Copy disabled (too large)
Download .txt
Showing preview only (12,540K chars total). Download the full file to get everything.
Repository: pha4ge/hAMRonization
Branch: master
Commit: c8184233c52b
Files: 98
Total size: 11.9 MB
Directory structure:
gitextract_2sm_t59v/
├── .dockerignore
├── .github/
│ ├── ISSUE_TEMPLATE/
│ │ └── bug_report.md
│ └── workflows/
│ ├── dockerpublish.yml
│ ├── python_publish.yml
│ └── test_package.yml
├── .gitignore
├── Dockerfile
├── LICENSE.txt
├── README.md
├── docs/
│ ├── hAMRonization_specification_details.csv
│ ├── interactive_report_demo.html
│ └── subgrant/
│ └── PHA4GE_AMR_SubGrant_Orientation.pptx
├── hAMRonization/
│ ├── AbricateIO.py
│ ├── AmrFinderPlusIO.py
│ ├── AmrPlusPlusIO.py
│ ├── AribaIO.py
│ ├── CSStarIO.py
│ ├── DeepArgIO.py
│ ├── FARGeneIO.py
│ ├── GrootIO.py
│ ├── Interfaces.py
│ ├── KmerResistanceIO.py
│ ├── MykrobeIO.py
│ ├── README.md
│ ├── ResFamsIO.py
│ ├── ResFinderIO.py
│ ├── RgiIO.py
│ ├── SraxIO.py
│ ├── Srst2IO.py
│ ├── StarAmrIO.py
│ ├── TBProfilerIO.py
│ ├── __init__.py
│ ├── constants.py
│ ├── hAMRonizedResult.py
│ ├── hamronize.py
│ └── summarize.py
├── schema/
│ ├── PHA4GE AMR Gene & Variant Specification.csv
│ ├── PHA4GE AMR Gene & Variant Specification.json
│ └── csv2json.py
├── setup.cfg
├── setup.py
└── test/
├── .gitignore
├── README.md
├── __init__.py
├── data/
│ ├── dummy/
│ │ ├── abricate/
│ │ │ └── report.tsv
│ │ ├── amrfinderplus/
│ │ │ └── report.tsv
│ │ ├── amrplusplus/
│ │ │ └── gene.tsv
│ │ ├── ariba/
│ │ │ ├── report.tsv
│ │ │ └── report_var.tsv
│ │ ├── deepARG/
│ │ │ ├── output.mapping.ARG.
│ │ │ └── output.mapping.potential.ARG
│ │ ├── fargene/
│ │ │ └── retrieved-genes-class_A-hmmsearched.out
│ │ ├── groot/
│ │ │ └── groot_report.tsv
│ │ ├── kmerresistance/
│ │ │ └── results.res
│ │ ├── mykrobe/
│ │ │ ├── empty.json
│ │ │ └── mykrobe.json
│ │ ├── pointfinder/
│ │ │ └── PointFinder_results.txt
│ │ ├── resfams/
│ │ │ └── resfams.tblout
│ │ ├── resfinder/
│ │ │ └── data_resfinder.json
│ │ ├── rgi/
│ │ │ ├── rgi.txt
│ │ │ ├── rgi_orf.txt
│ │ │ └── rgi_var.txt
│ │ ├── srax/
│ │ │ └── sraX_detected_ARGs.tsv
│ │ ├── srst2/
│ │ │ └── report.tsv
│ │ ├── sstar/
│ │ │ └── report.tsv
│ │ ├── staramr/
│ │ │ ├── pointfinder.tsv
│ │ │ └── resfinder.tsv
│ │ └── tbprofiler/
│ │ └── tbprofiler.json
│ └── raw_outputs/
│ ├── abricate/
│ │ └── report.tsv
│ ├── amrfinderplus/
│ │ ├── afp_non_coding.tsv
│ │ ├── empty_report_with_header.tsv
│ │ ├── hamronized_non_coding.tsv
│ │ ├── non_coding_contig.fasta
│ │ ├── report_nucleotide.tsv
│ │ └── report_protein.tsv
│ ├── amrplusplus/
│ │ └── gene.tsv
│ ├── ariba/
│ │ └── report.tsv
│ ├── deeparg/
│ │ ├── output.mapping.ARG
│ │ └── output.mapping.potential.ARG
│ ├── fargene/
│ │ └── retrieved-genes-class_A-hmmsearched.out
│ ├── groot/
│ │ └── report.tsv
│ ├── kmerresistance/
│ │ └── results.res
│ ├── mykrobe/
│ │ └── report.json
│ ├── pointfinder/
│ │ └── PointFinder_results.txt
│ ├── resfams/
│ │ └── resfams.tblout
│ ├── resfinder/
│ │ ├── ResFinder_results_tab.txt
│ │ └── data_resfinder.json
│ ├── rgi/
│ │ └── rgi.txt
│ ├── rgibwt/
│ │ └── Kp11_bwtoutput.gene_mapping_data.txt
│ ├── srax/
│ │ └── sraX_detected_ARGs.tsv
│ ├── srst2/
│ │ └── SAMN13064234_srst2_report.tsv
│ ├── sstar/
│ │ └── report.tsv
│ ├── staramr/
│ │ ├── pointfinder.tsv
│ │ └── resfinder.tsv
│ └── tbprofiler/
│ └── tbprofiler.json
├── run_integration_test.sh
├── test_interfaces.py
└── test_parsing_validity.py
================================================
FILE CONTENTS
================================================
================================================
FILE: .dockerignore
================================================
.*
*.egg-info
*.pyc
/docs
/test
/schema
__pycache__
================================================
FILE: .github/ISSUE_TEMPLATE/bug_report.md
================================================
---
name: Bug report
about: Create a report to help us improve
title: "[BUG]"
labels: bug
assignees: ''
---
**Describe the bug**
A clear and concise description of what the bug is.
**Input**
The command-line input you used to run your data.
**Input file**
The file you used as an input to run the data.
Either paste a sample or upload the file.
**Error log**
The error that is thrown when you run the process.
**hAMRonization Version**
The version of hAMRonization you are using. Please ensure it is up to date.
**Expected behavior**
A clear and concise description of what you expected to happen.
**Screenshots**
If applicable, add screenshots to help explain your problem.
**Desktop (please complete the following information):**
- OS: [e.g. iOS]
- Browser [e.g. chrome, safari]
- Version [e.g. 22]
**Additional context**
Add any other context about the problem here.
If applicable, include dependency versions such as pandas version and Python version.
================================================
FILE: .github/workflows/dockerpublish.yml
================================================
name: Publish Docker Image
on:
release:
types: [published]
jobs:
main:
runs-on: ubuntu-latest
steps:
-
name: Set up QEMU
uses: docker/setup-qemu-action@v1
-
name: Set up Docker Buildx
uses: docker/setup-buildx-action@v1
-
name: Login to DockerHub
uses: docker/login-action@v1
with:
username: ${{ secrets.DOCKERHUB_USERNAME }}
password: ${{ secrets.DOCKERHUB_TOKEN }}
-
name: Build and push
id: docker_build
uses: docker/build-push-action@v2
with:
push: true
tags: finlaymaguire/hamronization:latest
build-args: |
SOFTWARE_VERSION=${{ github.event.release.tag_name }}
-
name: Image digest
run: echo ${{ steps.docker_build.outputs.digest }}
================================================
FILE: .github/workflows/python_publish.yml
================================================
# This workflow will upload a Python Package using Twine when a release is created
# For more information see: https://help.github.com/en/actions/language-and-framework-guides/using-python-with-github-actions#publishing-to-package-registries
name: python_publish
on:
release:
types: [created]
jobs:
pypi-publish:
name: Publish release to PyPI
runs-on: ubuntu-latest
environment:
name: pypi
url: https://pypi.org/p/hAMRonization
permissions:
id-token: write
steps:
- uses: actions/checkout@v4
- name: Set up Python
uses: actions/setup-python@v4
with:
python-version: "3.x"
- name: Install dependencies
run: |
python -m pip install --upgrade pip
pip install setuptools wheel
- name: Build package
run: |
python setup.py sdist bdist_wheel # Could also be python -m build
- name: Publish package distributions to PyPI
uses: pypa/gh-action-pypi-publish@release/v1
================================================
FILE: .github/workflows/test_package.yml
================================================
#This workflow will install Python dependencies, run tests and lint with a variety of Python versions
# For more information see: https://help.github.com/actions/language-and-framework-guides/using-python-with-github-actions
name: test_package
on:
push:
branches: [ master ]
pull_request:
branches: [ master ]
jobs:
build:
runs-on: ubuntu-latest
strategy:
matrix:
python-version: [3.10.20, 3.11.15, 3.12.13, 3.13.13, 3.14.4]
steps:
- uses: actions/checkout@v2
- name: Set up Python ${{ matrix.python-version }}
uses: actions/setup-python@v1
with:
python-version: ${{ matrix.python-version }}
- name: Install dependencies
run: |
python -m pip install --upgrade pip
pip install flake8 pytest
pip install .
- name: Lint hAMRonization library with flake8
run: |
pushd hAMRonization
# stop the build if there are Python syntax errors or undefined names
flake8 . --count --select=E9,F63,F7,F82 --show-source --statistics
# exit-zero treats all errors as warnings. The GitHub editor is 127 chars wide
flake8 . --count --exit-zero --max-complexity=20 --max-line-length=127 --statistics
popd
- name: Run sanity tests
run: |
pushd test
pytest
popd
- name: Run crude test of CLI parser for all tools
run: |
pushd test
bash run_integration_test.sh
popd
#- name: Validate all harmonized .json files with SALAD schema
# run: |
# pip install schema_salad
# pushd test/data
# for f in *.harmonized.json; do schema-salad-tool ../../schema/antimicrobial_resistance_genomic_analysis_result.schema.yml ${f}; done
================================================
FILE: .gitignore
================================================
.DS_Store
*.egg-info
*.pyc
build
dist
================================================
FILE: Dockerfile
================================================
# base image
FROM docker.io/library/python:3.14.4-alpine3.23
# metadata
LABEL org.opencontainers.image.version=1.2.1
LABEL org.opencontainers.image.base.name="docker.io/library/python:3.14.4-alpine3.23"
LABEL org.opencontainers.image.base.digest="sha256:105efb1f600e4e5d216985f6eeda0ed853ff9b38e65877039781f448ed677a0f"
LABEL org.opencontainers.image.title="hAMRronization"
LABEL org.opencontainers.image.title="Parse multiple Antimicrobial Resistance Analysis Reports into a common data structure"
LABEL org.opencontainers.image.source="https://github.com/pha4ge/hAMRonization"
LABEL org.opencontainers.image.documentation="https://github.com/pha4ge/hAMRonization/blob/master/README.md"
LABEL org.opencontainers.image.licenses="LGPL-3.0-only"
LABEL org.opencontainers.image.authors="Finlay Maguire <finlaymaguire@gmail.com>, Marco van Zwetselaar <io@zwets.it>"
LABEL tags="Genomics"
# add bash so Nextflow can run the container
RUN apk add --no-cache bash && rm -rf /var/cache/apk/*
# set the working directory in the container
WORKDIR /hAMRonization
# copy the sources into the container
COPY . /hAMRonization/src
# install dependencies and clean all up
RUN python -m pip --no-cache-dir install ./src && rm -rf ./src
# command to run on container start without args
CMD ["hamronize", "--help"]
================================================
FILE: LICENSE.txt
================================================
GNU LESSER GENERAL PUBLIC LICENSE
Version 3, 29 June 2007
Copyright (C) 2007 Free Software Foundation, Inc. <https://fsf.org/>
Everyone is permitted to copy and distribute verbatim copies
of this license document, but changing it is not allowed.
This version of the GNU Lesser General Public License incorporates
the terms and conditions of version 3 of the GNU General Public
License, supplemented by the additional permissions listed below.
0. Additional Definitions.
As used herein, "this License" refers to version 3 of the GNU Lesser
General Public License, and the "GNU GPL" refers to version 3 of the GNU
General Public License.
"The Library" refers to a covered work governed by this License,
other than an Application or a Combined Work as defined below.
An "Application" is any work that makes use of an interface provided
by the Library, but which is not otherwise based on the Library.
Defining a subclass of a class defined by the Library is deemed a mode
of using an interface provided by the Library.
A "Combined Work" is a work produced by combining or linking an
Application with the Library. The particular version of the Library
with which the Combined Work was made is also called the "Linked
Version".
The "Minimal Corresponding Source" for a Combined Work means the
Corresponding Source for the Combined Work, excluding any source code
for portions of the Combined Work that, considered in isolation, are
based on the Application, and not on the Linked Version.
The "Corresponding Application Code" for a Combined Work means the
object code and/or source code for the Application, including any data
and utility programs needed for reproducing the Combined Work from the
Application, but excluding the System Libraries of the Combined Work.
1. Exception to Section 3 of the GNU GPL.
You may convey a covered work under sections 3 and 4 of this License
without being bound by section 3 of the GNU GPL.
2. Conveying Modified Versions.
If you modify a copy of the Library, and, in your modifications, a
facility refers to a function or data to be supplied by an Application
that uses the facility (other than as an argument passed when the
facility is invoked), then you may convey a copy of the modified
version:
a) under this License, provided that you make a good faith effort to
ensure that, in the event an Application does not supply the
function or data, the facility still operates, and performs
whatever part of its purpose remains meaningful, or
b) under the GNU GPL, with none of the additional permissions of
this License applicable to that copy.
3. Object Code Incorporating Material from Library Header Files.
The object code form of an Application may incorporate material from
a header file that is part of the Library. You may convey such object
code under terms of your choice, provided that, if the incorporated
material is not limited to numerical parameters, data structure
layouts and accessors, or small macros, inline functions and templates
(ten or fewer lines in length), you do both of the following:
a) Give prominent notice with each copy of the object code that the
Library is used in it and that the Library and its use are
covered by this License.
b) Accompany the object code with a copy of the GNU GPL and this license
document.
4. Combined Works.
You may convey a Combined Work under terms of your choice that,
taken together, effectively do not restrict modification of the
portions of the Library contained in the Combined Work and reverse
engineering for debugging such modifications, if you also do each of
the following:
a) Give prominent notice with each copy of the Combined Work that
the Library is used in it and that the Library and its use are
covered by this License.
b) Accompany the Combined Work with a copy of the GNU GPL and this license
document.
c) For a Combined Work that displays copyright notices during
execution, include the copyright notice for the Library among
these notices, as well as a reference directing the user to the
copies of the GNU GPL and this license document.
d) Do one of the following:
0) Convey the Minimal Corresponding Source under the terms of this
License, and the Corresponding Application Code in a form
suitable for, and under terms that permit, the user to
recombine or relink the Application with a modified version of
the Linked Version to produce a modified Combined Work, in the
manner specified by section 6 of the GNU GPL for conveying
Corresponding Source.
1) Use a suitable shared library mechanism for linking with the
Library. A suitable mechanism is one that (a) uses at run time
a copy of the Library already present on the user's computer
system, and (b) will operate properly with a modified version
of the Library that is interface-compatible with the Linked
Version.
e) Provide Installation Information, but only if you would otherwise
be required to provide such information under section 6 of the
GNU GPL, and only to the extent that such information is
necessary to install and execute a modified version of the
Combined Work produced by recombining or relinking the
Application with a modified version of the Linked Version. (If
you use option 4d0, the Installation Information must accompany
the Minimal Corresponding Source and Corresponding Application
Code. If you use option 4d1, you must provide the Installation
Information in the manner specified by section 6 of the GNU GPL
for conveying Corresponding Source.)
