(regex, repr_[::-1]).end()
if '\n' in repr_[left_lim:-right_lim]:
# The left side and right side aren't on the same line.
# To avoid weird cuts, e.g.:
# categoric...ore',
# we need to start the right side with an appropriate newline
# character so that it renders properly as:
# categoric...
# handle_unknown='ignore',
# so we add [^\n]*\n which matches until the next \n
regex += r'[^\n]*\n'
right_lim = re.match(regex, repr_[::-1]).end()
ellipsis = '...'
if left_lim + len(ellipsis) < len(repr_) - right_lim:
# Only add ellipsis if it results in a shorter repr
repr_ = repr_[:left_lim] + '...' + repr_[-right_lim:]
return repr_
def __getstate__(self):
try:
state = super().__getstate__()
except AttributeError:
state = self.__dict__.copy()
if type(self).__module__.startswith('sklearn.'):
return dict(state.items(), _sklearn_version=__version__)
else:
return state
def __setstate__(self, state):
if type(self).__module__.startswith('sklearn.'):
pickle_version = state.pop("_sklearn_version", "pre-0.18")
if pickle_version != __version__:
warnings.warn(
"Trying to unpickle estimator {0} from version {1} when "
"using version {2}. This might lead to breaking code or "
"invalid results. Use at your own risk.".format(
self.__class__.__name__, pickle_version, __version__),
UserWarning)
try:
super().__setstate__(state)
except AttributeError:
self.__dict__.update(state)
def _get_tags(self):
collected_tags = {}
for base_class in inspect.getmro(self.__class__):
if (hasattr(base_class, '_more_tags')
and base_class != self.__class__):
more_tags = base_class._more_tags(self)
collected_tags = _update_if_consistent(collected_tags,
more_tags)
if hasattr(self, '_more_tags'):
more_tags = self._more_tags()
collected_tags = _update_if_consistent(collected_tags, more_tags)
tags = _DEFAULT_TAGS.copy()
tags.update(collected_tags)
return tags
class ClassifierMixin:
"""Mixin class for all classifiers in scikit-learn."""
_estimator_type = "classifier"
def score(self, X, y, sample_weight=None):
"""Returns the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy
which is a harsh metric since you require for each sample that
each label set be correctly predicted.
Parameters
----------
X : array-like, shape = (n_samples, n_features)
Test samples.
y : array-like, shape = (n_samples) or (n_samples, n_outputs)
True labels for X.
sample_weight : array-like, shape = [n_samples], optional
Sample weights.
Returns
-------
score : float
Mean accuracy of self.predict(X) wrt. y.
"""
from .metrics import accuracy_score
return accuracy_score(y, self.predict(X), sample_weight=sample_weight)
class RegressorMixin:
"""Mixin class for all regression estimators in scikit-learn."""
_estimator_type = "regressor"
def score(self, X, y, sample_weight=None):
"""Returns the coefficient of determination R^2 of the prediction.
The coefficient R^2 is defined as (1 - u/v), where u is the residual
sum of squares ((y_true - y_pred) ** 2).sum() and v is the total
sum of squares ((y_true - y_true.mean()) ** 2).sum().
The best possible score is 1.0 and it can be negative (because the
model can be arbitrarily worse). A constant model that always
predicts the expected value of y, disregarding the input features,
would get a R^2 score of 0.0.
Parameters
----------
X : array-like, shape = (n_samples, n_features)
Test samples. For some estimators this may be a
precomputed kernel matrix instead, shape = (n_samples,
n_samples_fitted], where n_samples_fitted is the number of
samples used in the fitting for the estimator.
y : array-like, shape = (n_samples) or (n_samples, n_outputs)
True values for X.
sample_weight : array-like, shape = [n_samples], optional
Sample weights.
Returns
-------
score : float
R^2 of self.predict(X) wrt. y.
Notes
-----
The R2 score used when calling ``score`` on a regressor will use
``multioutput='uniform_average'`` from version 0.23 to keep consistent
with `metrics.r2_score`. This will influence the ``score`` method of
all the multioutput regressors (except for
`multioutput.MultiOutputRegressor`). To specify the default value
manually and avoid the warning, please either call `metrics.r2_score`
directly or make a custom scorer with `metrics.make_scorer` (the
built-in scorer ``'r2'`` uses ``multioutput='uniform_average'``).
"""
from .metrics import r2_score
from .metrics.regression import _check_reg_targets
y_pred = self.predict(X)
# XXX: Remove the check in 0.23
y_type, _, _, _ = _check_reg_targets(y, y_pred, None)
if y_type == 'continuous-multioutput':
warnings.warn("The default value of multioutput (not exposed in "
"score method) will change from 'variance_weighted' "
"to 'uniform_average' in 0.23 to keep consistent "
"with 'metrics.r2_score'. To specify the default "
"value manually and avoid the warning, please "
"either call 'metrics.r2_score' directly or make a "
"custom scorer with 'metrics.make_scorer' (the "
"built-in scorer 'r2' uses "
"multioutput='uniform_average').", FutureWarning)
return r2_score(y, y_pred, sample_weight=sample_weight,
multioutput='variance_weighted')
class ClusterMixin:
"""Mixin class for all cluster estimators in scikit-learn."""
_estimator_type = "clusterer"
def fit_predict(self, X, y=None):
"""Performs clustering on X and returns cluster labels.
Parameters
----------
X : ndarray, shape (n_samples, n_features)
Input data.
y : Ignored
not used, present for API consistency by convention.
Returns
-------
labels : ndarray, shape (n_samples,)
cluster labels
"""
# non-optimized default implementation; override when a better
# method is possible for a given clustering algorithm
self.fit(X)
return self.labels_
class BiclusterMixin:
"""Mixin class for all bicluster estimators in scikit-learn"""
@property
def biclusters_(self):
"""Convenient way to get row and column indicators together.
Returns the ``rows_`` and ``columns_`` members.
"""
return self.rows_, self.columns_
def get_indices(self, i):
"""Row and column indices of the i'th bicluster.
Only works if ``rows_`` and ``columns_`` attributes exist.
Parameters
----------
i : int
The index of the cluster.
Returns
-------
row_ind : np.array, dtype=np.intp
Indices of rows in the dataset that belong to the bicluster.
col_ind : np.array, dtype=np.intp
Indices of columns in the dataset that belong to the bicluster.
"""
rows = self.rows_[i]
columns = self.columns_[i]
return np.nonzero(rows)[0], np.nonzero(columns)[0]
def get_shape(self, i):
"""Shape of the i'th bicluster.
Parameters
----------
i : int
The index of the cluster.
Returns
-------
shape : (int, int)
Number of rows and columns (resp.) in the bicluster.
"""
indices = self.get_indices(i)
return tuple(len(i) for i in indices)
def get_submatrix(self, i, data):
"""Returns the submatrix corresponding to bicluster `i`.
Parameters
----------
i : int
The index of the cluster.
data : array
The data.
Returns
-------
submatrix : array
The submatrix corresponding to bicluster i.
Notes
-----
Works with sparse matrices. Only works if ``rows_`` and
``columns_`` attributes exist.
"""
from .utils.validation import check_array
data = check_array(data, accept_sparse='csr')
row_ind, col_ind = self.get_indices(i)
return data[row_ind[:, np.newaxis], col_ind]
class TransformerMixin:
"""Mixin class for all transformers in scikit-learn."""
def fit_transform(self, X, y=None, **fit_params):
"""Fit to data, then transform it.
Fits transformer to X and y with optional parameters fit_params
and returns a transformed version of X.
Parameters
----------
X : numpy array of shape [n_samples, n_features]
Training set.
y : numpy array of shape [n_samples]
Target values.
Returns
-------
X_new : numpy array of shape [n_samples, n_features_new]
Transformed array.
"""
# non-optimized default implementation; override when a better
# method is possible for a given clustering algorithm
if y is None:
# fit method of arity 1 (unsupervised transformation)
return self.fit(X, **fit_params).transform(X)
else:
# fit method of arity 2 (supervised transformation)
return self.fit(X, y, **fit_params).transform(X)
class DensityMixin:
"""Mixin class for all density estimators in scikit-learn."""
_estimator_type = "DensityEstimator"
def score(self, X, y=None):
"""Returns the score of the model on the data X
Parameters
----------
X : array-like, shape = (n_samples, n_features)
Returns
-------
score : float
"""
pass
class OutlierMixin:
"""Mixin class for all outlier detection estimators in scikit-learn."""
_estimator_type = "outlier_detector"
def fit_predict(self, X, y=None):
"""Performs fit on X and returns labels for X.
Returns -1 for outliers and 1 for inliers.
Parameters
----------
X : ndarray, shape (n_samples, n_features)
Input data.
y : Ignored
not used, present for API consistency by convention.
Returns
-------
y : ndarray, shape (n_samples,)
1 for inliers, -1 for outliers.
"""
# override for transductive outlier detectors like LocalOulierFactor
return self.fit(X).predict(X)
class MetaEstimatorMixin:
_required_parameters = ["estimator"]
"""Mixin class for all meta estimators in scikit-learn."""
class MultiOutputMixin(object):
"""Mixin to mark estimators that support multioutput."""
def _more_tags(self):
return {'multioutput': True}
class _UnstableArchMixin(object):
"""Mark estimators that are non-determinstic on 32bit or PowerPC"""
def _more_tags(self):
return {'non_deterministic': (
_IS_32BIT or platform.machine().startswith(('ppc', 'powerpc')))}
def is_classifier(estimator):
"""Returns True if the given estimator is (probably) a classifier.
Parameters
----------
estimator : object
Estimator object to test.
Returns
-------
out : bool
True if estimator is a classifier and False otherwise.
"""
return getattr(estimator, "_estimator_type", None) == "classifier"
def is_regressor(estimator):
"""Returns True if the given estimator is (probably) a regressor.
Parameters
----------
estimator : object
Estimator object to test.
Returns
-------
out : bool
True if estimator is a regressor and False otherwise.
"""
return getattr(estimator, "_estimator_type", None) == "regressor"
def is_outlier_detector(estimator):
"""Returns True if the given estimator is (probably) an outlier detector.
Parameters
----------
estimator : object
Estimator object to test.
Returns
-------
out : bool
True if estimator is an outlier detector and False otherwise.
"""
return getattr(estimator, "_estimator_type", None) == "outlier_detector"