5. Combined Libraries.
You may place library facilities that are a work based on the
Library side by side in a single library together with other library
facilities that are not Applications and are not covered by this
License, and convey such a combined library under terms of your
choice, if you do both of the following:
a) Accompany the combined library with a copy of the same work based
on the Library, uncombined with any other library facilities,
conveyed under the terms of this License.
b) Give prominent notice with the combined library that part of it
is a work based on the Library, and explaining where to find the
accompanying uncombined form of the same work.
6. Revised Versions of the GNU Lesser General Public License.
The Free Software Foundation may publish revised and/or new versions
of the GNU Lesser General Public License from time to time. Such new
versions will be similar in spirit to the present version, but may
differ in detail to address new problems or concerns.
Each version is given a distinguishing version number. If the
Library as you received it specifies that a certain numbered version
of the GNU Lesser General Public License "or any later version"
applies to it, you have the option of following the terms and
conditions either of that published version or of any later version
published by the Free Software Foundation. If the Library as you
received it does not specify a version number of the GNU Lesser
General Public License, you may choose any version of the GNU Lesser
General Public License ever published by the Free Software Foundation.
If the Library as you received it specifies that a proxy can decide
whether future versions of the GNU Lesser General Public License shall
apply, that proxy's public statement of acceptance of any version is
permanent authorization for you to choose that version for the
Library.
================================================
FILE: README.md
================================================

[](https://doi.org/10.1101/2024.03.07.583950)
[](https://zenodo.org/badge/latestdoi/248040662)
[](https://github.com/pha4ge/hAMRonization/blob/master/docs/subgrant/PHA4GE_AMR_SubGrant_Documentation.pdf)
[](https://github.com/pha4ge/hAMRonization/blob/master/docs/subgrant/PHA4GE_hAMRonization_espan%CC%83ol.pdf)
# hAMRonization
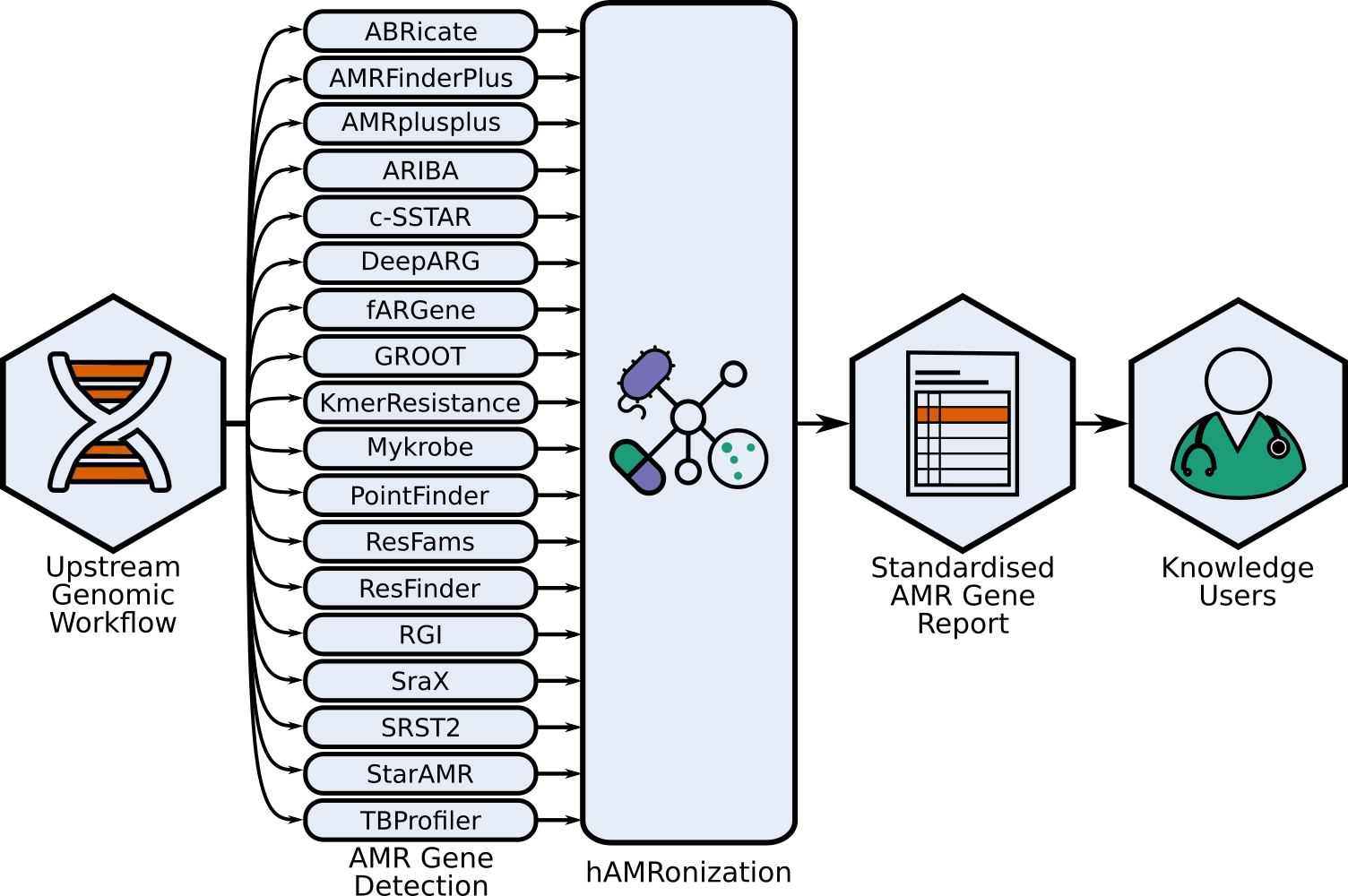
This repo contains the hAMRonization module and CLI parser tools combine the outputs of
17 disparate antimicrobial resistance gene detection tools into a single unified format.
This is an implementation of the [hAMRonization AMR detection specification scheme](docs/hAMRonization_specification_details.csv) which supports gene presence/absence resistance and mutational resistance (if supported by the underlying tool).
This supports a variety of summary options including an [interactive summary](https://finlaymagui.re/assets/interactive_report_demo.html).
## Installation
This tool requires python>=3.9 and [pandas](https://pandas.pydata.org/)
and the latest release can be installed directly from pip, conda, docker, this repository, or from the galaxy toolshed:
```
pip install hAMRonization
```
[](https://badge.fury.io/py/hamronization)
[](https://img.shields.io/pypi/dm/hAMRonization)
Or
```
conda create --name hamronization --channel conda-forge --channel bioconda hamronization
```



Or to install and run using docker, podman, singularity:
```
docker pull docker.io/finlaymaguire/hamronization:latest
docker run --rm docker.io/finlaymaguire/hamronization:latest hamronize --help
```
Or to install the latest development version:
```
git clone https://github.com/pha4ge/hAMRonization
pip install hAMRonization
```
Alternatively, hAMRonization can also be installed and used in [galaxy](https://galaxyproject.org/) via the [galaxy toolshed](https://toolshed.g2.bx.psu.edu/view/iuc/suite_hamronization/904ab154f8f4).
## Usage
**NOTE**: Only the output format used in the "last updated" version of the AMR prediction tool has been tested for accuracy. Older tool versions or updates which lead to a change in output format may not work.
In theory, this should only be a problem with major version changes but not all tools follow semantic versioning.
If you encounter any issues with newer tool versions then please create an issue in this repository.
```
usage: hamronize <tool> <options>
Convert AMR gene detection tool output(s) to hAMRonization specification format
options:
-h, --help show this help message and exit
-v, --version show program's version number and exit
Tools with hAMRonizable reports:
{abricate,amrfinderplus,amrplusplus,ariba,csstar,deeparg,fargene,groot,kmerresistance,resfams,resfinder,mykrobe,rgi,srax,srst2,staramr,tbprofiler,summarize}
abricate hAMRonize abricate's output report i.e., OUTPUT.tsv
amrfinderplus hAMRonize amrfinderplus's output report i.e., OUTPUT.tsv
amrplusplus hAMRonize amrplusplus's output report i.e., gene.tsv
ariba hAMRonize ariba's output report i.e., OUTDIR/OUTPUT.tsv
csstar hAMRonize csstar's output report i.e., OUTPUT.tsv
deeparg hAMRonize deeparg's output report i.e.,
OUTDIR/OUTPUT.mapping.ARG
fargene hAMRonize fargene's output report i.e., retrieved-
genes-*-hmmsearched.out
groot hAMRonize groot's output report i.e., OUTPUT.tsv (from `groot
report`)
kmerresistance hAMRonize kmerresistance's output report i.e., OUTPUT.res
resfams hAMRonize resfams's output report i.e., resfams.tblout
resfinder hAMRonize resfinder's JSON output report (use -j to produce)
mykrobe hAMRonize mykrobe's output report i.e., OUTPUT.json
rgi hAMRonize rgi's output report i.e., OUTPUT.txt or
OUTPUT_bwtoutput.gene_mapping_data.txt
srax hAMRonize srax's output report i.e., sraX_detected_ARGs.tsv
srst2 hAMRonize srst2's output report i.e., OUTPUT_srst2_report.tsv
staramr hAMRonize staramr's output report i.e., resfinder.tsv
tbprofiler hAMRonize tbprofiler's output report i.e., OUTPUT.results.json
summarize Provide a list of paths to the reports you wish to summarize
```
To look at a specific tool e.g. `abricate`:
```
>hamronize abricate -h
usage: hamronize abricate <options>
Applies hAMRonization specification to output from abricate (OUTPUT.tsv)
positional arguments:
report Path to tool report
optional arguments:
-h, --help show this help message and exit
--format FORMAT Output format (tsv or json)
--output OUTPUT Output location
--analysis_software_version ANALYSIS_SOFTWARE_VERSION
Input string containing the analysis_software_version for abricate
--reference_database_version REFERENCE_DATABASE_VERSION
Input string containing the reference_database_version for abricate
```
Therefore, hAMRonizing abricates output:
```
hamronize abricate ../test/data/raw_outputs/abricate/report.tsv --reference_database_version 3.2.5 --analysis_software_version 1.0.0 --format json
```
To parse multiple reports from the same tool at once just give a list of reports as the argument,
and they will be concatenated appropriately (i.e. only one header for tsv)
```
hamronize rgi --input_file_name rgi_report --analysis_software_version 6.0.0 --reference_database_version 3.2.5 test/data/raw_outputs/rgi/rgi.txt test/data/raw_outputs/rgibwt/Kp11_bwtoutput.gene_mapping_data.txt
```
You can summarize hAMRonized reports regardless of format using the 'summarize'
function:
```
> hamronize summarize -h
usage: hamronize summarize <options> <list of reports>
Concatenate and summarize AMR detection reports
positional arguments:
hamronized_reports list of hAMRonized reports
optional arguments:
-h, --help show this help message and exit
-t {tsv,json,interactive}, --summary_type {tsv,json,interactive}
Which summary report format to generate
-o OUTPUT, --output OUTPUT
Output file path for summary
```
This will take a list of report and create single sorted report in the
specified format just containing the unique entries across input reports.
This can handle mixed json and tsv hamronized report formats.
```
hamronize summarize -o combined_report.tsv -t tsv abricate.json ariba.tsv
```
The [interactive summary](https://finlaymagui.re/assets/interactive_report_demo.html) option will produce an html file that can be opened within the browser for navigable data exploration (feature developed
with @alexmanuele).
### Using within scripts
Alternatively, hAMRonization can be used within scripts (the metadata must contain the mandatory metadata that is not included in that tool's output, this can be checked by looking at the CLI flags in `hamronize <tool> --help`):
```
import hAMRonization
metadata = {"analysis_software_version": "1.0.1", "reference_database_version": "2019-Jul-28"}
parsed_report = hAMRonization.parse("abricate_report.tsv", metadata, "abricate")
```
The `parsed_report` is then a generator that yields hAMRonized result objects from the parsed report:
```
for result in parsed_report:
print(result)
```
Alternatively, you can use the `.write` attribute to export all results left in the generator to a file (if a filepath isn't provided, this will write to stdout).
```parsed_report.write('hAMRonized_abricate_report.tsv')```
You can also output a `json` formatted hAMRonized report:
`parsed_report.write('all_hAMRonized_abricate_report.json', output_format='json')`
If you want to write multiple reports to one file, this `.write` method can accept `append_mode=True` to append rather than overwrite the output file and not include the header (in tsv format).
`parsed_report.write('all_hAMRonized_abricate_report.tsv', append_mode=True)`
### Implemented Parsers
Currently implemented parsers and the last tool version for which they have been validated:
1. [abricate](hAMRonization/AbricateIO.py): last updated for v1.0.0
2. [amrfinderplus](hAMRonization/AmrFinderPlusIO.py): last updated for v4.0.3
3. [amrplusplus](hAMRonization/AmrPlusPlusIO.py): last updated for c6b097a
4. [ariba](hAMRonization/AribaIO.py): last updated for v2.14.6
5. [csstar](hAMRonization/CSStarIO.py): last updated for v2.1.0
6. [deeparg](hAMRonization/DeepArgIO.py): last updated for v1.0.2
7. [fargene](hAMRonization/FARGeneIO.py): last updated for v0.1
8. [groot](hAMRonization/GrootIO.py): last updated for v1.1.2
9. [kmerresistance](hAMRonization/KmerResistanceIO.py): late updated for v2.2.0
10. [mykrobe](test/data/raw_outputs/mykrobe/report.json): last updated for v0.8.1
11. ~pointfinder~ (removed, PointFinder is now integrated in ResFinder)
12. [resfams](hAMRonization/ResFamsIO.py): last updated for hmmer v3.3.2
13. [resfinder](hAMRonization/ResFinderIO.py): last updated for v4.6.0
14. [rgi](hAMRonization/RgiIO.py) (includes RGI-BWT) last updated for v5.2.0
15. [srax](hAMRonization/SraxIO.py): last updated for v1.5
16. [srst2](hAMRonization/Srst2IO.py): last updated for v0.2.0
17. [staramr](hAMRonization/StarAmrIO.py): last updated for v0.8.0
18. [tbprofilder](test/data/raw_outputs/tbprofiler/tbprofiler.json): last updated for v3.0.8
## Implementation Details
### hAMRonizedResult Data Structure
The hAMRonization specification is implemented in the [hAMRonizedResult dataclass](https://github.com/pha4ge/harmonized-amr-parsers/blob/master/hAMRonization/hAMRonization/hAMRonizedResult.py#L6).
This is a simple datastructure that uses positional and key-word args to distinguish mandatory from optional hAMRonization fields.
It also uses type-hinting to validate the supplied values are of the correct type
Each parser follows a similar strategy, using a common interface.
This has been designed to match the `biopython` `SeqIO` `parse` function
>>> import hAMRonization
>>> filename = "abricate_report.tsv"
>>> metadata = {"analysis_software_version": "1.0.1", "reference_database_version": "2019-Jul-28"}
>>> for result in hAMRonization.parse(filename, metadata, "abricate"):
... print(result)
Where the final argument to the `hAMRonization.parse` command is whichever tool is being parsed.
### hAMRonizedResultIterator
An abstract iterator is then implemented to ingest a given AMR tool's report
(via the appropriate subclassed implementation), hAMRonize results i.e. translate the
original inputs to the fields in the hAMRonization specification, and yield a stream of
hAMRonizedResult dataclasses.
This iterator also implements a write function to enable outputting the contents
to a output stream or filehandle in either tsv or json format.
### Tool-specific Iterators
Each tool has a specific subclass of this abstract hAMRonizedResultIterator e.g. `AbricateIO.AbricateIterator`.
These include an attribute containing the mapping of the tools original output report fields to the hAMRonized specification fields (`self.field_mapping`), as well as handling specifying any additional required metadata.
The `parse` method of these subclasses then implements the tool-specific parsing logic required.
This is typically a simple `csv.DictReader` but can be more complex such as the json parsing of `resfinder` output,
or the modification of output fields required to better fit some tools into the hAMRonization specification.
## Contributing
We welcome contributions for users in any form (from github issues flagging problems/requests) to pull requests of bug fixes or adding new parsers.
## Setting up a Development Environment
First fork this repository and set up a development environment (replacing `YOURUSERNAME` with your github username:
```
git clone https://github.com/YOURUSERNAME/hAMRonization
conda create -n hAMRonization python pip pytest flake8
conda activate hAMRonization
cd hAMRonization
pip install -e .
```
## Testing and Linting
On every commit github actions automatically runs tests and linting to check
the code.
You can manually run these in your development environment as well.
To run a full set of integration tests:
pushd test
bash run_integration_test.sh
popd
To run unit tests that verify parsing validity for each tool
as well as generation of valid summaries you can use pytest:
pip install pytest
pushd test
pytest
popd
Finally to run linting and check whether your code matches the project
code style:
pushd hAMRonization
flake8 . --count --select=E9,F63,F7,F82 --show-source --statistics
flake8 . --count --exit-zero --max-complexity=20 --max-line-length=127 --statistics
popd
## Adding a new parser
If you wish to add a parser for a new tool here are the main steps required:
1. Add an entry into `_RequiredToolMetadata` and `_FormatToIterator` in `hAMRonziation/__init__.py` which points to the appropriate `ToolNameIO.py` containing the tool's Iterator subclass
2. In `ToolNameIO.py` add a `required_metadata` list containing any mandatory fields not implemented by the tool
3. Then add a class `ToolNameIterator(hAMRonizedResultIterator)` and implement the `__init__` methods with the approriate mapping (`self.field_mapping`), and metadata (`self.metadata`).
4. To this class, add a `parse` method which reads an opened file stream into a dictionary per line/result (matching the keys of `self.field_mapping`) and yields the output of `self.hAMRonize` being applied to that dictionary.
5. To add a CLI parser for the tool, create a python file in the `parsers` directory:
```
from hAMRonization import Interfaces
if __name__ == '__main__':
Interfaces.cli_parser('toolname')
```
Alternatively, the `hAMRonized_parser.py` can be used as a common script interface to all implemented parsers.
6. Finally, following the template in `test/test_parsing_validity.py`, please generate a unit test that ensures the parser is working as you intend it to!
If you have any questions about any of this or need any help, please file an issue.
## FAQ
* What's the difference between an Antimicrobial Resistance 'Result' and 'Report'?
* For the purposes of this project, a 'Report' is an output file (or collection of files) from an AMR analysis tool.
A 'Result' is a single entry in a report. For example, a single line in an abricate report file is a single Antimicrobial
Resistance 'Result'.
### Known Issues
Here are some known issues that we would welcome input on trying to solve!
#### Limitations of specification
- mandatory fields: `gene_symbol` and `gene_name` are confusing and not usually both present (only consistently used in AFP). Means tools either need 1:2 mapping i.e. single output field maps to both `gene_symbol` and `gene_name` OR have fragile text splitting of single field that won't be robust to databases changes. Current solution is 1:2 mapping e.g. staramr
- inconsistent nomenclature of terms being used in specification fields: target, query, subject, reference. Need to stick to one name for sequence with which the database is being searched, and one the hit that results from that search.
- `sequence_identity`: is sequence type specific %id amino acids != %id nucleotide but does this matter?
- `coverage_depth` seems to include both tool fields that are average depth of read and just plain overall read-count,
- `contig_id` isn't general enough when some tools this ID naturally corresponds to a `read_name` (deepARG), individual ORF (resfams), or protein sequence (AFP with protein input): *change to `query_id_name` or similar?*
## Citation
If you use hAMRonization please cite the following publication:
> I Mendes et al. 2024. "hAMRonization: Enhancing antimicrobial resistance prediction using the PHA4GE AMR detection specification and tooling". bioRxiv. https://doi.org/10.1101/2024.03.07.583950
================================================
FILE: docs/hAMRonization_specification_details.csv
================================================
Interface Label,Required/Optional,Definition,Value Type,Example,Guidance,Comments,
Input File Name,Mandatory,Name or other identifier of an entry from a sample database.,String,ERR3581801,Tools can have multiple input files (genomic/amino acid). Multiple input files will be accomodated by their corresponding parser. ,,
Input Sequence ID,Optional,The identifier of a molecular sequence being analyzed.,String,DAAGAT010000041.1,Can be left blank if not provided.,,
Input Protein Start ,Optional,The position of the first amino acid in a protein sequence being analyzed (input protein sequence). Also known as the query protein start site in a BLAST search. ,Int,6,"The value should be biologically relevant, and should not start with 0. This field only applies to tools providing gene sequence outputs. Can be left blank if output not provided/applicable.","can start at 1 or 0, fmin/fmax (start at 0), NCBI starts at 1 but CARD database starts at 0",
Input Protein Stop ,Optional,The position of the last amino acid in a protein sequence being analyzed (input protein sequence). Also known as the query protein stop site in a BLAST search. ,Int,307,This field only applies to tools providing gene sequence outputs. Can be left blank if output not provided/applicable.,,
Input Gene Start ,Optional,The position of the first nucleotide in a gene sequence being analyzed (input gene sequence). Also known as the query gene start site in a BLAST search. ,Int,18,,,
Input Gene Stop,Optional,The position of the last nucelotide in a gene sequence being analyzed (input gene sequence). Also known as the query gene stop site in a BLAST search. ,Int,921,,,
Reference Protein Start ,Optional,The position of the first amino acid in a reference protein sequence (sequence being used for comparison). Also known as the subject protein start site in a BLAST search. ,Int,1,This field only applies to tools providing amino acid sequence outputs. Can be left blank if output not provided/applicable.,,
Reference Protein Stop ,Optional,The position of the last amino acid in a reference protein sequence (sequence being used for comparison). Also known as the subject protein stop site in a BLAST search.,Int,300,This field only applies to tools providing gene sequence outputs. Can be left blank if output not provided/applicable.,,
Reference Gene Start ,Optional,The position of the first nucleotide in a reference gene sequence (sequence being used for comparison). Also known as the subject gene start site in a BLAST search.,Int,1,,,
Reference Gene Stop ,Optional,The position of the last nucelotide in a reference sequence (sequence being used for comparison). Also known as the subject gene stop site in a BLAST search.,Int,900,,,
Strand Orientation,Optional,The orientation of a gene in a double stranded DNA replicon. ,String,"(+), sense","Values should be sense or antisense, or the corresponding short form ""+"" or ""-"". The terms ""positive"" and ""negative"" should be avoided. Can be left blank if not provided.","sense or antisense, or use short hand +/-",
Gene Symbol,Mandatory,The short name of a gene or gene product; a single word that does not contain white space characters. It is typically derived from the gene/gene product name.,String,"catA1, blaOXA-101","Acceptable values may represent gene symbols or gene product symbols. ""Gene symbol"" and ""allele"" (based on amino acid variations) may be used interchangeably due to nuances in resistance gene family nomenclature. Parsers should include values from ""gene symbol"" and ""allele"" fields in this category. Either the ""gene symbol"" or the ""gene name"" must be provided.","every aa diff for beta lactamases is a new allele, not the case for all amr gene families; novel allele, leave it blank","Comment: I don't think if it's a novel allele, this should be blank (hence why it's mandatory); it should instead say 'blaOXA', just without a number afterwards designating a specific allele (or it will alternatively list the allele with closest homology)"
Gene Name,Mandatory,"The name of a gene, (typically) assigned by a person and/or according to a naming scheme. It may contain white space characters and is typically more intuitive and readable than a gene symbol. It (typically) may be used to identify similar genes in different species and to derive a gene symbol.",String,type A-1 chloramphenicol O-acetyltransferase ,"The long name for the gene/protein. Gene names or gene product names are acceptable. Either the ""gene symbol"" or the ""gene name"" must be provided.",stress long form (describes protein); gene name or protein name?,
Coverage (depth),Optional,Coverage (read depth or depth) is the average number of reads representing a given nucleotide in the reconstructed sequence.,Float,56,The value is expressed as a fold value e.g. 56x.,,
Coverage (%),Optional,The percentage of the reference sequence covered by the sequence of interest.,Float,90,"The sequences can refer to reads and genome, nucelotides and genes, or amino acids and proteins. This value should be normalized and expressed as percentage. Do not include the ""%"" sign in the value. Either ""% Coverage (breadth)"" of ""Coverage (breadth) is required.","Comment: I think this should be mandatory; if you have a hit for a gene, the first question you have is the % identity and % length. This is necessary context to determine whether it's likely a functional protein.",
Coverage (ratio),Optional,The ratio of the reference sequence covered by the sequence of interest.,,450/500,"The sequences can refer to reads and genome, nucelotides and genes, or amino acids and proteins. This value should not be normalized and expressed as the ratio of actual positions being compared e.g. 450/500. Either ""% Coverage (breadth)"" of ""Coverage (breadth) is required.",,
Sequence Identity,Optional,Sequence identity is the number (%) of matches (identical characters) in positions from an alignment of two molecular sequences.,Float,1,"The sequences can refer to reads and genome, nucelotides and genes, or amino acids and proteins. The value should be expressed as a percentage. Do not include the ""%"" sign in the value.",,
Reference Database ID,Mandatory,"Database containing references genomes for genome annotation, gene identification, characterization etc.",String,"ncbi, ResFinder",,,
Reference Database Version,Mandatory,Version of a particular database.,String,,,ISSUE: No standards available for database versioning,
Reference Accession,Mandatory,"A persistent, unique identifier of a molecular sequence database entry.",String,NF000491.1 ,,,
Reference Gene Length,Optional,The length (number of positions) of a gene sequence being used as a reference for comparison.,Int,657,,"more specific term under ""Genetic Sequence Length (NCIT:C135487)""",
Reference Protein Length,Optional,The length (number of positions) of a protein sequence being used as a reference for comparison.,Int,219,,,
Input Gene Length,Optional,The length (number of positions) of a gene sequence being analyzed (input gene sequence).,Int,657,,"more specific term under ""Genetic Sequence Length (NCIT:C135487)""",
Input Protein Length,Optional,The length (number of positions) of a protein sequence being analyzed (input protein sequence).,Int,219,,,
Drug Class,Optional,"A set of medications and other compounds that have similar chemical structures, the same mechanism of action (i.e., bind to the same biological target), a related mode of action, and/or are used to treat the same disease.",String,Phenicol,Standardized terms from the ChEBI ontology should be used. Find terms using https://www.ebi.ac.uk/ols/ontologies/chebi. Can be left blank if not provided.,,
Antimicrobial Agent ,Optional,"A substance that kills or slows the growth of microorganisms, including bacteria, viruses, fungi and protozoans.",String,CHLORAMPHENICOL,This should describe a specific agent. Standardized terms from the ChEBI ontology should be used. Find terms using https://www.ebi.ac.uk/ols/ontologies/chebi. Can be left blank if not provided.,,
Resistance mechanism,Optional,Cellular processes in a pathogen that result in antimicrobial drug resistance.,String,target alteration,Standardized terms from the ChEBI ontology should be used. Find terms using https://www.ebi.ac.uk/ols/ontologies/aro. Can be left blank if not provided. ,,
Analysis Software Name,Mandatory,"The name of a computer package, application, method or function.",String,amrfinder,Not typically included in an AMR prediction output. The user may have to provide the value.,,
Analysis Software Version,Mandatory,"A version number is a unique number or set of numbers assigned to a specific release of a software program, file, firmware, device driver, or even hardware. Typically, as updates and entirely new editions of a program or driver are released, the version number will increase.",String,1.2.5,Not typically included in an AMR prediction output. The user may have to provide the value. Semantic versioning is strongly recommended.,https://semver.org/,"Comment: the version of the gene database is more typical than the software, so this should perhaps not be required"
Genetic Variation Type,Mandatory,"The type of genetic variant (e.g. gene presence, gene absence, protein variant, nucleotide variant).",String,protein_mutation,,,
Variant Frequency,Optional,The frequency of the variant in the data.,Float,0.5,,,
Nucleotide Mutation,Optional,The nucleotide sequence change(s) detected in the sequence being analyzed compared to a reference in HGVS format.,String,c.1349C>T,,,
Nucleotide Mutation Interpretation,Optional,The description of the HGVS encoded nucelotide mutation(s) for clinical interpretation. ,String,This is a subst found in rpoB at position 1349 where the reference has a C and the sample has a T,,,
Protein Mutation,Optional,The protein sequence change(s) detected in the sequence being analyzed compared to a reference in HGVS format.,String,p.Ser450Leu,,,
Protein Mutation Interpretation,Optional,The description of the HGVS encoded protein mutation(s) for clinical interpretation. ,String,This is a amino acid subst found in rpoB at position 450 where the reference has a Serine and the sample has a Leucine,,,
================================================
FILE: docs/interactive_report_demo.html
================================================
<!DOCTYPE html>
<html>
<head>
<title>hAMRonized Results</title>
<link rel='icon' href="https://pha4ge.org/wp-content/uploads/2020/06/logo-clear.png">
<!-- Required meta tags -->
<meta charset="utf-8">
<meta name="viewport" content="width=device-width, initial-scale=1, shrink-to-fit=no">
<!-- Bootstrap CSS -->
<link rel="stylesheet" href="https://stackpath.bootstrapcdn.com/bootstrap/4.3.1/css/bootstrap.min.css" integrity="sha384-ggOyR0iXCbMQv3Xipma34MD+dH/1fQ784/j6cY/iJTQUOhcWr7x9JvoRxT2MZw1T" crossorigin="anonymous">
<!-- custom CSS. Must be after botostrap -->
<!-- pha4ge dark blue: #1c4f77 -->
<!-- pha4ge light blue: #4879a1 -->
<style>
.search_hit{
background-color: paleturquoise;
}
.selected{
background-color: gold;
}
.search_hit.selected{
background-color: springgreen;
}
.amr_hit:hover{
background-color: lemonchiffon;
}
td
{
padding:0 15px;
}
.phage-blue{
background-color: #1c4f77;
color: white;
}
.phage-blue-light{
background-color: #4879a1;
color: white;
}
.sticky{
position: sticky;
top: 6rem;
}
.selection-bar{
max-height: 48rem;
}
.data-display{
max-height: 36rem;
}
.scroll-x{
overflow-x: auto;
}
.scroll-y{
overflow-y: auto;
}
/* custom table bg-colors
currently broken. TODO make it work... breaks hover.
.table-striped>tbody>tr:nth-child(even)>td,
.table-striped>tbody>tr:nth-child(even)>th {
background-color: ##99d8c9; // Choose your own color here
}
.table-striped>tbody>tr:nth-child(odd)>td,
.table-striped>tbody>tr:nth-child(odd)>th {
background-color: #e5f5f9; // Choose your own color here
} */
</style>
</head>
<body>
<!-- Navbar -->
<nav class="navbar sticky-top navbar-light bg-light">
<a class="navbar-brand" href="#">
<img src="https://pha4ge.org/wp-content/uploads/2020/04/logob.png" width="320" height="74" alt="">
</a>
<div class="form-inline my-2 my-lg-0">
<input id="gene-search" class="form-control mr-sm-2" onkeyup="geneSearch()" placeholder="Search" aria-label="Search">
<button id="results-only" class="btn btn-outline-success my-2 my-sm-0" onclick="showOnlyResults()">Show Only Genomes With Hits</button>
<button id="restore-table" class="btn btn-outline-primary my-2 my-sm-0" onclick="restoreTable()" style="display:none">Restore Results</button>
</div>
</nav>
<div class="container-flex">
<div class="row">
<div class="col col-lg-9 col-md-12" id="dynamic-table">
<!--<input type='button' onclick='CreateTableFromJSON()' value='make table'/>-->
</div>
<div class="col">
<div class="sticky selection-bar scroll-y">
<div class="row">
<div class="col">
<div class="card">
<div class="card-header phage-blue-light">
Search Results
</div>
<ul class="list-group list-group-flush">
<li class="list-group-item d-flex justify-content-between align-items-center">Total hits:<span class="badge badge-primary badge-pill" id="nhits-badge">0</span></li>
<li class="list-group-item d-flex justify-content-between align-items-center">Genomes with hits:<span class="badge badge-primary badge-pill" id="ngenomes-badge">0</span></li>
<li class="list-group-item d-flex justify-content-between align-items-center">Tools with hits:<span class="badge badge-primary badge-pill" id="ntools-badge">0</span></li>
<li class="list-group-item d-flex justify-content-between align-items-center">
<div>
<span data-toggle="tooltip" data-placement="bottom" class="badge badge-dark text-light badge-pill" title="Indicates the number of genomes in which at least one tool found a hit matching the search query, but not all tools did.">
?
</span>
Differential results:
</div>
<span class="badge badge-primary badge-pill" id="ndiff-badge">0</span>
</li>
</ul>
</div>
</div>
</div>
<div class="row">
<div class="col">
<div class="card">
<div class="card-header phage-blue-light">
Selected
</div>
<ul id="selected-genes" class="list-group list-group-flush">
<!--populated by JS -->
</ul>
<div class="row">
<div class="col-6">
<button type="button" class="btn btn-info btn-block selection-button" onclick="displayAmrData()" disabled>Compare</button>
</div>
<div class="col-6">
<button type="button" class="btn btn-warning btn-block selection-button" onclick="clearSelected()" disabled>Clear</button>
</div>
</div>
</div>
</div>
</div>
</div>
</div>
</div>
</div>
<div id="select-container" class="container-flex phage-blue-light">
<div class="row bg-light">
<div class="container-fluid" id="data-display">
</div>
</div>
</div>
<!-- JavaScript -->
<!-- jQuery first, then Popper.js, then Bootstrap JS -->
<script src="https://code.jquery.com/jquery-3.4.1.min.js" integrity="sha256-CSXorXvZcTkaix6Yvo6HppcZGetbYMGWSFlBw8HfCJo=" crossorigin="anonymous"></script>
<script src="https://cdnjs.cloudflare.com/ajax/libs/popper.js/1.14.7/umd/popper.min.js" integrity="sha384-UO2eT0CpHqdSJQ6hJty5KVphtPhzWj9WO1clHTMGa3JDZwrnQq4sF86dIHNDz0W1" crossorigin="anonymous"></script>
<script src="https://stackpath.bootstrapcdn.com/bootstrap/4.3.1/js/bootstrap.min.js" integrity="sha384-JjSmVgyd0p3pXB1rRibZUAYoIIy6OrQ6VrjIEaFf/nJGzIxFDsf4x0xIM+B07jRM" crossorigin="anonymous"></script>
<script type='text/javascript'>
//parse JSON
var hamronized_data = JSON.parse('[{"input_file_name": "ERR873305", "abricate: config 0": [{"input_file_name": "ERR873305", "gene_symbol": "aac(6\')-29a", "gene_name": "aminoglycoside N-acetyltransferase AAC(6\')-29a", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_048575.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 72.0, "query_stop_nt": 473.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "aph(3\')-IIb", "gene_name": "aminoglycoside O-phosphotransferase APH(3\')-IIb", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047424.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 99.26, "contig_id": "gnl|BUGS|ERR873305_93", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 12299.0, "query_stop_nt": 13105.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "KANAMYCIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaOXA-395", "gene_name": "OXA-50 family oxacillin-hydrolyzing class D beta-lactamase OXA-395", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_049684.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_15", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 18488.0, "query_stop_nt": 19276.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaPDC-158", "gene_name": "class C beta-lactamase PDC-158", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_050831.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 75.51, "contig_id": "gnl|BUGS|ERR873305_303", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 436.0, "query_stop_nt": 1448.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 84.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CEPHALOSPORIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaPDC-55", "gene_name": "class C beta-lactamase PDC-55", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_049929.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.77, "contig_id": "gnl|BUGS|ERR873305_60", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 60.0, "query_stop_nt": 1282.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CEPHALOSPORIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaVIM-2", "gene_name": "subclass B1 metallo-beta-lactamase VIM-2", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_050347.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 604.0, "query_stop_nt": 1404.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CARBAPENEM", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "catB7", "gene_name": "type B-4 chloramphenicol O-acetyltransferase CatB7", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047614.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873305_63", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 29357.0, "query_stop_nt": 29995.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CHLORAMPHENICOL", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "cmlB1", "gene_name": "chloramphenicol efflux MFS transporter CmlB1", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047658.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 79.5, "contig_id": "gnl|BUGS|ERR873305_74", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6748.0, "query_stop_nt": 7957.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 95.58, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CHLORAMPHENICOL", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "crpP", "gene_name": "ciprofloxacin resistance protein CrpP", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_062203.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873305_43", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 4289.0, "query_stop_nt": 4486.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "FLUOROQUINOLONE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "fosA-354827590", "gene_name": "FosA family fosfomycin resistance glutathione transferase", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047883.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873305_36", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8371.0, "query_stop_nt": 8778.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "FOSFOMYCIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "sul1", "gene_name": "sulfonamide-resistant dihydropteroate synthase Sul1", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_048082.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 1932.0, "query_stop_nt": 2771.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "SULFONAMIDE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}], "amrfinderplus: config 0": [{"input_file_name": "ERR873305", "gene_symbol": "aac(6\')-29a", "gene_name": "aminoglycoside N-acetyltransferase AAC(6\')-29a", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_064190968.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 72.0, "query_stop_nt": 470.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 133.0, "reference_protein_length": null, "target_gene_length": 133.0, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": "AMINOGLYCOSIDE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "aph(3\')-IIb", "gene_name": "aminoglycoside O-phosphotransferase APH(3\')-IIb", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003113011.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 99.25, "contig_id": "gnl|BUGS|ERR873305_93", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 12302.0, "query_stop_nt": 13105.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 268.0, "reference_protein_length": null, "target_gene_length": 268.0, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": "KANAMYCIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaOXA-395", "gene_name": "OXA-50 family oxacillin-hydrolyzing class D beta-lactamase OXA-395", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_015649877.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_15", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 18491.0, "query_stop_nt": 19276.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 262.0, "reference_protein_length": null, "target_gene_length": 262.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "BETA-LACTAM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaPDC-3", "gene_name": "class C extended-spectrum beta-lactamase PDC-3", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003121934.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_60", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 92.0, "query_stop_nt": 1282.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 397.0, "reference_protein_length": null, "target_gene_length": 397.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "CEPHALOSPORIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaVIM-2", "gene_name": "subclass B1 metallo-beta-lactamase VIM-2", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003108247.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 604.0, "query_stop_nt": 1401.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 266.0, "reference_protein_length": null, "target_gene_length": 266.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "CARBAPENEM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "catB7", "gene_name": "type B-4 chloramphenicol O-acetyltransferase CatB7", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003112709.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 99.06, "contig_id": "gnl|BUGS|ERR873305_63", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 29357.0, "query_stop_nt": 29992.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 212.0, "reference_protein_length": null, "target_gene_length": 212.0, "target_protein_length": null, "drug_class": "PHENICOL", "antimicrobial_agent": "CHLORAMPHENICOL", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "crpP", "gene_name": "ciprofloxacin resistance protein CrpP", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_033179079.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 98.46, "contig_id": "gnl|BUGS|ERR873305_43", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 4289.0, "query_stop_nt": 4483.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 65.0, "reference_protein_length": null, "target_gene_length": 65.0, "target_protein_length": null, "drug_class": "FLUOROQUINOLONE", "antimicrobial_agent": "FLUOROQUINOLONE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "fosA", "gene_name": "FosA family fosfomycin resistance glutathione transferase", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003082280.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 98.52, "contig_id": "gnl|BUGS|ERR873305_36", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8374.0, "query_stop_nt": 8778.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 135.0, "reference_protein_length": null, "target_gene_length": 135.0, "target_protein_length": null, "drug_class": "FOSFOMYCIN", "antimicrobial_agent": "FOSFOMYCIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "qacEdelta1", "gene_name": "quaternary ammonium compound efflux SMR transporter QacE delta 1", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_000679427.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 1591.0, "query_stop_nt": 1935.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 115.0, "reference_protein_length": null, "target_gene_length": 115.0, "target_protein_length": null, "drug_class": "QUATERNARY AMMONIUM", "antimicrobial_agent": "QUATERNARY AMMONIUM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "sul1", "gene_name": "sulfonamide-resistant dihydropteroate synthase Sul1", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_000259031.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 1932.0, "query_stop_nt": 2768.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 279.0, "reference_protein_length": null, "target_gene_length": 279.0, "target_protein_length": null, "drug_class": "SULFONAMIDE", "antimicrobial_agent": "SULFONAMIDE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}], "csstar: config 0": [{"input_file_name": "ERR873305", "gene_symbol": "aac(6\')", "gene_name": "aac(6\')-29a", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "aac(6\')-29a", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 396.0, "reference_protein_length": null, "target_gene_length": 396.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "aph(3\')", "gene_name": "aph(3\')-IIb", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "aph(3\')-IIb", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.257, "contig_id": "gnl|BUGS|ERR873305_93", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 807.0, "reference_protein_length": null, "target_gene_length": 807.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "bcr1", "gene_name": "bcr1", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "bcr1", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_152", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 1209.0, "reference_protein_length": null, "target_gene_length": 1209.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaOXA", "gene_name": "blaOXA-395", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaOXA-395", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_15", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 789.0, "reference_protein_length": null, "target_gene_length": 789.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaPAO", "gene_name": "blaPAO", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaPAO", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.497, "contig_id": "gnl|BUGS|ERR873305_60", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 1194.0, "reference_protein_length": null, "target_gene_length": 1194.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaVIM", "gene_name": "blaVIM-2", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaVIM-2", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 801.0, "reference_protein_length": null, "target_gene_length": 801.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "catB7", "gene_name": "catB7", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "catB7", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.531, "contig_id": "gnl|BUGS|ERR873305_63", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 639.0, "reference_protein_length": null, "target_gene_length": 639.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "crpP", "gene_name": "crpP", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "crpP", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 98.485, "contig_id": "gnl|BUGS|ERR873305_43", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 198.0, "reference_protein_length": null, "target_gene_length": 198.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "fosATR", "gene_name": "fosATR", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "fosATR", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 98.529, "contig_id": "gnl|BUGS|ERR873305_36", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 408.0, "reference_protein_length": null, "target_gene_length": 408.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "sul1", "gene_name": "sul1", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "sul1", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.885, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 867.0, "reference_protein_length": null, "target_gene_length": 867.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}], "resfinder.py: config 0": [{"input_file_name": "ERR873305", "gene_symbol": "aac(6\')-29a", "gene_name": "aac(6\')-29a_1_AF263519", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF263519", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 78.0, "query_stop_nt": 473.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 396.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Warning: gene is missing from Notes file. Please inform curator.", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaVIM-2", "gene_name": "blaVIM-2_1_AF302086", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF302086", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 604.0, "query_stop_nt": 1404.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 801.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Beta-lactam resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "catB7", "gene_name": "catB7_1_AF036933", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF036933", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873305_63", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 29357.0, "query_stop_nt": 29995.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 639.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Phenicol resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "crpP", "gene_name": "crpP_1_HM560971", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "HM560971", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873305_43", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 4289.0, "query_stop_nt": 4486.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 198.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Warning: gene is missing from Notes file. Please inform curator.", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "fosA", "gene_name": "fosA_4_ACWU01000146", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "ACWU01000146", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873305_36", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8371.0, "query_stop_nt": 8778.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 408.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Fosfomycin resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "sul1", "gene_name": "sul1_2_U12338", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "U12338", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 1932.0, "query_stop_nt": 2771.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 840.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Sulphonamide resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}], "staramr: config 0": [{"input_file_name": "ERR873305", "gene_symbol": "aac(6\')-29a", "gene_name": "aac(6\')-29a", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF263519", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 78.0, "query_stop_nt": 473.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "gentamicin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "aph(3\')-IIb", "gene_name": "aph(3\')-IIb", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "CP006832", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.26, "contig_id": "gnl|BUGS|ERR873305_93", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 13105.0, "query_stop_nt": 12299.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "kanamycin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaOXA-50", "gene_name": "blaOXA-50", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AY306133", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_15", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 19276.0, "query_stop_nt": 18488.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaPAO", "gene_name": "blaPAO", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AY083592", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.5, "contig_id": "gnl|BUGS|ERR873305_60", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 1282.0, "query_stop_nt": 89.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin, amoxicillin/clavulanic acid, cefoxitin, ceftriaxone", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "blaVIM-2", "gene_name": "blaVIM-2", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF302086", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 604.0, "query_stop_nt": 1404.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin, amoxicillin/clavulanic acid, cefoxitin, ceftriaxone, meropenem", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "catB7", "gene_name": "catB7", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF036933", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873305_63", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 29357.0, "query_stop_nt": 29995.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "chloramphenicol", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "crpP", "gene_name": "crpP", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "HM560971", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873305_43", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 4289.0, "query_stop_nt": 4486.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "unknown[crpP_1_HM560971]", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "fosA", "gene_name": "fosA", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "ACWU01000146", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873305_36", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8778.0, "query_stop_nt": 8371.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "fosfomycin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873305", "gene_symbol": "sul1", "gene_name": "sul1", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "U12338", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873305_223", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 1932.0, "query_stop_nt": 2771.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "sulfisoxazole", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}]}, {"input_file_name": "ERR873306", "abricate: config 0": [{"input_file_name": "ERR873306", "gene_symbol": "aac(6\')-29a", "gene_name": "aminoglycoside N-acetyltransferase AAC(6\')-29a", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_048575.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 9090.0, "query_stop_nt": 9491.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "aph(3\')-IIb", "gene_name": "aminoglycoside O-phosphotransferase APH(3\')-IIb", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047424.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 99.26, "contig_id": "gnl|BUGS|ERR873306_28", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 14225.0, "query_stop_nt": 15031.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "KANAMYCIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaOXA-395", "gene_name": "OXA-50 family oxacillin-hydrolyzing class D beta-lactamase OXA-395", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_049684.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_21", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 56773.0, "query_stop_nt": 57561.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaPDC-158", "gene_name": "class C beta-lactamase PDC-158", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_050831.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 75.51, "contig_id": "gnl|BUGS|ERR873306_157", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 438.0, "query_stop_nt": 1450.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 84.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CEPHALOSPORIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaPDC-55", "gene_name": "class C beta-lactamase PDC-55", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_049929.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.77, "contig_id": "gnl|BUGS|ERR873306_28", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 27345.0, "query_stop_nt": 28567.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CEPHALOSPORIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaVIM-2", "gene_name": "subclass B1 metallo-beta-lactamase VIM-2", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_050347.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8159.0, "query_stop_nt": 8959.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CARBAPENEM", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "catB7", "gene_name": "type B-4 chloramphenicol O-acetyltransferase CatB7", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047614.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873306_16", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 51123.0, "query_stop_nt": 51761.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CHLORAMPHENICOL", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "cmlB1", "gene_name": "chloramphenicol efflux MFS transporter CmlB1", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047658.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 79.5, "contig_id": "gnl|BUGS|ERR873306_56", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 15916.0, "query_stop_nt": 17125.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 95.58, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CHLORAMPHENICOL", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "crpP", "gene_name": "ciprofloxacin resistance protein CrpP", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_062203.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873306_10", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 3961.0, "query_stop_nt": 4158.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "FLUOROQUINOLONE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "fosA-354827590", "gene_name": "FosA family fosfomycin resistance glutathione transferase", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047883.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873306_5", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 20865.0, "query_stop_nt": 21272.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "FOSFOMYCIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "sul1", "gene_name": "sulfonamide-resistant dihydropteroate synthase Sul1", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_048082.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6792.0, "query_stop_nt": 7631.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "SULFONAMIDE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}], "amrfinderplus: config 0": [{"input_file_name": "ERR873306", "gene_symbol": "aac(6\')-29a", "gene_name": "aminoglycoside N-acetyltransferase AAC(6\')-29a", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_064190968.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 9093.0, "query_stop_nt": 9491.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 133.0, "reference_protein_length": null, "target_gene_length": 133.0, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": "AMINOGLYCOSIDE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "aph(3\')-IIb", "gene_name": "aminoglycoside O-phosphotransferase APH(3\')-IIb", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003113011.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 99.25, "contig_id": "gnl|BUGS|ERR873306_28", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 14225.0, "query_stop_nt": 15028.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 268.0, "reference_protein_length": null, "target_gene_length": 268.0, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": "KANAMYCIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaOXA-395", "gene_name": "OXA-50 family oxacillin-hydrolyzing class D beta-lactamase OXA-395", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_015649877.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_21", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 56773.0, "query_stop_nt": 57558.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 262.0, "reference_protein_length": null, "target_gene_length": 262.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "BETA-LACTAM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaPDC-3", "gene_name": "class C extended-spectrum beta-lactamase PDC-3", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003121934.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_28", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 27377.0, "query_stop_nt": 28567.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 397.0, "reference_protein_length": null, "target_gene_length": 397.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "CEPHALOSPORIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaVIM-2", "gene_name": "subclass B1 metallo-beta-lactamase VIM-2", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003108247.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8162.0, "query_stop_nt": 8959.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 266.0, "reference_protein_length": null, "target_gene_length": 266.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "CARBAPENEM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "catB7", "gene_name": "type B-4 chloramphenicol O-acetyltransferase CatB7", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003112709.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 99.06, "contig_id": "gnl|BUGS|ERR873306_16", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 51126.0, "query_stop_nt": 51761.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 212.0, "reference_protein_length": null, "target_gene_length": 212.0, "target_protein_length": null, "drug_class": "PHENICOL", "antimicrobial_agent": "CHLORAMPHENICOL", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "crpP", "gene_name": "ciprofloxacin resistance protein CrpP", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_033179079.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 98.46, "contig_id": "gnl|BUGS|ERR873306_10", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 3961.0, "query_stop_nt": 4155.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 65.0, "reference_protein_length": null, "target_gene_length": 65.0, "target_protein_length": null, "drug_class": "FLUOROQUINOLONE", "antimicrobial_agent": "FLUOROQUINOLONE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "fosA", "gene_name": "FosA family fosfomycin resistance glutathione transferase", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003082280.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 98.52, "contig_id": "gnl|BUGS|ERR873306_5", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 20868.0, "query_stop_nt": 21272.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 135.0, "reference_protein_length": null, "target_gene_length": 135.0, "target_protein_length": null, "drug_class": "FOSFOMYCIN", "antimicrobial_agent": "FOSFOMYCIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "qacEdelta1", "gene_name": "quaternary ammonium compound efflux SMR transporter QacE delta 1", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_000679427.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 7628.0, "query_stop_nt": 7972.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 115.0, "reference_protein_length": null, "target_gene_length": 115.0, "target_protein_length": null, "drug_class": "QUATERNARY AMMONIUM", "antimicrobial_agent": "QUATERNARY AMMONIUM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "sul1", "gene_name": "sulfonamide-resistant dihydropteroate synthase Sul1", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_000259031.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6795.0, "query_stop_nt": 7631.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 279.0, "reference_protein_length": null, "target_gene_length": 279.0, "target_protein_length": null, "drug_class": "SULFONAMIDE", "antimicrobial_agent": "SULFONAMIDE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}], "csstar: config 0": [{"input_file_name": "ERR873306", "gene_symbol": "aac(6\')", "gene_name": "aac(6\')-29a", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "aac(6\')-29a", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 396.0, "reference_protein_length": null, "target_gene_length": 396.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "aph(3\')", "gene_name": "aph(3\')-IIb", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "aph(3\')-IIb", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.257, "contig_id": "gnl|BUGS|ERR873306_28", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 807.0, "reference_protein_length": null, "target_gene_length": 807.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "bcr1", "gene_name": "bcr1", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "bcr1", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_102", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 1209.0, "reference_protein_length": null, "target_gene_length": 1209.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaOXA", "gene_name": "blaOXA-395", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaOXA-395", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_21", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 789.0, "reference_protein_length": null, "target_gene_length": 789.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaPAO", "gene_name": "blaPAO", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaPAO", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.497, "contig_id": "gnl|BUGS|ERR873306_28", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 1194.0, "reference_protein_length": null, "target_gene_length": 1194.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaVIM", "gene_name": "blaVIM-2", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaVIM-2", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 801.0, "reference_protein_length": null, "target_gene_length": 801.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "catB7", "gene_name": "catB7", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "catB7", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.531, "contig_id": "gnl|BUGS|ERR873306_16", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 639.0, "reference_protein_length": null, "target_gene_length": 639.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "crpP", "gene_name": "crpP", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "crpP", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 98.485, "contig_id": "gnl|BUGS|ERR873306_10", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 198.0, "reference_protein_length": null, "target_gene_length": 198.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "fosATR", "gene_name": "fosATR", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "fosATR", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 98.529, "contig_id": "gnl|BUGS|ERR873306_5", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 408.0, "reference_protein_length": null, "target_gene_length": 408.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "sul1", "gene_name": "sul1", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "sul1", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.885, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 867.0, "reference_protein_length": null, "target_gene_length": 867.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}], "resfinder.py: config 0": [{"input_file_name": "ERR873306", "gene_symbol": "aac(6\')-29a", "gene_name": "aac(6\')-29a_1_AF263519", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF263519", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 9090.0, "query_stop_nt": 9485.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 396.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Warning: gene is missing from Notes file. Please inform curator.", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaVIM-2", "gene_name": "blaVIM-2_1_AF302086", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF302086", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8159.0, "query_stop_nt": 8959.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 801.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Beta-lactam resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "catB7", "gene_name": "catB7_1_AF036933", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF036933", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873306_16", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 51123.0, "query_stop_nt": 51761.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 639.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Phenicol resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "crpP", "gene_name": "crpP_1_HM560971", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "HM560971", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873306_10", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 3961.0, "query_stop_nt": 4158.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 198.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Warning: gene is missing from Notes file. Please inform curator.", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "fosA", "gene_name": "fosA_4_ACWU01000146", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "ACWU01000146", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873306_5", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 20865.0, "query_stop_nt": 21272.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 408.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Fosfomycin resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "sul1", "gene_name": "sul1_2_U12338", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "U12338", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6792.0, "query_stop_nt": 7631.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 840.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Sulphonamide resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}], "staramr: config 0": [{"input_file_name": "ERR873306", "gene_symbol": "aac(6\')-29a", "gene_name": "aac(6\')-29a", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF263519", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 9485.0, "query_stop_nt": 9090.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "gentamicin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "aph(3\')-IIb", "gene_name": "aph(3\')-IIb", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "CP006832", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.26, "contig_id": "gnl|BUGS|ERR873306_28", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 14225.0, "query_stop_nt": 15031.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "kanamycin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaOXA-50", "gene_name": "blaOXA-50", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AY306133", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_21", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 56773.0, "query_stop_nt": 57561.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaPAO", "gene_name": "blaPAO", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AY083592", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.5, "contig_id": "gnl|BUGS|ERR873306_28", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 28567.0, "query_stop_nt": 27374.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin, amoxicillin/clavulanic acid, cefoxitin, ceftriaxone", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "blaVIM-2", "gene_name": "blaVIM-2", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF302086", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8959.0, "query_stop_nt": 8159.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin, amoxicillin/clavulanic acid, cefoxitin, ceftriaxone, meropenem", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "catB7", "gene_name": "catB7", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF036933", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873306_16", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 51761.0, "query_stop_nt": 51123.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "chloramphenicol", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "crpP", "gene_name": "crpP", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "HM560971", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873306_10", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 3961.0, "query_stop_nt": 4158.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "unknown[crpP_1_HM560971]", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "fosA", "gene_name": "fosA", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "ACWU01000146", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873306_5", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 21272.0, "query_stop_nt": 20865.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "fosfomycin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873306", "gene_symbol": "sul1", "gene_name": "sul1", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "U12338", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873306_117", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 7631.0, "query_stop_nt": 6792.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "sulfisoxazole", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}]}, {"input_file_name": "ERR873307", "abricate: config 0": [{"input_file_name": "ERR873307", "gene_symbol": "aac(6\')-29a", "gene_name": "aminoglycoside N-acetyltransferase AAC(6\')-29a", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_048575.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 9090.0, "query_stop_nt": 9491.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "aph(3\')-IIb", "gene_name": "aminoglycoside O-phosphotransferase APH(3\')-IIb", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047424.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 99.26, "contig_id": "gnl|BUGS|ERR873307_37", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 49533.0, "query_stop_nt": 50339.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "KANAMYCIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaOXA-395", "gene_name": "OXA-50 family oxacillin-hydrolyzing class D beta-lactamase OXA-395", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_049684.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_20", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 56757.0, "query_stop_nt": 57545.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaPDC-158", "gene_name": "class C beta-lactamase PDC-158", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_050831.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 75.51, "contig_id": "gnl|BUGS|ERR873307_180", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 438.0, "query_stop_nt": 1450.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 84.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CEPHALOSPORIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaPDC-55", "gene_name": "class C beta-lactamase PDC-55", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_049929.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.77, "contig_id": "gnl|BUGS|ERR873307_37", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 35997.0, "query_stop_nt": 37219.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CEPHALOSPORIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaVIM-2", "gene_name": "subclass B1 metallo-beta-lactamase VIM-2", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_050347.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8159.0, "query_stop_nt": 8959.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CARBAPENEM", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "catB7", "gene_name": "type B-4 chloramphenicol O-acetyltransferase CatB7", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047614.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873307_27", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 70629.0, "query_stop_nt": 71267.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CHLORAMPHENICOL", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "cmlB1", "gene_name": "chloramphenicol efflux MFS transporter CmlB1", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047658.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 79.5, "contig_id": "gnl|BUGS|ERR873307_61", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 15945.0, "query_stop_nt": 17154.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 95.58, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CHLORAMPHENICOL", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "crpP", "gene_name": "ciprofloxacin resistance protein CrpP", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_062203.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873307_7", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 145576.0, "query_stop_nt": 145773.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "FLUOROQUINOLONE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "fosA-354827590", "gene_name": "FosA family fosfomycin resistance glutathione transferase", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047883.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873307_8", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 109287.0, "query_stop_nt": 109694.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "FOSFOMYCIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "sul1", "gene_name": "sulfonamide-resistant dihydropteroate synthase Sul1", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_048082.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6792.0, "query_stop_nt": 7631.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "SULFONAMIDE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}], "amrfinderplus: config 0": [{"input_file_name": "ERR873307", "gene_symbol": "aac(6\')-29a", "gene_name": "aminoglycoside N-acetyltransferase AAC(6\')-29a", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_064190968.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 9093.0, "query_stop_nt": 9491.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 133.0, "reference_protein_length": null, "target_gene_length": 133.0, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": "AMINOGLYCOSIDE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "aph(3\')-IIb", "gene_name": "aminoglycoside O-phosphotransferase APH(3\')-IIb", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003113011.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 99.25, "contig_id": "gnl|BUGS|ERR873307_37", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 49536.0, "query_stop_nt": 50339.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 268.0, "reference_protein_length": null, "target_gene_length": 268.0, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": "KANAMYCIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaOXA-395", "gene_name": "OXA-50 family oxacillin-hydrolyzing class D beta-lactamase OXA-395", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_015649877.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_20", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 56757.0, "query_stop_nt": 57542.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 262.0, "reference_protein_length": null, "target_gene_length": 262.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "BETA-LACTAM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaPDC-3", "gene_name": "class C extended-spectrum beta-lactamase PDC-3", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003121934.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_37", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 35997.0, "query_stop_nt": 37187.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 397.0, "reference_protein_length": null, "target_gene_length": 397.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "CEPHALOSPORIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaVIM-2", "gene_name": "subclass B1 metallo-beta-lactamase VIM-2", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003108247.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8162.0, "query_stop_nt": 8959.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 266.0, "reference_protein_length": null, "target_gene_length": 266.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "CARBAPENEM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "catB7", "gene_name": "type B-4 chloramphenicol O-acetyltransferase CatB7", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003112709.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 99.06, "contig_id": "gnl|BUGS|ERR873307_27", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 70629.0, "query_stop_nt": 71264.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 212.0, "reference_protein_length": null, "target_gene_length": 212.0, "target_protein_length": null, "drug_class": "PHENICOL", "antimicrobial_agent": "CHLORAMPHENICOL", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "crpP", "gene_name": "ciprofloxacin resistance protein CrpP", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_033179079.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 98.46, "contig_id": "gnl|BUGS|ERR873307_7", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 145579.0, "query_stop_nt": 145773.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 65.0, "reference_protein_length": null, "target_gene_length": 65.0, "target_protein_length": null, "drug_class": "FLUOROQUINOLONE", "antimicrobial_agent": "FLUOROQUINOLONE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "fosA", "gene_name": "FosA family fosfomycin resistance glutathione transferase", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003082280.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 98.52, "contig_id": "gnl|BUGS|ERR873307_8", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 109287.0, "query_stop_nt": 109691.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 135.0, "reference_protein_length": null, "target_gene_length": 135.0, "target_protein_length": null, "drug_class": "FOSFOMYCIN", "antimicrobial_agent": "FOSFOMYCIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "qacEdelta1", "gene_name": "quaternary ammonium compound efflux SMR transporter QacE delta 1", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_000679427.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 7628.0, "query_stop_nt": 7972.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 115.0, "reference_protein_length": null, "target_gene_length": 115.0, "target_protein_length": null, "drug_class": "QUATERNARY AMMONIUM", "antimicrobial_agent": "QUATERNARY AMMONIUM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "sul1", "gene_name": "sulfonamide-resistant dihydropteroate synthase Sul1", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_000259031.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6795.0, "query_stop_nt": 7631.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 279.0, "reference_protein_length": null, "target_gene_length": 279.0, "target_protein_length": null, "drug_class": "SULFONAMIDE", "antimicrobial_agent": "SULFONAMIDE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}], "csstar: config 0": [{"input_file_name": "ERR873307", "gene_symbol": "aac(6\')", "gene_name": "aac(6\')-29a", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "aac(6\')-29a", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 396.0, "reference_protein_length": null, "target_gene_length": 396.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "aph(3\')", "gene_name": "aph(3\')-IIb", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "aph(3\')-IIb", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.257, "contig_id": "gnl|BUGS|ERR873307_37", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 807.0, "reference_protein_length": null, "target_gene_length": 807.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "bcr1", "gene_name": "bcr1", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "bcr1", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_112", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 1209.0, "reference_protein_length": null, "target_gene_length": 1209.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaOXA", "gene_name": "blaOXA-395", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaOXA-395", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_20", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 789.0, "reference_protein_length": null, "target_gene_length": 789.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaPAO", "gene_name": "blaPAO", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaPAO", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.497, "contig_id": "gnl|BUGS|ERR873307_37", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 1194.0, "reference_protein_length": null, "target_gene_length": 1194.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaVIM", "gene_name": "blaVIM-2", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "blaVIM-2", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 801.0, "reference_protein_length": null, "target_gene_length": 801.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "catB7", "gene_name": "catB7", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "catB7", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.531, "contig_id": "gnl|BUGS|ERR873307_27", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 639.0, "reference_protein_length": null, "target_gene_length": 639.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "crpP", "gene_name": "crpP", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "crpP", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 98.485, "contig_id": "gnl|BUGS|ERR873307_7", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 198.0, "reference_protein_length": null, "target_gene_length": 198.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "fosATR", "gene_name": "fosATR", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "fosATR", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 98.529, "contig_id": "gnl|BUGS|ERR873307_8", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 408.0, "reference_protein_length": null, "target_gene_length": 408.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "sul1", "gene_name": "sul1", "reference_database_id": "ResGANNOT", "reference_database_version": "2020-Nov-05", "reference_accession": "sul1", "analysis_software_name": "csstar", "analysis_software_version": "2.0.0", "sequence_identity": 99.885, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": null, "query_stop_nt": null, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": null, "coverage_ratio": null, "reference_gene_length": 867.0, "reference_protein_length": null, "target_gene_length": 867.0, "target_protein_length": null, "drug_class": null, "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "csstar: config 0"}], "resfinder.py: config 0": [{"input_file_name": "ERR873307", "gene_symbol": "aac(6\')-29a", "gene_name": "aac(6\')-29a_1_AF263519", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF263519", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 9090.0, "query_stop_nt": 9485.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 396.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Warning: gene is missing from Notes file. Please inform curator.", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaVIM-2", "gene_name": "blaVIM-2_1_AF302086", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF302086", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8159.0, "query_stop_nt": 8959.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 801.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Beta-lactam resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "catB7", "gene_name": "catB7_1_AF036933", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "AF036933", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873307_27", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 70629.0, "query_stop_nt": 71267.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 639.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Phenicol resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "crpP", "gene_name": "crpP_1_HM560971", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "HM560971", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873307_7", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 145576.0, "query_stop_nt": 145773.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 198.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Warning: gene is missing from Notes file. Please inform curator.", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "fosA", "gene_name": "fosA_4_ACWU01000146", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "ACWU01000146", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873307_8", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 109287.0, "query_stop_nt": 109694.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 408.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Fosfomycin resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "sul1", "gene_name": "sul1_2_U12338", "reference_database_id": "resfinder", "reference_database_version": "2a8dd7fc7a8c", "reference_accession": "U12338", "analysis_software_name": "resfinder.py", "analysis_software_version": 3.2, "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6792.0, "query_stop_nt": 7631.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 840.0, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "Sulphonamide resistance", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "resfinder.py: config 0"}], "staramr: config 0": [{"input_file_name": "ERR873307", "gene_symbol": "aac(6\')-29a", "gene_name": "aac(6\')-29a", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF263519", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 9485.0, "query_stop_nt": 9090.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "gentamicin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "aph(3\')-IIb", "gene_name": "aph(3\')-IIb", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "CP006832", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.26, "contig_id": "gnl|BUGS|ERR873307_37", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 50339.0, "query_stop_nt": 49533.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "kanamycin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaOXA-50", "gene_name": "blaOXA-50", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AY306133", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_20", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 56757.0, "query_stop_nt": 57545.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaPAO", "gene_name": "blaPAO", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AY083592", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.5, "contig_id": "gnl|BUGS|ERR873307_37", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 35997.0, "query_stop_nt": 37190.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin, amoxicillin/clavulanic acid, cefoxitin, ceftriaxone", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "blaVIM-2", "gene_name": "blaVIM-2", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF302086", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 8959.0, "query_stop_nt": 8159.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "ampicillin, amoxicillin/clavulanic acid, cefoxitin, ceftriaxone, meropenem", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "catB7", "gene_name": "catB7", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "AF036933", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873307_27", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 70629.0, "query_stop_nt": 71267.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "chloramphenicol", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "crpP", "gene_name": "crpP", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "HM560971", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873307_7", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 145773.0, "query_stop_nt": 145576.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "unknown[crpP_1_HM560971]", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "fosA", "gene_name": "fosA", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "ACWU01000146", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873307_8", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 109287.0, "query_stop_nt": 109694.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "fosfomycin", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}, {"input_file_name": "ERR873307", "gene_symbol": "sul1", "gene_name": "sul1", "reference_database_id": "resfinder", "reference_database_version": "e8f1eb2585cd9610c4034a54ce7fc4f93aa95535", "reference_accession": "U12338", "analysis_software_name": "staramr", "analysis_software_version": "0.7.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873307_125", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 7631.0, "query_stop_nt": 6792.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": null, "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": 1.0, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "sulfisoxazole", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "staramr: config 0"}]}, {"input_file_name": "ERR873308", "abricate: config 0": [{"input_file_name": "ERR873308", "gene_symbol": "aac(6\')-29b", "gene_name": "aminoglycoside N-acetyltransferase AAC(6\')-29b", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_048576.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_120", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 5561.0, "query_stop_nt": 5962.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "aph(3\')-IIb", "gene_name": "aminoglycoside O-phosphotransferase APH(3\')-IIb", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047424.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 99.26, "contig_id": "gnl|BUGS|ERR873308_38", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 14272.0, "query_stop_nt": 15078.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "KANAMYCIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "blaOXA-395", "gene_name": "OXA-50 family oxacillin-hydrolyzing class D beta-lactamase OXA-395", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_049684.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_20", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 36358.0, "query_stop_nt": 37146.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "blaPDC-158", "gene_name": "class C beta-lactamase PDC-158", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_050831.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 75.51, "contig_id": "gnl|BUGS|ERR873308_155", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 438.0, "query_stop_nt": 1450.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 84.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CEPHALOSPORIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "blaPDC-55", "gene_name": "class C beta-lactamase PDC-55", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_049929.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.77, "contig_id": "gnl|BUGS|ERR873308_38", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 27392.0, "query_stop_nt": 28614.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CEPHALOSPORIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "blaVIM-2", "gene_name": "subclass B1 metallo-beta-lactamase VIM-2", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_050347.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_168", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 85.0, "query_stop_nt": 885.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CARBAPENEM", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "catB7", "gene_name": "type B-4 chloramphenicol O-acetyltransferase CatB7", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047614.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 99.53, "contig_id": "gnl|BUGS|ERR873308_13", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 51113.0, "query_stop_nt": 51751.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CHLORAMPHENICOL", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "cmlB1", "gene_name": "chloramphenicol efflux MFS transporter CmlB1", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047658.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 79.5, "contig_id": "gnl|BUGS|ERR873308_34", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 48285.0, "query_stop_nt": 49494.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 95.58, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "CHLORAMPHENICOL", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "crpP", "gene_name": "ciprofloxacin resistance protein CrpP", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_062203.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.48, "contig_id": "gnl|BUGS|ERR873308_6", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 3991.0, "query_stop_nt": 4188.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "FLUOROQUINOLONE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "fosA-354827590", "gene_name": "FosA family fosfomycin resistance glutathione transferase", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_047883.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 98.53, "contig_id": "gnl|BUGS|ERR873308_40", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 20865.0, "query_stop_nt": 21272.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "FOSFOMYCIN", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "sul1", "gene_name": "sulfonamide-resistant dihydropteroate synthase Sul1", "reference_database_id": "ncbi", "reference_database_version": "2020-Apr-19", "reference_accession": "NG_048082.1", "analysis_software_name": "abricate", "analysis_software_version": "1.0.1", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_110", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6792.0, "query_stop_nt": 7631.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": null, "reference_protein_length": null, "target_gene_length": null, "target_protein_length": null, "drug_class": "SULFONAMIDE", "antimicrobial_agent": null, "resistance_mechanism": null, "config_display_name": "abricate: config 0"}], "amrfinderplus: config 0": [{"input_file_name": "ERR873308", "gene_symbol": "aac(6\')-29b", "gene_name": "aminoglycoside N-acetyltransferase AAC(6\')-29b", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_064190969.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_120", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 5561.0, "query_stop_nt": 5959.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 133.0, "reference_protein_length": null, "target_gene_length": 133.0, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": "AMINOGLYCOSIDE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "aph(3\')-IIb", "gene_name": "aminoglycoside O-phosphotransferase APH(3\')-IIb", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003113011.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 99.25, "contig_id": "gnl|BUGS|ERR873308_38", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 14272.0, "query_stop_nt": 15075.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 268.0, "reference_protein_length": null, "target_gene_length": 268.0, "target_protein_length": null, "drug_class": "AMINOGLYCOSIDE", "antimicrobial_agent": "KANAMYCIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "blaOXA-395", "gene_name": "OXA-50 family oxacillin-hydrolyzing class D beta-lactamase OXA-395", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_015649877.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_20", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 36361.0, "query_stop_nt": 37146.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 262.0, "reference_protein_length": null, "target_gene_length": 262.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "BETA-LACTAM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "blaPDC-3", "gene_name": "class C extended-spectrum beta-lactamase PDC-3", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003121934.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_38", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 27424.0, "query_stop_nt": 28614.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 397.0, "reference_protein_length": null, "target_gene_length": 397.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "CEPHALOSPORIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "blaVIM-2", "gene_name": "subclass B1 metallo-beta-lactamase VIM-2", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003108247.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_168", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 85.0, "query_stop_nt": 882.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 266.0, "reference_protein_length": null, "target_gene_length": 266.0, "target_protein_length": null, "drug_class": "BETA-LACTAM", "antimicrobial_agent": "CARBAPENEM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "catB7", "gene_name": "type B-4 chloramphenicol O-acetyltransferase CatB7", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003112709.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 99.06, "contig_id": "gnl|BUGS|ERR873308_13", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 51116.0, "query_stop_nt": 51751.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 212.0, "reference_protein_length": null, "target_gene_length": 212.0, "target_protein_length": null, "drug_class": "PHENICOL", "antimicrobial_agent": "CHLORAMPHENICOL", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "crpP", "gene_name": "ciprofloxacin resistance protein CrpP", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_033179079.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 98.46, "contig_id": "gnl|BUGS|ERR873308_6", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 3991.0, "query_stop_nt": 4185.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "+", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 65.0, "reference_protein_length": null, "target_gene_length": 65.0, "target_protein_length": null, "drug_class": "FLUOROQUINOLONE", "antimicrobial_agent": "FLUOROQUINOLONE", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "fosA", "gene_name": "FosA family fosfomycin resistance glutathione transferase", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_003082280.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 98.52, "contig_id": "gnl|BUGS|ERR873308_40", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 20868.0, "query_stop_nt": 21272.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 135.0, "reference_protein_length": null, "target_gene_length": 135.0, "target_protein_length": null, "drug_class": "FOSFOMYCIN", "antimicrobial_agent": "FOSFOMYCIN", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "qacEdelta1", "gene_name": "quaternary ammonium compound efflux SMR transporter QacE delta 1", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_000679427.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_110", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 7628.0, "query_stop_nt": 7972.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "coverage_depth": null, "coverage_percentage": 100.0, "coverage_ratio": null, "reference_gene_length": 115.0, "reference_protein_length": null, "target_gene_length": 115.0, "target_protein_length": null, "drug_class": "QUATERNARY AMMONIUM", "antimicrobial_agent": "QUATERNARY AMMONIUM", "resistance_mechanism": null, "config_display_name": "amrfinderplus: config 0"}, {"input_file_name": "ERR873308", "gene_symbol": "sul1", "gene_name": "sulfonamide-resistant dihydropteroate synthase Sul1", "reference_database_id": "NCBI Reference Gene Database", "reference_database_version": "2020-03-20.1", "reference_accession": "WP_000259031.1", "analysis_software_name": "amrfinderplus", "analysis_software_version": "3.6.10", "sequence_identity": 100.0, "contig_id": "gnl|BUGS|ERR873308_110", "query_start_aa": null, "query_stop_aa": null, "query_start_nt": 6795.0, "query_stop_nt": 7631.0, "subject_start_aa": null, "subject_stop_aa": null, "subject_start_nt": null, "subject_stop_nt": null, "strand_orientation": "-", "c
gitextract_2sm_t59v/
├── .dockerignore
├── .github/
│ ├── ISSUE_TEMPLATE/
│ │ └── bug_report.md
│ └── workflows/
│ ├── dockerpublish.yml
│ ├── python_publish.yml
│ └── test_package.yml
├── .gitignore
├── Dockerfile
├── LICENSE.txt
├── README.md
├── docs/
│ ├── hAMRonization_specification_details.csv
│ ├── interactive_report_demo.html
│ └── subgrant/
│ └── PHA4GE_AMR_SubGrant_Orientation.pptx
├── hAMRonization/
│ ├── AbricateIO.py
│ ├── AmrFinderPlusIO.py
│ ├── AmrPlusPlusIO.py
│ ├── AribaIO.py
│ ├── CSStarIO.py
│ ├── DeepArgIO.py
│ ├── FARGeneIO.py
│ ├── GrootIO.py
│ ├── Interfaces.py
│ ├── KmerResistanceIO.py
│ ├── MykrobeIO.py
│ ├── README.md
│ ├── ResFamsIO.py
│ ├── ResFinderIO.py
│ ├── RgiIO.py
│ ├── SraxIO.py
│ ├── Srst2IO.py
│ ├── StarAmrIO.py
│ ├── TBProfilerIO.py
│ ├── __init__.py
│ ├── constants.py
│ ├── hAMRonizedResult.py
│ ├── hamronize.py
│ └── summarize.py
├── schema/
│ ├── PHA4GE AMR Gene & Variant Specification.csv
│ ├── PHA4GE AMR Gene & Variant Specification.json
│ └── csv2json.py
├── setup.cfg
├── setup.py
└── test/
├── .gitignore
├── README.md
├── __init__.py
├── data/
│ ├── dummy/
│ │ ├── abricate/
│ │ │ └── report.tsv
│ │ ├── amrfinderplus/
│ │ │ └── report.tsv
│ │ ├── amrplusplus/
│ │ │ └── gene.tsv
│ │ ├── ariba/
│ │ │ ├── report.tsv
│ │ │ └── report_var.tsv
│ │ ├── deepARG/
│ │ │ ├── output.mapping.ARG.
│ │ │ └── output.mapping.potential.ARG
│ │ ├── fargene/
│ │ │ └── retrieved-genes-class_A-hmmsearched.out
│ │ ├── groot/
│ │ │ └── groot_report.tsv
│ │ ├── kmerresistance/
│ │ │ └── results.res
│ │ ├── mykrobe/
│ │ │ ├── empty.json
│ │ │ └── mykrobe.json
│ │ ├── pointfinder/
│ │ │ └── PointFinder_results.txt
│ │ ├── resfams/
│ │ │ └── resfams.tblout
│ │ ├── resfinder/
│ │ │ └── data_resfinder.json
│ │ ├── rgi/
│ │ │ ├── rgi.txt
│ │ │ ├── rgi_orf.txt
│ │ │ └── rgi_var.txt
│ │ ├── srax/
│ │ │ └── sraX_detected_ARGs.tsv
│ │ ├── srst2/
│ │ │ └── report.tsv
│ │ ├── sstar/
│ │ │ └── report.tsv
│ │ ├── staramr/
│ │ │ ├── pointfinder.tsv
│ │ │ └── resfinder.tsv
│ │ └── tbprofiler/
│ │ └── tbprofiler.json
│ └── raw_outputs/
│ ├── abricate/
│ │ └── report.tsv
│ ├── amrfinderplus/
│ │ ├── afp_non_coding.tsv
│ │ ├── empty_report_with_header.tsv
│ │ ├── hamronized_non_coding.tsv
│ │ ├── non_coding_contig.fasta
│ │ ├── report_nucleotide.tsv
│ │ └── report_protein.tsv
│ ├── amrplusplus/
│ │ └── gene.tsv
│ ├── ariba/
│ │ └── report.tsv
│ ├── deeparg/
│ │ ├── output.mapping.ARG
│ │ └── output.mapping.potential.ARG
│ ├── fargene/
│ │ └── retrieved-genes-class_A-hmmsearched.out
│ ├── groot/
│ │ └── report.tsv
│ ├── kmerresistance/
│ │ └── results.res
│ ├── mykrobe/
│ │ └── report.json
│ ├── pointfinder/
│ │ └── PointFinder_results.txt
│ ├── resfams/
│ │ └── resfams.tblout
│ ├── resfinder/
│ │ ├── ResFinder_results_tab.txt
│ │ └── data_resfinder.json
│ ├── rgi/
│ │ └── rgi.txt
│ ├── rgibwt/
│ │ └── Kp11_bwtoutput.gene_mapping_data.txt
│ ├── srax/
│ │ └── sraX_detected_ARGs.tsv
│ ├── srst2/
│ │ └── SAMN13064234_srst2_report.tsv
│ ├── sstar/
│ │ └── report.tsv
│ ├── staramr/
│ │ ├── pointfinder.tsv
│ │ └── resfinder.tsv
│ └── tbprofiler/
│ └── tbprofiler.json
├── run_integration_test.sh
├── test_interfaces.py
└── test_parsing_validity.py
SYMBOL INDEX (107 symbols across 25 files)
FILE: hAMRonization/AbricateIO.py
class AbricateIterator (line 10) | class AbricateIterator(hAMRonizedResultIterator):
method __init__ (line 11) | def __init__(self, source, metadata):
method parse (line 36) | def parse(self, handle):
FILE: hAMRonization/AmrFinderPlusIO.py
class AmrFinderPlusIterator (line 20) | class AmrFinderPlusIterator(hAMRonizedResultIterator):
method __init__ (line 61) | def __init__(self, source, metadata):
method parse (line 72) | def parse(self, handle):
FILE: hAMRonization/AmrPlusPlusIO.py
class AmrPlusPlusIterator (line 14) | class AmrPlusPlusIterator(hAMRonizedResultIterator):
method __init__ (line 15) | def __init__(self, source, metadata):
method parse (line 35) | def parse(self, handle):
FILE: hAMRonization/AribaIO.py
class AribaIterator (line 20) | class AribaIterator(hAMRonizedResultIterator):
method __init__ (line 21) | def __init__(self, source, metadata):
method parse (line 63) | def parse(self, handle):
FILE: hAMRonization/CSStarIO.py
class CSStarIterator (line 16) | class CSStarIterator(hAMRonizedResultIterator):
method __init__ (line 17) | def __init__(self, source, metadata):
method parse (line 34) | def parse(self, handle):
FILE: hAMRonization/DeepArgIO.py
class DeepArgIterator (line 14) | class DeepArgIterator(hAMRonizedResultIterator):
method __init__ (line 15) | def __init__(self, source, metadata):
method parse (line 41) | def parse(self, handle):
FILE: hAMRonization/FARGeneIO.py
class FARGeneIOIterator (line 15) | class FARGeneIOIterator(hAMRonizedResultIterator):
method __init__ (line 16) | def __init__(self, source, metadata):
method parse (line 56) | def parse(self, handle):
FILE: hAMRonization/GrootIO.py
class GrootIterator (line 16) | class GrootIterator(hAMRonizedResultIterator):
method __init__ (line 17) | def __init__(self, source, metadata):
method parse (line 36) | def parse(self, handle):
FILE: hAMRonization/Interfaces.py
class hAMRonizedResultIterator (line 15) | class hAMRonizedResultIterator(ABC):
method __init__ (line 23) | def __init__(self, source, field_map, metadata):
method hAMRonize (line 59) | def hAMRonize(self, report_data, metadata, field_map_override=None):
method __next__ (line 78) | def __next__(self):
method __iter__ (line 85) | def __iter__(self):
method parse (line 94) | def parse(self, handle):
method write (line 99) | def write(
function generate_tool_subparser (line 190) | def generate_tool_subparser(subparser, analysis_tool):
function generic_cli_interface (line 224) | def generic_cli_interface():
FILE: hAMRonization/KmerResistanceIO.py
class KmerResistanceIterator (line 14) | class KmerResistanceIterator(hAMRonizedResultIterator):
method __init__ (line 15) | def __init__(self, source, metadata):
method parse (line 42) | def parse(self, handle):
FILE: hAMRonization/MykrobeIO.py
class MykrobeIterator (line 11) | class MykrobeIterator(hAMRonizedResultIterator):
method __init__ (line 42) | def __init__(self, source, metadata):
method parse (line 67) | def parse(self, handle):
FILE: hAMRonization/ResFamsIO.py
class ResFamsIterator (line 14) | class ResFamsIterator(hAMRonizedResultIterator):
method __init__ (line 15) | def __init__(self, source, metadata):
method parse (line 51) | def parse(self, handle):
FILE: hAMRonization/ResFinderIO.py
class ResFinderIterator (line 20) | class ResFinderIterator(hAMRonizedResultIterator):
method __init__ (line 26) | def __init__(self, source, metadata):
method parse (line 32) | def parse(self, handle):
function _get_start_pos (line 201) | def _get_start_pos(p0, p1):
function _get_end_pos (line 205) | def _get_end_pos(p0, p1):
function _get_strand (line 209) | def _get_strand(p0, p1):
function _get_length (line 213) | def _get_length(p0, p1):
function _safe_round (line 217) | def _safe_round(n, d):
function _empty_to_none (line 221) | def _empty_to_none(s):
function _condense_notes (line 225) | def _condense_notes(notes, pmids):
function fold (line 235) | def fold(fun, acc, lst): # Python's missing function (why have map but ...
FILE: hAMRonization/RgiIO.py
class RgiIterator (line 20) | class RgiIterator(hAMRonizedResultIterator):
method __init__ (line 21) | def __init__(self, source, metadata):
method parse (line 110) | def parse(self, handle):
FILE: hAMRonization/SraxIO.py
class SraxIterator (line 15) | class SraxIterator(hAMRonizedResultIterator):
method __init__ (line 16) | def __init__(self, source, metadata):
method parse (line 35) | def parse(self, handle):
FILE: hAMRonization/Srst2IO.py
class Srst2Iterator (line 14) | class Srst2Iterator(hAMRonizedResultIterator):
method __init__ (line 15) | def __init__(self, source, metadata):
method parse (line 40) | def parse(self, handle):
FILE: hAMRonization/StarAmrIO.py
class StarAmrIterator (line 14) | class StarAmrIterator(hAMRonizedResultIterator):
method __init__ (line 15) | def __init__(self, source, metadata):
method parse (line 41) | def parse(self, handle):
FILE: hAMRonization/TBProfilerIO.py
class TBProfilerIterator (line 10) | class TBProfilerIterator(hAMRonizedResultIterator):
method __init__ (line 11) | def __init__(self, source, metadata):
method parse (line 33) | def parse(self, handle):
FILE: hAMRonization/__init__.py
function parse (line 85) | def parse(handle, metadata, tool):
FILE: hAMRonization/hAMRonizedResult.py
class hAMRonizedResult (line 8) | class hAMRonizedResult:
method __post_init__ (line 54) | def __post_init__(self):
FILE: hAMRonization/hamronize.py
function main (line 6) | def main():
FILE: hAMRonization/summarize.py
function format_interactive_json (line 13) | def format_interactive_json(combined_records):
function generate_interactive_report (line 75) | def generate_interactive_report(combined_report_data):
function check_report_type (line 676) | def check_report_type(file_path):
function summarize_reports (line 694) | def summarize_reports(report_paths, summary_type, output_path=None):
FILE: schema/csv2json.py
function string_list_to_list (line 15) | def string_list_to_list(string):
function interface_label_to_property_key (line 21) | def interface_label_to_property_key(interface_label):
function parse_properties_table (line 28) | def parse_properties_table(path_to_properties_table):
function get_required_fields (line 129) | def get_required_fields(path_to_properties_table):
function main (line 141) | def main(args):
FILE: test/test_interfaces.py
function not_raises (line 11) | def not_raises(exception, msg):
function test_empty_input_report (line 18) | def test_empty_input_report():
function test_summarize_empty_reports (line 51) | def test_summarize_empty_reports():
FILE: test/test_parsing_validity.py
function not_raises (line 7) | def not_raises(exception, msg):
function test_abricate (line 14) | def test_abricate():
function test_amrfinderplus (line 65) | def test_amrfinderplus():
function test_amrplusplus (line 116) | def test_amrplusplus():
function test_ariba (line 164) | def test_ariba():
function test_ariba_var (line 213) | def test_ariba_var():
function test_kmerresistance (line 247) | def test_kmerresistance():
function test_resfinder (line 295) | def test_resfinder():
function test_rgi_variants (line 418) | def test_rgi_variants():
function test_rgi_orf_mode (line 448) | def test_rgi_orf_mode():
function test_rgi (line 482) | def test_rgi():
function test_srax (line 533) | def test_srax():
function test_groot (line 585) | def test_groot():
function test_deeparg (line 636) | def test_deeparg():
function test_srst2 (line 684) | def test_srst2():
function test_csstar (line 732) | def test_csstar():
function test_tbprofiler (line 781) | def test_tbprofiler():
function test_mykrobe_empty (line 831) | def test_mykrobe_empty():
function test_mykrobe (line 838) | def test_mykrobe():
function test_staramr_resfinder (line 889) | def test_staramr_resfinder():
function test_staramr_pointfinder (line 916) | def test_staramr_pointfinder():
function test_resfams (line 945) | def test_resfams():
function test_fargene (line 965) | def test_fargene():
Copy disabled (too large)
Download .json
Condensed preview — 98 files, each showing path, character count, and a content snippet. Download the .json file for the full structured content (13,446K chars).
[
{
"path": ".dockerignore",
"chars": 52,
"preview": ".*\n*.egg-info\n*.pyc\n/docs\n/test\n/schema\n__pycache__\n"
},
{
"path": ".github/ISSUE_TEMPLATE/bug_report.md",
"chars": 969,
"preview": "---\nname: Bug report\nabout: Create a report to help us improve\ntitle: \"[BUG]\"\nlabels: bug\nassignees: ''\n\n---\n\n**Describe"
},
{
"path": ".github/workflows/dockerpublish.yml",
"chars": 854,
"preview": "name: Publish Docker Image\n\non:\n release:\n types: [published]\n\njobs:\n main:\n runs-on: ubuntu-latest\n steps:\n "
},
{
"path": ".github/workflows/python_publish.yml",
"chars": 1019,
"preview": "# This workflow will upload a Python Package using Twine when a release is created\n# For more information see: https://h"
},
{
"path": ".github/workflows/test_package.yml",
"chars": 1770,
"preview": "#This workflow will install Python dependencies, run tests and lint with a variety of Python versions\n# For more informa"
},
{
"path": ".gitignore",
"chars": 38,
"preview": ".DS_Store\n*.egg-info\n*.pyc\nbuild\ndist\n"
},
{
"path": "Dockerfile",
"chars": 1302,
"preview": "# base image\nFROM docker.io/library/python:3.14.4-alpine3.23\n\n# metadata\nLABEL org.opencontainers.image.version=1.2.1\nLA"
},
{
"path": "LICENSE.txt",
"chars": 7653,
"preview": " GNU LESSER GENERAL PUBLIC LICENSE\n Version 3, 29 June 2007\n\n Copyright (C) 2007"
},
{
"path": "README.md",
"chars": 16937,
"preview": "\n[:\n hAMRonization.Interfaces.generic_cli_interface()\n\n\nif __n"
},
{
"path": "hAMRonization/summarize.py",
"chars": 32432,
"preview": "#!/usr/bin/env python\n\nimport csv\nimport pandas as pd\nimport os\nimport sys\nimport json\nfrom .hAMRonizedResult import hAM"
},
{
"path": "schema/PHA4GE AMR Gene & Variant Specification.csv",
"chars": 10763,
"preview": "Interface Label,Required/Optional,Definition,Ontology,Value Type,Example,Guidance,Values\r\nAnalysis Software Name,Require"
},
{
"path": "schema/PHA4GE AMR Gene & Variant Specification.json",
"chars": 12132,
"preview": "{\n \"$schema\": \"http://json-schema.org/draft/2019-09/schema#\",\n \"version\": \"2022-05-13T15:07:33.790091\",\n \"type\""
},
{
"path": "schema/csv2json.py",
"chars": 5202,
"preview": "#!/usr/bin/env python\n\nimport argparse\nimport csv\nimport json\nimport re\nfrom datetime import datetime\nfrom ast import li"
},
{
"path": "setup.cfg",
"chars": 229,
"preview": "[metadata]\n# This includes the license file(s) in the wheel.\n# # https://wheel.readthedocs.io/en/stable/user_guide.html#"
},
{
"path": "setup.py",
"chars": 1943,
"preview": "import re\nfrom distutils.core import setup\n\nwith open('README.md') as fh:\n long_description = fh.read()\n\nwith open('h"
},
{
"path": "test/.gitignore",
"chars": 78,
"preview": "/hamronized_*.json\n/hamronized_*.tsv\n/summary.html\n/summary.json\n/summary.tsv\n"
},
{
"path": "test/README.md",
"chars": 1772,
"preview": "# hARMonization Testing\n\nTo ensure correct functionality and reproducibly of the parsing modules implemented in hAMRoniz"
},
{
"path": "test/__init__.py",
"chars": 0,
"preview": ""
},
{
"path": "test/data/dummy/abricate/report.tsv",
"chars": 302,
"preview": "#FILE\tSEQUENCE\tSTART\tEND\tSTRAND\tGENE\tCOVERAGE\tCOVERAGE_MAP\tGAPS\t%COVERAGE\t%IDENTITY\tDATABASE\tACCESSION\tPRODUCT\tRESISTANC"
},
{
"path": "test/data/dummy/amrfinderplus/report.tsv",
"chars": 570,
"preview": "Protein id\tContig id\tStart\tStop\tStrand\tElement symbol\tElement name\tScope\tType\tSubtype\tClass\tSubclass\tMethod\tTarget lengt"
},
{
"path": "test/data/dummy/amrplusplus/gene.tsv",
"chars": 140,
"preview": "Sample\tGene\tHits\tGene Fraction\nDummy\tMEG_4334|Multi-compound|Drug_and_biocide_resistance|Drug_and_biocide_RND_efflux_pum"
},
{
"path": "test/data/dummy/ariba/report.tsv",
"chars": 607,
"preview": "#ariba_ref_name\tref_name\tgene\tvar_only\tflag\treads\tcluster\tref_len\tref_base_assembled\tpc_ident\tctg\tctg_len\tctg_cov\tknown_"
},
{
"path": "test/data/dummy/ariba/report_var.tsv",
"chars": 769,
"preview": "#ariba_ref_name\tref_name\tgene\tvar_only\tflag\treads\tcluster\tref_len\tref_base_assembled\tpc_ident\tctg\tctg_len\tctg_cov\tknown_"
},
{
"path": "test/data/dummy/deepARG/output.mapping.ARG.",
"chars": 287,
"preview": "#ARG\tquery-start\tquery-end\tread_id\tpredicted_ARG-class\tbest-hit\tprobability\tidentity\talignment-length\talignment-bitscore"
},
{
"path": "test/data/dummy/deepARG/output.mapping.potential.ARG",
"chars": 972609,
"preview": "#ARG\tquery-start\tquery-end\tread_id\tpredicted_ARG-class\tbest-hit\tprobability\tidentity\talignment-length\talignment-bitscore"
},
{
"path": "test/data/dummy/fargene/retrieved-genes-class_A-hmmsearched.out",
"chars": 1600,
"preview": "# --- fu"
},
{
"path": "test/data/dummy/groot/groot_report.tsv",
"chars": 60,
"preview": "OqxA.3003470.EU370913.4407527-4408202.4553\t266\t1176\t3D648M6D"
},
{
"path": "test/data/dummy/kmerresistance/results.res",
"chars": 235,
"preview": "#Template\tScore\tExpected\tTemplate_length\tTemplate_Identity\tTemplate_Coverage\tQuery_Identity\tQuery_Coverage\tDepth\tq_value"
},
{
"path": "test/data/dummy/mykrobe/empty.json",
"chars": 3,
"preview": "{}\n"
},
{
"path": "test/data/dummy/mykrobe/mykrobe.json",
"chars": 12350,
"preview": "{\n \"SRR6916544\": {\n \"susceptibility\": {\n \"Rifampicin\": {\n \"predict\": \"r\",\n "
},
{
"path": "test/data/dummy/pointfinder/PointFinder_results.txt",
"chars": 141,
"preview": "Mutation\tNucleotide change\tAmino acid change\tResistance\tPMID\ngyrA p.G81D\tGGT -> GAT\tG -> D\tCiprofloxacin,Nalidixic acid,"
},
{
"path": "test/data/dummy/resfams/resfams.tblout",
"chars": 1237,
"preview": "# --- full sequence ---- --- best 1 domai"
},
{
"path": "test/data/dummy/resfinder/data_resfinder.json",
"chars": 43958,
"preview": "{\"type\": \"software_result\", \"databases\": {\"ResFinder-2.4.0\": {\"type\": \"database\", \"database_name\": \"ResFinder\", \"databas"
},
{
"path": "test/data/dummy/rgi/rgi.txt",
"chars": 2862,
"preview": "ORF_ID\tContig\tStart\tStop\tOrientation\tCut_Off\tPass_Bitscore\tBest_Hit_Bitscore\tBest_Hit_ARO\tBest_Identities\tARO\tModel_type"
},
{
"path": "test/data/dummy/rgi/rgi_orf.txt",
"chars": 2066,
"preview": "ORF_ID\tContig\tStart\tStop\tOrientation\tCut_Off\tPass_Bitscore\tBest_Hit_Bitscore\tBest_Hit_ARO\tBest_Identities\tARO\tModel_type"
},
{
"path": "test/data/dummy/rgi/rgi_var.txt",
"chars": 2291,
"preview": "ORF_ID\tContig\tStart\tStop\tOrientation\tCut_Off\tPass_Bitscore\tBest_Hit_Bitscore\tBest_Hit_ARO\tBest_Identities\tARO\tModel_type"
},
{
"path": "test/data/dummy/srax/sraX_detected_ARGs.tsv",
"chars": 236,
"preview": "Locus ID\t# Sequences\tARG\tCoverage (%)\tIdentity (%)\tDrug class\tGene accession ID\tGene description\tAMR detection model\n6\t1"
},
{
"path": "test/data/dummy/srst2/report.tsv",
"chars": 216,
"preview": "Sample\tDB\tgene\tallele\tcoverage\tdepth\tdiffs\tuncertainty\tdivergence\tlength\tmaxMAF\tclusterid\tseqid\tannotation\nDummy\tResFind"
},
{
"path": "test/data/dummy/sstar/report.tsv",
"chars": 44,
"preview": "oqxA*\toqxA*\tNZ_LR792628.1\t99.575%\t1176\t1176\n"
},
{
"path": "test/data/dummy/staramr/pointfinder.tsv",
"chars": 252,
"preview": "Isolate ID\tGene\tPredicted Phenotype\tType\tPosition\tMutation\t%Identity\t%Overlap\tHSP Length/Total Length\tContig\tStart\tEnd\nS"
},
{
"path": "test/data/dummy/staramr/resfinder.tsv",
"chars": 1408,
"preview": "Isolate ID\tGene\tPredicted Phenotype\t%Identity\t%Overlap\tHSP Length/Total Length\tContig\tStart\tEnd\tAccession\tSequence\nGCF_9"
},
{
"path": "test/data/dummy/tbprofiler/tbprofiler.json",
"chars": 2053,
"preview": "{\n \"qc\": {\n \"pct_reads_mapped\": 97.16,\n \"num_reads_mapped\": 4232866,\n \"median_coverage\": 121,\n \"gene_covera"
},
{
"path": "test/data/raw_outputs/abricate/report.tsv",
"chars": 1464,
"preview": "#FILE\tSEQUENCE\tSTART\tEND\tSTRAND\tGENE\tCOVERAGE\tCOVERAGE_MAP\tGAPS\t%COVERAGE\t%IDENTITY\tDATABASE\tACCESSION\tPRODUCT\tRESISTANC"
},
{
"path": "test/data/raw_outputs/amrfinderplus/afp_non_coding.tsv",
"chars": 517,
"preview": "Protein id\tContig id\tStart\tStop\tStrand\tElement symbol\tElement name\tScope\tType\tSubtype\tClass\tSubclass\tMethod\tTarget lengt"
},
{
"path": "test/data/raw_outputs/amrfinderplus/empty_report_with_header.tsv",
"chars": 309,
"preview": "Protein id\tContig id\tStart\tStop\tStrand\tElement symbol\tElement name\tScope\tType\tSubtype\tClass\tSubclass\tMethod\tTarget lengt"
},
{
"path": "test/data/raw_outputs/amrfinderplus/hamronized_non_coding.tsv",
"chars": 1061,
"preview": "input_file_name\tgene_symbol\tgene_name\treference_database_name\treference_database_version\treference_accession\tanalysis_so"
},
{
"path": "test/data/raw_outputs/amrfinderplus/non_coding_contig.fasta",
"chars": 3084,
"preview": ">DAWXTK010000082.1:68-2970 TPA_asm: Neisseria gonorrhoeae strain WHO_V SAMEA3905804-rid23293863.denovo.083, whole genome"
},
{
"path": "test/data/raw_outputs/amrfinderplus/report_nucleotide.tsv",
"chars": 19240,
"preview": "Protein id\tContig id\tStart\tStop\tStrand\tElement symbol\tElement name\tScope\tType\tSubtype\tClass\tSubclass\tMethod\tTarget lengt"
},
{
"path": "test/data/raw_outputs/amrfinderplus/report_protein.tsv",
"chars": 3451,
"preview": "Protein id\tContig id\tStart\tStop\tStrand\tElement symbol\tElement name\tScope\tType\tSubtype\tClass\tSubclass\tMethod\tTarget lengt"
},
{
"path": "test/data/raw_outputs/amrplusplus/gene.tsv",
"chars": 11425,
"preview": "Sample\tGene\tHits\tGene Fraction\nKp85\tMEG_106|Drugs|Aminoglycosides|Aminoglycoside_N-acetyltransferases|AAC6-PRIME\t14\t86.4"
},
{
"path": "test/data/raw_outputs/ariba/report.tsv",
"chars": 96632,
"preview": "#ariba_ref_name\tref_name\tgene\tvar_only\tflag\treads\tcluster\tref_len\tref_base_assembled\tpc_ident\tctg\tctg_len\tctg_cov\tknown_"
},
{
"path": "test/data/raw_outputs/deeparg/output.mapping.ARG",
"chars": 5390679,
"preview": "#ARG\tquery-start\tquery-end\tread_id\tpredicted_ARG-class\tbest-hit\tprobability\tidentity\talignment-length\talignment-bitscore"
},
{
"path": "test/data/raw_outputs/deeparg/output.mapping.potential.ARG",
"chars": 972609,
"preview": "#ARG\tquery-start\tquery-end\tread_id\tpredicted_ARG-class\tbest-hit\tprobability\tidentity\talignment-length\talignment-bitscore"
},
{
"path": "test/data/raw_outputs/fargene/retrieved-genes-class_A-hmmsearched.out",
"chars": 3580,
"preview": "# --- fu"
},
{
"path": "test/data/raw_outputs/groot/report.tsv",
"chars": 1530,
"preview": "efpA.3003955.AL123456.3.3153038-3154631.5259\t82\t1593\t1582M11D\nMycobacterium_tuberculosis_16S.3003481.AL123456.3.1471846-"
},
{
"path": "test/data/raw_outputs/kmerresistance/results.res",
"chars": 6831,
"preview": "#Template\tScore\tExpected\tTemplate_length\tTemplate_Identity\tTemplate_Coverage\tQuery_Identity\tQuery_Coverage\tDepth\tq_value"
},
{
"path": "test/data/raw_outputs/mykrobe/report.json",
"chars": 3862,
"preview": "{\n \"my_sample_name\": {\n \"susceptibility\": {\n \"Ofloxacin\": {\n \"predict\": \"S\"\n "
},
{
"path": "test/data/raw_outputs/pointfinder/PointFinder_results.txt",
"chars": 1238,
"preview": "Mutation\tNucleotide change\tAmino acid change\tResistance\tPMID\n16s_rrna r.1192c>t\tc -> t\trna mutations\tspectinomycin\t24982"
},
{
"path": "test/data/raw_outputs/resfams/resfams.tblout",
"chars": 781240,
"preview": "# --- full sequence ---- --- best 1 domai"
},
{
"path": "test/data/raw_outputs/resfinder/ResFinder_results_tab.txt",
"chars": 2034,
"preview": "Resistance gene\tIdentity\tAlignment Length/Gene Length\tCoverage\tPosition in reference\tContig\tPosition in contig\tPhenotype"
},
{
"path": "test/data/raw_outputs/resfinder/data_resfinder.json",
"chars": 47222,
"preview": "{\"type\": \"software_result\", \"databases\": {\"ResFinder-2.4.0\": {\"type\": \"database\", \"database_name\": \"ResFinder\", \"databas"
},
{
"path": "test/data/raw_outputs/rgi/rgi.txt",
"chars": 17853,
"preview": "ORF_ID\tContig\tStart\tStop\tOrientation\tCut_Off\tPass_Bitscore\tBest_Hit_Bitscore\tBest_Hit_ARO\tBest_Identities\tARO\tModel_type"
},
{
"path": "test/data/raw_outputs/rgibwt/Kp11_bwtoutput.gene_mapping_data.txt",
"chars": 44717,
"preview": "ARO Term\tARO Accession\tReference Model Type\tReference DB\tAlleles with Mapped Reads\tReference Allele(s) Identity to CARD "
},
{
"path": "test/data/raw_outputs/srax/sraX_detected_ARGs.tsv",
"chars": 963,
"preview": "Locus ID\t# Sequences\tARG\tCoverage (%)\tIdentity (%)\tDrug class\tGene accession ID\tGene description\tAMR detection model\n1\t1"
},
{
"path": "test/data/raw_outputs/srst2/SAMN13064234_srst2_report.tsv",
"chars": 1763,
"preview": "Sample\tDB\tgene\tallele\tcoverage\tdepth\tdiffs\tuncertainty\tdivergence\tlength\tmaxMAF\tclusterid\tseqid\tannotation\nSRR10313716\tR"
},
{
"path": "test/data/raw_outputs/sstar/report.tsv",
"chars": 1186,
"preview": "blaSHV*\tblaSHV-100*\tNODE_11_length_157792_cov_39.2115\t95.111%\t900\t900\ncatB3?$\tcatB3?$\tNODE_51_length_1731_cov_22.5767\t99"
},
{
"path": "test/data/raw_outputs/staramr/pointfinder.tsv",
"chars": 367,
"preview": "Isolate ID\tGene\tPredicted Phenotype\tType\tPosition\tMutation\t%Identity\t%Overlap\tHSP Length/Total Length\tContig\tStart\tEnd\nS"
},
{
"path": "test/data/raw_outputs/staramr/resfinder.tsv",
"chars": 15057,
"preview": "Isolate ID\tGene\tPredicted Phenotype\t%Identity\t%Overlap\tHSP Length/Total Length\tContig\tStart\tEnd\tAccession\tSequence\nGDW22"
},
{
"path": "test/data/raw_outputs/tbprofiler/tbprofiler.json",
"chars": 21635,
"preview": "{\n \"qc\": {\n \"pct_reads_mapped\": 97.16,\n \"num_reads_mapped\": 4232866,\n \"median_coverage\": 121,\n \"gene_covera"
},
{
"path": "test/run_integration_test.sh",
"chars": 9082,
"preview": "#!/bin/bash\nset -e\n\nhamronize tbprofiler data/raw_outputs/tbprofiler/tbprofiler.json --format json --output hamronized_t"
},
{
"path": "test/test_interfaces.py",
"chars": 3889,
"preview": "import pytest\nimport json\nimport os\nimport csv\nfrom contextlib import contextmanager\nimport hAMRonization\nfrom hAMRoniza"
},
{
"path": "test/test_parsing_validity.py",
"chars": 42310,
"preview": "import pytest\nfrom contextlib import contextmanager\nimport hAMRonization\n\n\n@contextmanager\ndef not_raises(exception, msg"
}
]
// ... and 1 more files (download for full content)
About this extraction
This page contains the full source code of the pha4ge/hAMRonization GitHub repository, extracted and formatted as plain text for AI agents and large language models (LLMs). The extraction includes 98 files (11.9 MB), approximately 3.1M tokens, and a symbol index with 107 extracted functions, classes, methods, constants, and types. Use this with OpenClaw, Claude, ChatGPT, Cursor, Windsurf, or any other AI tool that accepts text input. You can copy the full output to your clipboard or download it as a .txt file.
Extracted by GitExtract — free GitHub repo to text converter for AI. Built by Nikandr Surkov.